Abstract
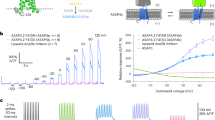
Accurate optical reporting of electrical activity in genetically defined neuronal populations is a long-standing goal in neuroscience. We developed Accelerated Sensor of Action Potentials 1 (ASAP1), a voltage sensor design in which a circularly permuted green fluorescent protein is inserted in an extracellular loop of a voltage-sensing domain, rendering fluorescence responsive to membrane potential. ASAP1 demonstrated on and off kinetics of ∼2 ms, reliably detected single action potentials and subthreshold potential changes, and tracked trains of action potential waveforms up to 200 Hz in single trials. With a favorable combination of brightness, dynamic range and speed, ASAP1 enables continuous monitoring of membrane potential in neurons at kilohertz frame rates using standard epifluorescence microscopy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
GenBank/EMBL/DDBJ
Referenced accessions
NCBI Reference Sequence
References
Magee, J.C. Dendritic integration of excitatory synaptic input. Nat. Rev. Neurosci. 1, 181–190 (2000).
Zecevic, D. et al. Imaging nervous system activity with voltage-sensitive dyes. Curr. Protoc. Neurosci. 6, Unit 6.17 (2003).
Castro-Alamancos, M.A. Cortical up and activated states: implications for sensory information processing. Neuroscientist 15, 625–634 (2009).
Branco, T. & Hausser, M. Synaptic integration gradients in single cortical pyramidal cell dendrites. Neuron 69, 885–892 (2011).
Puig, M.V., Ushimaru, M. & Kawaguchi, Y. Two distinct activity patterns of fast-spiking interneurons during neocortical UP states. Proc. Natl. Acad. Sci. USA 105, 8428–8433 (2008).
Royer, S. et al. Control of timing, rate and bursts of hippocampal place cells by dendritic and somatic inhibition. Nat. Neurosci. 15, 769–775 (2012).
Zhou, F.W. & Roper, S.N. Altered firing rates and patterns in interneurons in experimental cortical dysplasia. Cereb. Cortex 21, 1645–1658 (2011).
Chen, T.W. et al. Ultrasensitive fluorescent proteins for imaging neuronal activity. Nature 499, 295–300 (2013).
Murthy, V.N., Sejnowski, T.J. & Stevens, C.F. Dynamics of dendritic calcium transients evoked by quantal release at excitatory hippocampal synapses. Proc. Natl. Acad. Sci. USA 97, 901–906 (2000).
Kralj, J.M., Douglass, A.D., Hochbaum, D.R., Maclaurin, D. & Cohen, A.E. Optical recording of action potentials in mammalian neurons using a microbial rhodopsin. Nat. Methods 9, 90–95 (2012).
Gong, Y., Li, J.Z. & Schnitzer, M.J. Enhanced archaerhodopsin fluorescent protein voltage indicators. PLoS ONE 8, e66959 (2013).
Maclaurin, D., Venkatachalam, V., Lee, H. & Cohen, A.E. Mechanism of voltage-sensitive fluorescence in a microbial rhodopsin. Proc. Natl. Acad. Sci. USA 110, 5939–5944 (2013).
Akemann, W. et al. Imaging neural circuit dynamics with a voltage-sensitive fluorescent protein. J. Neurophysiol. 108, 2323–2337 (2012).
Barnett, L., Platisa, J., Popovic, M., Pieribone, V.A. & Hughes, T. A fluorescent, genetically encoded voltage probe capable of resolving action potentials. PLoS ONE 7, e43454 (2012).
Jin, L. et al. Single action potentials and subthreshold electrical events imaged in neurons with a fluorescent protein voltage probe. Neuron 75, 779–785 (2012).
Lam, A.J. et al. Improving FRET dynamic range with bright green and red fluorescent proteins. Nat. Methods 9, 1005–1012 (2012).
Lundby, A., Mutoh, H., Dimitrov, D., Akemann, W. & Knopfel, T. Engineering of a genetically encodable fluorescent voltage sensor exploiting fast Ci-VSP voltage-sensing movements. PLoS ONE 3, e2514 (2008).
Tsutsui, H. et al. Improved detection of electrical activity with a voltage probe based on a voltage-sensing phosphatase. J. Physiol. 591, 4427–4437 (2013).
Gautam, S.G., Perron, A., Mutoh, H. & Knopfel, T. Exploration of fluorescent protein voltage probes based on circularly permuted fluorescent proteins. Front. Neuroeng. 2, 14 (2009).
Staff, N.P., Jung, H.Y., Thiagarajan, T., Yao, M. & Spruston, N. Resting and active properties of pyramidal neurons in subiculum and CA1 of rat hippocampus. J. Neurophysiol. 84, 2398–2408 (2000).
Cao, G. et al. Genetically targeted optical electrophysiology in intact neural circuits. Cell 154, 904–913 (2013).
Jensen, M.Ø. et al. Mechanism of voltage gating in potassium channels. Science 336, 229–233 (2012).
Li, Q. et al. Structural mechanism of voltage-dependent gating in an isolated voltage-sensing domain. Nat. Struct. Mol. Biol. 21, 244–252 (2014).
Tian, L. et al. Imaging neural activity in worms, flies and mice with improved GCaMP calcium indicators. Nat. Methods 6, 875–881 (2009).
Dimitrov, D. et al. Engineering and characterization of an enhanced fluorescent protein voltage sensor. PLoS ONE 2, e440 (2007).
Cabantous, S., Terwilliger, T.C. & Waldo, G.S. Protein tagging and detection with engineered self-assembling fragments of green fluorescent protein. Nat. Biotechnol. 23, 102–107 (2005).
Akerboom, J. et al. Optimization of a GCaMP calcium indicator for neural activity imaging. J. Neurosci. 32, 13819–13840 (2012).
Akerboom, J. et al. Genetically encoded calcium indicators for multi-color neural activity imaging and combination with optogenetics. Front. Mol. Neurosci. 6, 2 (2013).
Siegel, M.S. & Isacoff, E.Y. A genetically encoded optical probe of membrane voltage. Neuron 19, 735–741 (1997).
Ataka, K. & Pieribone, V.A. A genetically targetable fluorescent probe of channel gating with rapid kinetics. Biophys. J. 82, 509–516 (2002).
Baker, B.J. et al. Three fluorescent protein voltage sensors exhibit low plasma membrane expression in mammalian cells. J. Neurosci. Methods 161, 32–38 (2007).
Perron, A., Mutoh, H., Launey, T. & Knopfel, T. Red-shifted voltage-sensitive fluorescent proteins. Chem. Biol. 16, 1268–1277 (2009).
Gong, Y., Wagner, M.J., Li, J.Z. & Schnitzer, M.J. Imaging neural spiking in brain tissue using FRET-opsin protein voltage sensors. Nat. Commun. advance online publication, 10.1038/ncomms4674 (22 April 2014).
Knöpfel, T. Genetically encoded optical indicators for the analysis of neuronal circuits. Nat. Rev. Neurosci. 13, 687–700 (2012).
Pédelacq, J.D., Cabantous, S., Tran, T., Terwilliger, T.C. & Waldo, G.S. Engineering and characterization of a superfolder green fluorescent protein. Nat. Biotechnol. 24, 79–88 (2006).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using microManager. Curr Protoc Mol Biol 14, Unit 14.20 (2010).
Jiang, M. & Chen, G. High Ca2+-phosphate transfection efficiency in low-density neuronal cultures. Nat. Protoc. 1, 695–700 (2006).
Watkins, R.J. et al. A novel interaction between FRMD7 and CASK: evidence for a causal role in idiopathic infantile nystagmus. Hum. Mol. Genet. 22, 2105–2118 (2013).
Wouterlood, F.G., Boekel, A.J., Kajiwara, R. & Belien, J.A. Counting contacts between neurons in 3D in confocal laser scanning images. J. Neurosci. Methods 171, 296–308 (2008).
Lin, M.Z. et al. Autofluorescent proteins with excitation in the optical window for intravital imaging in mammals. Chem. Biol. 16, 1169–1179 (2009).
Shcherbo, D. et al. Bright far-red fluorescent protein for whole-body imaging. Nat. Methods 4, 741–746 (2007).
Butko, M.T. et al. Fluorescent and photo-oxidizing TimeSTAMP tags track protein fates in light and electron microscopy. Nat. Neurosci. 15, 1742–1751 (2012).
Huang, Y.M. & Bystroff, C. Complementation and reconstitution of fluorescence from circularly permuted and truncated green fluorescent protein. Biochemistry 48, 929–940 (2009).
Acknowledgements
We thank the following for providing rats or dissociated neurons: H. Park and Y. Geng (Stanford), M. Hintze and S. Ganesan (Stanford), and C. Ramakrishnan and H. Swanson (Stanford). We also thank X. Ding (Tsinghua University) for assistance with cloning ASAP1 variants with cpsepHluorin A227D, Y. Geng (Stanford) for providing microfluidic chambers, J. Chu (Stanford) for a purified preparation of Clover GFP, V. Pieribone (Yale University) for the generous gift of GgVSD(153Q), DrVSD(R153Q) and XlVSD(R152Q), L. Oltrogge (Stanford) for the generous gift of cpsfGFP-OPT, J. Bant (Stanford) for advice on electrophysiological recordings, and members of the Lin laboratory for comments on the manuscript. This work was supported by DARPA (M.Z.L., M.J.S.), National Science Foundation grant 1134416 (F.S.-P., M.Z.L.), a Stanford Graduate Fellowship (J.D.M.), a Walter V. and Idun Berry Postdoctoral Fellowship (Y.Y.), a Stanford University Bio-X Interdisciplinary Initiatives Project grant (M.Z.L., M.J.S.), the Stanford CNC Program (Y.G., J.D.M., M.J.S.), the Howard Hughes Medical Institute (M.J.S.) and the National Academy of Sciences Keck Futures Initiative (Y.G., J.D.M., M.J.S.). M.Z.L. receives funding from the Rita Allen Foundation.
Author information
Authors and Affiliations
Contributions
M.Z.L. conceived the study. F.S.-P. and M.Z.L. designed ASAP1. F.S.-P., J.D.M., Y.Y. and Y.G. designed and performed experiments, and analyzed data. M.Z.L. and M.J.S. provided ideas and advice. F.S.-P. and M.Z.L. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
F.S.-P. and M.Z.L. have filed a patent application for a voltage sensor design based on the results reported in this paper.
Integrated supplementary information
Supplementary Figure 1 Development of ASAP1-class voltage sensors.
(a) Alignment of the S3-S4 loop region of voltage-sensing domains (VSDs) from voltage sensitive phosphatases of four different species. Low, intermediate and high levels of homology are represented by white, grey and black background color, respectively. The locations of S3 and S4 were predicted by the Transmembrane Prediction Tool of Geneious v.7 (Biomatters). (b) G. gallus VSD with the R153Q mutation and an insertion of circularly permuted green fluorescent protein (cpGFP) from GCaMP3 at 4, 5, and 6 amino acids downstream of S3 exhibited bright membrane-localized fluorescence in HEK293A cells. Insertion at other positions caused poor membrane localization or dimness. Scale bar, 10 μm. (c) Fluorescence response in HEK293A cells to voltage steps from –150 mV to +50 mV relative to the holding potential of –70 mV for three sensor variants with different insertion points of cpGFP into G. gallus VSD (GgVSD). All three variants produced similar fluorescence responses, with an ~3% decrease in intensity from –70 mV to +30 mV. Samples sizes (n) correspond to individual HEK293A cells. Error bars are mean ± standard error of the mean (SEM). (d) We constructed sensor variants in which the fluorescent protein, originally cpGFP from GCaMP3, was replaced with other circularly-permuted GFPs. All fluorescent proteins were inserted between Ala-147 and Thr-148 of GgVSD. To normalize for variability in transfection efficiency or expression in HEK293A cells, each sensor was coexpressed with mCherry-CAAX using an internal ribosome entry site (see Supplementary Fig. 5a). For each sensor, the green and red fluorescence images were identically scaled and merged. The relative ratio of green to red fluorescence, an indication of sensor brightness, was larger for the sensor containing the OPT variant of superfolder GFP (cpsfGFP-OPT). Substitution with a circularly permuted version of Clover (cpClover) or of the superecliptic pHluorin A227D from ArcLight (cpsepHluorin A227D) largely abolished membrane-localized GFP fluorescence. Findings were reproducible across multiple (n > 5) fields of view, with representative images shown here. Scale bar, 10 μm.
Supplementary Figure 2 Voltage response characteristics and membrane localization of ASAP1 variants with alternative voltage-sensing domains.
(a) Substituting the voltage-sensing domain (VSD) from Gallus gallus with the VSD of Xenopus laevis or Danio rerio produced membrane-localized fluorescence in HEK293A cells, while substituting with the VSD of Ciona intestinalis resulted in loss of membrane localization for all insertion positions tested, 1-12 amino acids downstream of S3. Reverting the R153Q mutation in ASAP1 produced comparable membrane localization to the R153Q sensor. Scale bar, 10 μm. (b) ASAP1 variants with VSD from alternative species show reduced fluorescence dynamic range in HEK293A cells to voltage steps from –150 mV to +50 mV relative to –70 mV. Holding potential was –11 mV, close to HEK293A resting potential (typically –10 to –20 mV). Samples sizes (n) correspond to individual HEK293A cells. Error bars are mean ± SEM. (c) Reverting the R153Q mutation in ASAP did not improve fluorescence dynamic range in HEK293A to voltage steps from –150 mV to +50 mV relative to –70 mV. Holding potential was –11 mV. For all panels, cspfGFP correspond to the OPT variant used in ASAP1. Samples sizes (n) correspond to individual HEK293A cells. Error bars, mean ± SEM.
Supplementary Figure 3 Voltage response characteristics and membrane localization of ASAP1 variants with different VSD-GFP junctions.
(a) Changing termini for circular permutants of sfGFP-OPT did not disrupt efficient membrane localization in HEK293A cells. Numbers indicate location of the new N- and C-termini using non-permuted sfGFP numbering. (b) These variants exhibited reduced responses in HEK293A to voltage steps. (c) Same data replotted to better show reversal of the fluorescence response in sensors with cpsfGFP-OPT 147-146, 148-147 and 149-147. (d) Variants with deletion of one amino acid at either side of the VSD-GFP or GFP-VSD junction showed efficient membrane localization in HEK293A cells. (e) Deletion of the N-terminal amino acid of cpsfGFP-OPT (Phe-145) dramatically reduced fluorescence response in HEK293A cells. Deletion of any of the other three junctional amino acids did not improve fluorescence responses within the physiological range (–100mV to +50mV). cspfGFP correspond to the OPT variant used in ASAP1. For all panels, samples sizes (n) correspond to individual HEK293A cells. Error bars are mean ± SEM
Supplementary Figure 4 Voltage response characteristics and membrane localization of ASAP1 variants with conservative mutations at the cspfGFP-OPT amino terminus.
(a) Conservative mutations of the Phe-145 of cpsfGFP-OPT immediately C-terminal to the VSD-GFP junction to leucine (F145L), methionine (F145M) or tyrosine (F145Y) produced qualitatively comparable membrane localization to ASAP1 in HEK293A cells. Scale bar, 10 μm. (b) F145I, F145L, F145M, and F145Y reduced fluorescence responses in HEK293A cells to voltage steps from –150 to +50 mV. Samples sizes (n) correspond to individual cells. Error bars are mean ± SEM.
Supplementary Figure 5 Quantification of ASAP1 membrane localization in dissociated neuronal cultures.
(a) Schematic of plasmid used for the membrane localization experiments. An internal ribosome entry site (IRES) allows the CAG promoter to drive expression of both ASAP1 and a fusion of mCherry RFP and a farnesylation motif (CAAX). (b) ASAP1 levels, detected by antibody staining, were larger following neuron membrane permeabilization, suggesting that a fraction of sensor molecules remained in the cytoplasm. We used mCherry-CAAX fluorescence as our measure of plasmid expression. n = 22 fixed hippocampal neurons per condition (intact or permeabilized). All neurons were from the same litter. (c) Same data as in (b), but without normalization of fluorescence levels with cell size. (d) Across all cells tested, expression-adjusted ASAP1 levels in intact cells were 42 ± 5 % (mean ± SEM) of those following permeabilization (p < 0.001). Normalization for expression level was performed by dividing fluorescence values for antibody-conjugated ASAP1 with the fluorescence values of mCherry-CAAX from the same cell. (e) We quantified the membrane localization of mCherry-CAAX in permeabilized cells (n = 22 fixed neurons) by measuring red fluorescence at the plasma membrane (cell perimeter) and cytoplasm. We quantified ASAP1 localization in the same cells using the boundaries for the cell membrane and cytoplasm defined on the mCherry-CAAX image. We observed that membrane localization of ASAP1 was nearly as efficient (91.0 ± 1.5 %, mean ± SEM) as mCherry-CAAX in permeabilized cells.
Supplementary Figure 6 Quantification of membrane capacitance of ASAP1-transfected neurons in dissociated cultures.
No significant increase in membrane capacitance was observed upon increasing ASAP1 expression, determined as GFP fluorescence intensity per pixel at the membrane (a) or total cellular GFP fluorescence (b). Cells with zero ASAP1 fluorescence correspond to control neurons transfected with a plasmid expressing cytoplasmic Clover GFP (n = 5 control neurons and 12 ASAP1-expressing neurons). We used the membrane resistance (Rm) as a coarse correlate of cell size, and reploted the data from (a) and (b) after normalizing capacitance with Rm (c,d). Linear trendlines and correlation coefficients are y = 0.069x + 71.35, R2= 0.01 (a); y = −0.180x + 75.55, R2= 0.06 (b); y = 0.002x + 0.285, R2= 0.07 (c); y = −0.002x + 0.363, R2= 0.06 (d). (e) Over all cells tested, membrane capacitance of ASAP1-expressing neurons was 74.5 ± 6.4 pF (mean ± standard error of the mean, n = 12 neurons), statistically indistinguishable from control neurons (69.9 ± 6.4 pF, n = 5; p = 0.65). (f) Similarly, the Rm-normalized membrane capacitance of ASAP1-expressing neurons was 0.36 ± 0.7 pF/MΩ, statistically indistinguishable from control neurons (0.29 ± 0.05 pF/MΩ, p = 0.65, 2-tail Mann-Whitney U test). ASAP1-expressing cells tested in this experiment were estimated to contain between ~20–300 sensor molecules per um2 of membrane, with an average of 106 ± 21 (Methods).
Supplementary Figure 7 ASAP1's faster kinetics allow improved resolution of a 100-Hz, three-AP waveform sequence in cultured hippocampal neurons.
ASAP1 (top) and ArcLight Q239 (bottom) fluorescence responses to a three-AP waveform sequence following a series of overlapping subthreshold depolarizations. Command voltage is shown on the left-most plots. Command AP spike full width at half-maximum was 1.8 ms. We collected fluorescence traces for this figure and Fig. 3c from distinct neurons.
Supplementary Figure 8 Imaging spontaneous neuronal activity in cultured hippocampal neurons with ASAP1 and ArcLight Q239.
Fluorescence responses of ASAP1 (a) and ArcLight Q239 (b) to spontaneous electrical activity. ArcLight Q239 did not produce an apparent fluorescence response to the single AP at ∼3,800ms. Fluorescence traces are unfiltered. We collected fluorescence traces for this figure and Fig. 4a from distinct neurons.
Supplementary Figure 9 ASAP1 photobleaching kinetics.
Dissociated hippocampal neurons expressing ASAP1 were illuminated continuously with 470- to 490-nm light at 0.12 mW/mm2. Each marker color corresponds to one of four individual neurons imaged every 10 s for 4.5-15 min. Fluorescence was normalized to 1.0 at t = 0. Photobleaching time constants (τ) were estimated from mono-exponential fits (dashed lines) as 41.7 min (red trace), 35.7 min (blue trace), 32.3 min (green trace), and 30.3 min (gray trace).
Supplementary Figure 10 Continuous multi-minute imaging of neuronal activity with ASAP1.
Fluorescence responses of ASAP1 to spontaneous electrical activity in two cultured hippocampal neurons at the beginning (top traces) and end (bottom traces) of 15 min of continuous illumination. Illumination power was 0.023 mW/mm2 (a) and 0.036 mW/mm2 (b). Traces were not corrected for photobleaching over the 30-s interval but were noise-filtered with LOWESS smoothing. We collected fluorescence traces for this figure and Fig. 4c from distinct neurons.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 22345 kb)
Supplementary Methods Checklist
(PDF 396 kb)
Rights and permissions
About this article
Cite this article
St-Pierre, F., Marshall, J., Yang, Y. et al. High-fidelity optical reporting of neuronal electrical activity with an ultrafast fluorescent voltage sensor. Nat Neurosci 17, 884–889 (2014). https://doi.org/10.1038/nn.3709
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nn.3709
This article is cited by
-
Widefield imaging of rapid pan-cortical voltage dynamics with an indicator evolved for one-photon microscopy
Nature Communications (2023)
-
A minimal-complexity light-sheet microscope maps network activity in 3D neuronal systems
Scientific Reports (2022)
-
Next generation genetically encoded fluorescent sensors for serotonin
Nature Communications (2022)
-
Monitoring of compound resting membrane potentials of cell cultures with ratiometric genetically encoded voltage indicators
Communications Biology (2021)
-
Enabling comprehensive optogenetic studies of mouse hearts by simultaneous opto-electrical panoramic mapping and stimulation
Nature Communications (2021)