Abstract
We report an atmospheric pressure (AP) matrix-assisted laser desorption/ionization (MALDI) mass spectrometry imaging (MSI) setup with a lateral resolution of 1.4 μm, a mass resolution greater than 100,000, and accuracy below ±2 p.p.m. We achieved this by coupling a focusing objective with a numerical aperture (NA) of 0.9 at 337 nm and a free working distance of 18 mm in coaxial geometry to an orbitrap mass spectrometer and optimizing the matrix application. We demonstrate improvement in image contrast, lateral resolution, and ion yield per unit area compared with a state-of-the-art commercial MSI source. We show that our setup can be used to detect metabolites, lipids, and small peptides, as well as to perform tandem MS experiments with 1.5-μm2 sampling areas. To showcase these capabilities, we identified subcellular lipid, metabolite, and peptide distributions that differentiate, for example, cilia and oral groove in Paramecium caudatum.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Spengler, B. Mass spectrometry imaging of biomolecular information. Anal. Chem. 87, 64–82 (2015).
Chughtai, K. & Heeren, R.M. Mass spectrometric imaging for biomedical tissue analysis. Chem. Rev. 110, 3237–3277 (2010).
Norris, J.L. & Caprioli, R.M. Analysis of tissue specimens by matrix-assisted laser desorption/ionization imaging mass spectrometry in biological and clinical research. Chem. Rev. 113, 2309–2342 (2013).
Dreisewerd, K. Recent methodological advances in MALDI mass spectrometry. Anal. Bioanal. Chem. 406, 2261–2278 (2014).
Spengler, B., Hubert, M. & Kaufmann, R. Maldi ion imaging and biological ion imaging with a new scanning UV-laser microprobe. In Proc. 42nd ASMS Conference on Mass Spectrometry and Allied Topics abstract 1041 (American Society for Mass Spectrometry, 1994).
Laiko, V.V., Baldwin, M.A. & Burlingame, A.L. Atmospheric pressure matrix-assisted laser desorption/ionization mass spectrometry. Anal. Chem. 72, 652–657 (2000).
Wiseman, J.M., Ifa, D.R., Song, Q. & Cooks, R.G. Tissue imaging at atmospheric pressure using desorption electrospray ionization (DESI) mass spectrometry. Angew. Chem. Int. Edn. Engl. 45, 7188–7192 (2006).
Kettling, H. et al. MALDI mass spectrometry imaging of bioactive lipids in mouse brain with a Synapt G2-S mass spectrometer operated at elevated pressure: improving the analytical sensitivity and the lateral resolution to ten micrometers. Anal. Chem. 86, 7798–7805 (2014).
Jones, E.E. et al. MALDI imaging mass spectrometry profiling of proteins and lipids in clear cell renal cell carcinoma. Proteomics 14, 924–935 (2014).
Northen, T.R. et al. Clathrate nanostructures for mass spectrometry. Nature 449, 1033–1036 (2007).
Bouschen, W., Schulz, O., Eikel, D. & Spengler, B. Matrix vapor deposition/recrystallization and dedicated spray preparation for high-resolution scanning microprobe matrix-assisted laser desorption/ionization imaging mass spectrometry (SMALDI-MS) of tissue and single cells. Rapid Commun. Mass Spectrom. 24, 355–364 (2010).
Schober, Y., Guenther, S., Spengler, B. & Römpp, A. Single cell matrix-assisted laser desorption/ionization mass spectrometry imaging. Anal. Chem. 84, 6293–6297 (2012).
Spraggins, J.M. et al. MALDI FTICR IMS of intact proteins: using mass accuracy to link protein images with proteomics data. J. Am. Soc. Mass Spectrom. 26, 974–985 (2015).
Römpp, A., Guenther, S., Takats, Z. & Spengler, B. Mass spectrometry imaging with high resolution in mass and space (HR(2) MSI) for reliable investigation of drug compound distributions on the cellular level. Anal. Bioanal. Chem. 401, 65–73 (2011).
Aichler, M. & Walch, A. MALDI imaging mass spectrometry: current frontiers and perspectives in pathology research and practice. Lab. Invest. 95, 422–431 (2015).
Römpp, A. et al. Histology by mass spectrometry: label-free tissue characterization obtained from high-accuracy bioanalytical imaging. Angew. Chem. Int. Edn. Engl. 49, 3834–3838 (2010).
Eberlin, L.S. et al. Alteration of the lipid profile in lymphomas induced by MYC overexpression. Proc. Natl. Acad. Sci. USA 111, 10450–10455 (2014).
Bhandari, D.R., Schott, M., Römpp, A., Vilcinskas, A. & Spengler, B. Metabolite localization by atmospheric pressure high-resolution scanning microprobe matrix-assisted laser desorption/ionization mass spectrometry imaging in whole-body sections and individual organs of the rove beetle Paederus riparius. Anal. Bioanal. Chem. 407, 2189–2201 (2015).
Shariatgorji, M. et al. Direct targeted quantitative molecular imaging of neurotransmitters in brain tissue sections. Neuron 84, 697–707 (2014).
Passarelli, M.K., Ewing, A.G. & Winograd, N. Single-cell lipidomics: characterizing and imaging lipids on the surface of individual Aplysia californica neurons with cluster secondary ion mass spectrometry. Anal. Chem. 85, 2231–2238 (2013).
Cajka, T. & Fiehn, O. Toward merging untargeted and targeted methods in mass spectrometry-based metabolomics and lipidomics. Anal. Chem. 88, 524–545 (2016).
Angelo, M. et al. Multiplexed ion beam imaging of human breast tumors. Nat. Med. 20, 436–442 (2014).
Abbassi-Ghadi, N. et al. Discrimination of lymph node metastases using desorption electrospray ionisation-mass spectrometry imaging. Chem. Commun. (Camb.) 50, 3661–3664 (2014).
Lanni, E.J. et al. MALDI-guided SIMS: multiscale imaging of metabolites in bacterial biofilms. Anal. Chem. 86, 9139–9145 (2014).
Khalil, S.M., Römpp, A., Pretzel, J., Becker, K. & Spengler, B. Phospholipid topography of whole-body sections of the Anopheles stephensi mosquito, characterized by high-resolution atmospheric-pressure scanning microprobe matrix-assisted laser desorption/ionization mass spectrometry imaging. Anal. Chem. 87, 11309–11316 (2015).
Li, B., Bhandari, D.R., Janfelt, C., Römpp, A. & Spengler, B. Natural products in Glycyrrhiza glabra (licorice) rhizome imaged at the cellular level by atmospheric pressure matrix-assisted laser desorption/ionization tandem mass spectrometry imaging. Plant J. 80, 161–171 (2014).
Bjarnholt, N., Li, B., D'Alvise, J. & Janfelt, C. Mass spectrometry imaging of plant metabolites—principles and possibilities. Nat. Prod. Rep. 31, 818–837 (2014).
Jaskolla, T.W., Lehmann, W.-D. & Karas, M. 4-Chloro-α-cyanocinnamic acid is an advanced, rationally designed MALDI matrix. Proc. Natl. Acad. Sci. USA 105, 12200–12205 (2008).
Soltwisch, J. et al. Mass spectrometry imaging with laser-induced postionization. Science 348, 211–215 (2015).
Yang, J. & Caprioli, R.M. Matrix sublimation/recrystallization for imaging proteins by mass spectrometry at high spatial resolution. Anal. Chem. 83, 5728–5734 (2011).
Liebisch, G. et al. Shorthand notation for lipid structures derived from mass spectrometry. J. Lipid Res. 54, 1523–1530 (2013).
Race, A.M. & Bunch, J. Optimisation of colour schemes to accurately display mass spectrometry imaging data based on human colour perception. Anal. Bioanal. Chem. 407, 2047–2054 (2015).
Zhao, P., Pedersen, M., Hardeberg, J.Y. & Thomas, J.-B. Measuring the relative image contrast of projection displays. J. Imaging Sci. Technol. 59, 30404-1–30404-13 (2015).
Zhang, W. & Bergholm, F. Multi-scale blur estimation and edge type classification for scene analysis. Int. J. Comput. Vis. 24, 219–250 (1997).
Qiao, H., Spicer, V. & Ens, W. The effect of laser profile, fluence, and spot size on sensitivity in orthogonal-injection matrix-assisted laser desorption/ionization time-of-flight mass spectrometry. Rapid Commun. Mass Spectrom. 22, 2779–2790 (2008).
Kaneshiro, E.S. Lipids of Paramecium. J. Lipid Res. 28, 1241–1258 (1987).
Hell, S.W. Nanoscopy with focused light (Nobel lecture). Angew. Chem. Int. Edn. Engl. 54, 8054–8066 (2015).
Ogrinc Potocčˇnik, N., Porta, T., Becker, M., Heeren, R.M.A. & Ellis, S.R. Use of advantageous, volatile matrices enabled by next-generation high-speed matrix-assisted laser desorption/ionization time-of-flight imaging employing a scanning laser beam. Rapid Commun. Mass Spectrom. 29, 2195–2203 (2015).
Zenobi, R. Single-cell metabolomics: analytical and biological perspectives. Science 342, 1243259 (2013).
Kompauer, M., Heiles, S. & Spengler, B. AP-MALDI MSI of lipids in mouse brain tissue sections. Protoc. Exch. http://dx.doi.org/10.1038/protex.2016.074 (2016).
Sjövall, P., Johansson, B. & Lausmaa, J. Localization of lipids in freeze-dried mouse brain sections by imaging TOF-SIMS. Appl. Surf. Sci. 252, 6966–6974 (2006).
Zavalin, A. et al. Direct imaging of single cells and tissue at sub-cellular spatial resolution using transmission geometry MALDI MS. J. Mass Spectrom. 47, 1473–1481 (2012).
Hankin, J.A., Barkley, R.M. & Murphy, R.C. Sublimation as a method of matrix application for mass spectrometric imaging. J. Am. Soc. Mass Spectrom. 18, 1646–1652 (2007).
Koestler, M. et al. A high-resolution scanning microprobe matrix-assisted laser desorption/ionization ion source for imaging analysis on an ion trap/Fourier transform ion cyclotron resonance mass spectrometer. Rapid Commun. Mass Spectrom. 22, 3275–3285 (2008).
Schürenberg, M., Schulz, T., Dreisewerd, K. & Hillenkamp, F. Matrix-assisted laser desorption/ionization in transmission geometry. Rapid Commun. Mass Spectrom. 10, 1873–1880 (1996).
Hillenkamp, F., Unsöld, E., Kaufmann, R. & Nitsche, R. Laser microprobe mass analysis of organic materials. Nature 256, 119–120 (1975).
Sun, L., Zhu, G. & Dovichi, N.J. Comparison of the LTQ-Orbitrap Velos and the Q-Exactive for proteomic analysis of 1-1000ng RAW 264.7 cell lysate digests. Rapid Commun. Mass Spectrom. 27, 157–162 (2013).
Paschke, C. et al. Mirion—a software package for automatic processing of mass spectrometric images. J. Am. Soc. Mass Spectrom. 24, 1296–1306 (2013).
Acknowledgements
Financial support from the Deutsche Forschungsgemeinschaft (DFG) under grant Sp314/13-1 is gratefully acknowledged. S.H. is grateful to the German National Academy of Sciences Leopoldina (LPDR 2014-01) for a postdoctoral scholarship. The authors are grateful to W. Kummer (Institute of Anatomy and Cell Biology, Justus Liebig University Giessen, Giessen, Germany) and his group members for providing mouse brain and kidney samples.
Author information
Authors and Affiliations
Contributions
B.S. supervised the project; B.S., S.H. and M.K. designed the experiment; B.S. designed the new instrumentation, and M.K. and S.H. set it up and performed all experiments; M.K. and S.H. performed the data analysis; and S.H., M.K. and B.S. discussed the findings and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
B.S. is a consultant of TransMIT GmbH Giessen. The PhD work of M.K. is funded by TransMIT GmbH.
Integrated supplementary information
Supplementary Figure 1 Laser focus characterization.
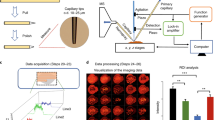
a) Experimental setup to visualize the laser beam profile at the focal plane of the centrally bored new objective. (b) Single laser pulse focus at 337 nm visualized with a calibrated beam profile system. (c) Laser ablation craters on red dye after moving the stage with 10 μm increments in x and y direction. 15 laser pulses with ~20 nJ per pulse were applied to each spot. (d) Focal depth determination on red pencil dye with 10 laser pulses of 20 nJ per pulse. The focal plane was reached between 4-6 μm z-distance increase.
Supplementary Figure 2 Matrix application methods.
(a) Optical images of mouse brain tissue sections after matrix application. Sublimation of CHCA (left); sublimation+recrystallization of CHCA (right). (b) Optical images of mouse brain tissue sections after matrix application. Pneumatic spraying of DHB (left) and CHCA (right).
Supplementary Figure 3 Matrix application method and intensity comparison.
MALDI mass spectra averaged over 100 pixels, obtained from mouse brain tissue using 20 laser pulses per pixel with 10 nJ per pulse, thereby probing ~1.5 μm2 of the tissue per ablation spot. Lipid intensities obtained by (MA)LDI MS from mouse brain tissue (a) without matrix, (b) after sublimation coating with DHB matrix, (c) after sublimation coating followed by recrystallization of DHB matrix and (d) after pneumatic spraying of DHB matrix. Selected lipid ion signals were labeled with the corresponding m/z values. Absolute signal intensity gain is reflected by scale values.
Supplementary Figure 4 On-tissue lateral resolution, image contrast and ion yield per unit area.
MS gray scale images of lipid ions shown in Fig. 3 of the manuscript. Mass spectra obtained from a single-pixel during MSI experiments, shown for the AP-SMALDI10 and the experimental AP MSI setup, respectively. Red lines indicate the data points used for contrast and edge width analysis.
Supplementary Figure 5 Single-cell MSI measurement and corresponding optical images of Paramecium caudatum.
(a) Optical Image of Paramecium caudatum revealing irregularities on the surface of the organism. (b-d) Corresponding gray scale MS images (100x100 pixels) obtained with 3 μm step size. A total of 30 laser pulses per spot with 15 nJ per pulse were applied. [DG(31:0)+NH4]+ (m/z 572.5240; red), [PC(34:1)+Na]+ (m/z 782.5668; green) and [Cer(d35:2)+H]+ (m/z 550.5195; blue) were selected for the three images. (e) Paramecium caudatum after the MSI measurement shown in Fig. 5a. Scale bars in all images are 100 μm.
Supplementary Figure 6 MS images of metabolites and peptides from Paramecium caudatum at 3-μm step size (100 × 100 pixels).
(1-8) Gray scale MS images: the ion signal assignment is given in Supplementary Table 2 according to the number code. Notably, metabolites, peptides and phospholipids were detected that are solely localized in the Paramecium caudatum organism. Signals assigned to the same analyte with differing ionic attachment show similar lateral distributions (see (1) and (2)). Scale bars in all images are 100 μm.
Supplementary Figure 7 Grayscale MS, optical, and RGB MS images of Paramecium caudatum and Rotifera.
(a) Optical image before MSI measurement of Paramecium caudatum in the process of being devoured by a Rotifera predator. (b) RGB image (175x150 pixels) obtained with 2 μm step size from Paramecium caudatum being devoured by Rotifera. A total of 30 laser pulses per spot with 10 nJ per pulse were used. [PE(35:2)+H]+ (m/z 730.5377; red) and [PC(34:1)+Na]+ (m/z 782.5668; green) were superimposed. (c-d) Corresponding gray scale images highlighting that the imaged lipids are characteristic for Paramecium caudatum and Rotifera, respectively. Scale bars in all images are 100 μm.
Supplementary Figure 8 Schematic of the experimental AP-MALDI MSI source.
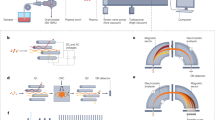
The setup features a (1) N2-laser system; (2) a beam shaping setup; (3) 45° mirror with central bore; (4) newly developed focusing objective with central bore; (5) an aperture; (6) a high precision sample movement stage with sample holder and (7, 8) a camera-mirror assembly to visualize the sample surface. Upon laser irradiation ions are ejected in reflection-mode and transferred into the inlet capillary (9). A DC voltage (HV) is applied between sample holder and inlet capillary of the mass spectrometer. The entire setup is attached to a Q Exactive mass spectrometer. Ion source and mass spectrometer are not to scale.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Tables 1–3, Supplementary Notes 1–3 and Supplementary Protocol. (PDF 6624 kb)
Source data
Rights and permissions
About this article
Cite this article
Kompauer, M., Heiles, S. & Spengler, B. Atmospheric pressure MALDI mass spectrometry imaging of tissues and cells at 1.4-μm lateral resolution. Nat Methods 14, 90–96 (2017). https://doi.org/10.1038/nmeth.4071
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.4071
This article is cited by
-
Metabolic heterogeneity in cancer
Nature Metabolism (2024)
-
Gel-assisted mass spectrometry imaging enables sub-micrometer spatial lipidomics
Nature Communications (2024)
-
Reproducible mass spectrometry data processing and compound annotation in MZmine 3
Nature Protocols (2024)
-
Multiscale biochemical mapping of the brain through deep-learning-enhanced high-throughput mass spectrometry
Nature Methods (2024)
-
Mass spectrometry imaging for spatially resolved multi-omics molecular mapping
npj Imaging (2024)