Abstract
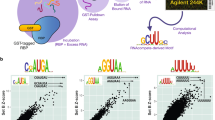
Human nuclei contain three RNA polymerases (I, II and III) that transcribe different groups of genes; the active forms of all three are difficult to isolate because they are bound to the substructure. Here we describe a purification approach for isolating active RNA polymerase complexes from mammalian cells. After isolation, we analyzed their protein content by mass spectrometry. Each complex represents part of the core of a transcription factory. For example, the RNA polymerase II complex contains subunits unique to RNA polymerase II plus various transcription factors but shares a number of ribonucleoproteins with the other polymerase complexes; it is also rich in polymerase II transcripts. We also describe a native chromosome conformation capture method to confirm that the complexes remain attached to the same pairs of DNA templates found in vivo.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Roeder, R.G. The eukaryotic transcriptional machinery: complexities and mechanisms unforeseen. Nat. Med. 9, 1239–1244 (2003).
Cramer, P. et al. Structure of eukaryotic RNA polymerases. Annu. Rev. Biophys. 37, 337–352 (2008).
Das, R. et al. SR proteins function in coupling RNAP II transcription to pre-mRNA splicing. Mol. Cell 26, 867–881 (2007).
Shi, Y. et al. Molecular architecture of the human pre-mRNA 3′ processing complex. Mol. Cell 33, 365–376 (2009).
Cook, P.R. A model for all genomes; the role of transcription factories. J. Mol. Biol. 395, 1–10 (2010).
Chakalova, L. & Fraser, P. Organization of transcription. Cold Spring Harb. Perspect. Biol. 2, a000729 (2010).
Sutherland, H. & Bickmore, W.A. Transcription factories: gene expression in unions? Nat. Rev. Genet. 10, 457–466 (2009).
Jackson, D.A., Iborra, F.J., Manders, E.M.M. & Cook, P.R. Numbers and organization of RNA polymerases, nascent transcripts and transcription units in HeLa nuclei. Mol. Biol. Cell 9, 1523–1536 (1998).
Kimura, H., Tao, Y., Roeder, R.G. & Cook, P.R. Quantitation of RNA polymerase II and its transcription factors in an HeLa cell: little soluble holoenzyme but significant amounts of polymerases attached to the nuclear substructure. Mol. Cell. Biol. 19, 5383–5392 (1999).
Jackson, D.A. & Cook, P.R. Transcription occurs at a nucleoskeleton. EMBO J. 4, 919–925 (1985).
Ahmad, Y., Boisvert, F.M., Gregor, P., Cobley, A. & Lamond, A.I. NOPdb: Nucleolar Proteome Database–2008 update. Nucleic Acids Res. 37, D181–D184 (2009).
Eskiw, C.H., Rapp, A., Carter, D.R.F. & Cook, P.R. RNA polymerase II activity is located on the surface of ∼87 nm protein-rich transcription factories. J. Cell Sci. 121, 1999–2007 (2008).
Novakova, Z., Man, P., Novak, P., Hozak, P. & Hodny, Z. Separation of nuclear protein complexes by blue native polyacrylamide gel electrophoresis. Electrophoresis 2, 1277–1287 (2006).
Trudgian, D.C. et al. CPFP – The Oxford Central Proteomics Facility Pipeline. Clin. Proteomics 5 (suppl. 1), 94 (2009).
Griffin, N.M. et al. Label-free, normalized quantification of complex mass spectrometry data for proteomic analysis. Nat. Biotechnol. 28, 83–89 (2010).
Hopper, A.K., Pai, D.A. & Engelke, D.R. Cellular dynamics of tRNAs and their genes. FEBS Lett. 584, 310–317 (2010).
Iborra, F.J., Escargueil, A.E., Kwek, K.Y., Akoulitchev, A. & Cook, P.R. Molecular cross-talk between the transcription, translation, and nonsense-mediated decay machineries. J. Cell Sci. 117, 899–906 (2004).
Dekker, J., Rippe, K., Dekker, M. & Kleckner, N. Capturing chromosome conformation. Science 295, 1306–1311 (2002).
Cullen, K.E., Kladde, M.P. & Seyfred, M.A. Interaction between transcription regulatory regions of prolactin chromatin. Science 261, 203–206 (1993).
Papantonis, A. et al. Active RNA polymerases: mobile or immobile molecular machines? PLoS Biol. 8, e1000419 (2010).
Zheng, B., Han, M., Bernier, M. & Wen, J.K. Nuclear actin and actin-binding proteins in the regulation of transcription and gene expression. FEBS J. 276, 2669–2685 (2009).
Hou, C. & Corces, V.G. Nups take leave of the nuclear envelope to regulate transcription. Cell 140, 306–308 (2010).
Nadano, D., Aoki, C., Yoshinaka, T., Irie, S. & Sato, T.A. Electrophoretic characterization of ribosomal subunits and protein in apoptosis: specific downregulation of S11 in staurosporine-treated human breast carcinoma cells. Biochemistry 40, 15184–15193 (2001).
Maclean, B., Eng, J.K., Beavis, R.C. & McIntosh, M. General framework for developing and evaluating database scoring algorithms using the TANDEM search engine. Bioinformatics 22, 2830–2832 (2006).
Geer, L.Y. et al. Open mass spectrometry search algorithm. J. Proteome Res. 3, 958–964 (2004).
Keller, A., Nesvizhskii, A.I., Kolker, E. & Aebersold, R. Empirical statistical model to estimate the accuracy of peptide identifications made by MS/MS and database search. Anal. Chem. 74, 5383–5392 (2002).
Shteynberg, D. et al. iProphet: Improved validation of peptide and protein IDs in the trans-proteomic pipeline. Poster session at: HUPO 7th Annual World Congress (August 16–20, Amsterdam 2008).
Elias, J.E. & Gygi, S.P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: a practical and powerful approach to multiple testing. J. R. Stat. Soc., B 57, 289–300 (1995).
Acknowledgements
We thank J. Bartlett for technical assistance, M. Vigneron (Institut de Génétique et de Biologie Moléculaire et Cellulaire, Strasbourg) for the 7C2 antibody, B. Thomas, D. Trudgian, G. Ridlova and M. Dreger for help with proteomics, M. Shaw for help with electron microscopy, and the Medical Research Council (S.M. and B.D.), EP Abraham Research Fund (B.D.), Biotechnology and Biological Sciences Research Council (A.P.), Wellcome Trust (A.P.) and Felix Scholarship Trust of Oxford University (S.B.) for support.
Author information
Authors and Affiliations
Contributions
Experiments were designed by S.M., B.D., A.P., S.B. and P.R.C. S.M. developed the isolation procedure and carried out many of the validation experiments, S.M. and B.D. performed gel electrophoreses and mass spectrometry, A.P. developed native 3C and carried out RT-PCR, S.B. did the light microscopy, and I.M.C. developed software. All authors wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–6, Supplementary Tables 1–3, Supplementary Note (PDF 1947 kb)
Supplementary Table 4
Experiment 1: output from the CPFP (FDRs for complexes I, II and III were 0.84, 0.8 and 0.82% respectively), and results from the SI analysis. (XLSX 1665 kb)
Supplementary Table 5
Experiment 2: output from the CPFP (FDRs for complexes I, II and III were all 0.8%). (XLSX 103 kb)
Supplementary Table 6
Experiment 3: output from the CPFP (FDRs for complexes II and III were 0.75 and 0.65%, respectively), and results from the SI analysis. (XLSX 243 kb)
Supplementary Table 7
Comparison of the proteomes seen in all three experiments. (XLSX 1036 kb)
Rights and permissions
About this article
Cite this article
Melnik, S., Deng, B., Papantonis, A. et al. The proteomes of transcription factories containing RNA polymerases I, II or III. Nat Methods 8, 963–968 (2011). https://doi.org/10.1038/nmeth.1705
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1705
This article is cited by
-
Alternative polyadenylation by sequential activation of distal and proximal PolyA sites
Nature Structural & Molecular Biology (2022)
-
CGGBP1 regulates CTCF occupancy at repeats
Epigenetics & Chromatin (2019)
-
Topological organization and dynamic regulation of human tRNA genes during macrophage differentiation
Genome Biology (2017)
-
Closing the loop: 3C versus DNA FISH
Genome Biology (2016)
-
Nuclear organization and 3D chromatin architecture in cognition and neuropsychiatric disorders
Molecular Brain (2016)