Abstract
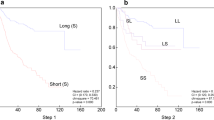
Diffuse large B-cell lymphoma (DLBCL), the most common lymphoid malignancy in adults, is curable in less than 50% of patients. Prognostic models based on pre-treatment characteristics, such as the International Prognostic Index (IPI), are currently used to predict outcome in DLBCL. However, clinical outcome models identify neither the molecular basis of clinical heterogeneity, nor specific therapeutic targets. We analyzed the expression of 6,817 genes in diagnostic tumor specimens from DLBCL patients who received cyclophosphamide, adriamycin, vincristine and prednisone (CHOP)-based chemotherapy, and applied a supervised learning prediction method to identify cured versus fatal or refractory disease. The algorithm classified two categories of patients with very different five-year overall survival rates (70% versus 12%). The model also effectively delineated patients within specific IPI risk categories who were likely to be cured or to die of their disease. Genes implicated in DLBCL outcome included some that regulate responses to B-cell–receptor signaling, critical serine/threonine phosphorylation pathways and apoptosis. Our data indicate that supervised learning classification techniques can predict outcome in DLBCL and identify rational targets for intervention.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
A clinical evaluation of the International Lymphoma Study Group classification of non-Hodgkin's lymphoma. The Non-Hodgkin's Lymphoma Classification Project. Blood 89, 3909–3918 (1997).
Shipp, M., Harris, N. & Mauch, P. The non-Hodgkin's lymphomas. in Cancer Principles & Practices of Oncology (eds. DeVita, V.T., Hellman, S. & Rosenberg, S.A) 2165–2220 (Lippincott, Philadelphia, 1997).
A predictive model for aggressive non-Hodgkin's lymphoma. The International NHL Prognostic Factors Project. N. Engl. J. Med. 329, 987–994 (1993).
Shipp, M.A. et al. International consensus conference on high-dose therapy with hematopoietic stem cell transplantation in aggressive non-Hodgkin's lymphomas: Report of the jury. J. Clin. Oncol. 17, 423–429 (1999).
Yakushijin, Y. et al. A directly spliced exon 10-containing CD44 variant promotes the metastasis and homotypic aggregation of aggressive non-Hodgkin's lymphoma. Blood 91, 4282–4291 (1998).
Terol, M.-J. et al. Expression of the adhesion molecule ICAM-1 in non-Hodgkin's lymphoma: Relationship with tumor dissemination and prognostic importance. J. Clin. Oncol. 16, 35–40 (1998).
Gascoyne, R. et al. Prognostic significance of bcl-2 protein expression and bcl-2 gene rearrangement in diffuse aggressive non-Hodgkin's lymphoma. Blood 90, 244–251 (1997).
Kramer, M. et al. Clinical relevance of BCL2, BCL6, and MYC rearrangements in diffuse large B-cell lymphoma. Blood 92, 3152–3162 (1998).
Ambrosini, G., Adida, C. & Altieri, D. A novel anti-apoptosis gene, survivin, expressed in cancer and lymphoma. Nature Med. 3, 917–921 (1997).
Salven, P., Teerenhovi, L. & Joensuu, H. A high pretreatment serum vascular endothelial growth factor concentration is associated with poor outcome in non-Hodgkin's lymphoma. Blood 90, 3167–3172 (1997).
Aguiar, R. et al. BAL is a novel risk-related gene in diffuse large B-cell lymphomas which enhances cellular migration. Blood 96, 4328–4334 (2000).
Alizadeh, A. et al. Distinct types of diffuse large B-cell lymphoma identified by gene expression profiling. Nature 4051, 503–511 (2000).
Kuppers, R., Klein, U., Hansmann, M.-L. & Rajewsky, K. Cellular origin or human B-cell lymphomas. N. Engl. J. Med. 341, 1520–1529 (1999).
Golub, T. et al. Molecular classification of cancer: Class discovery and class prediction by gene expression monitoring. Science 286, 531–537 (1999).
Cleary, M. et al. Clustering of extensive somatic mutations in the variable region of an immunoglobulin heavy chain gene from a human B cell lymphoma. Cell 44, 97–106 (1986).
Wood, L. et al. HMG-I/Y, a new c-Myc target gene and potential oncogene. Mol. Cell Biol. 20, 5490–5502 (2000).
Ilangumaran, S., Borisch, B. & Hoessli, D. Signal transduction via CD44: role of plasma membrane microdomains. Leuk. Lymphoma 35, 455–469 (1999).
Matarrese, P. et al. Galectin-3 overexpression protects from apoptosis by improving cell adhesion properties. Internal J. Cancer 85, 545–554 (2000).
Zhang, H. et al. Structural basis of BFL-1 for its interaction with BAX and its anti-apoptotic action in mammalian and yeast cells. J. Biol. Chem. 275, 11092–11099 (2000).
Lee, H., Dadgostar, H., Cheng, Q., Shu, J. & Cheng, G. NF-κB-mediated up-regulation of Bcl-x and Bfl-1/A1 is required for CD40 survival signaling in B lymphocytes. Proc. Natl. Acad. Sci. USA 96, 9136–9141 (1999).
Wang, J., Saunthararajah, Y., Redner, R.L. & Liu, J.M. Inhibitors of histone deacetylase relieve ETO-mediated repression and induce differentiation of AML1-ETO leukemia cells. Cancer Res. 59, 2766–2769 (1999).
Zong, W., Edelstein, L., Chen, C., Bash, J. & Gelinas, C. The prosurvival Bcl-2 homolog Bfl-1/A1 is a direct transcriptional target of NF-κB that blocks TNFα-induced apoptosis. Genes Devel. 13, 382–387 (1999).
Finger, L. et al. The human PD-1 gene: complete cDNA, genomic organization, and developmentally regulated expression in B cell progenitors. Gene 197, 177–187 (1997).
Kitson, J. et al. A death-domain-containing receptor that mediates apoptosis. Nature 384, 372–375 (1996).
Wellmann, A. et al. Detection of differentially expressed genes in lymphomas using cDNA arrays: identification of clusterin as a new diagnostic marker for anaplastic large-cell lymphomas. Blood 96, 398–404 (2000).
Sommers, C. et al. A role for the Tec family tyrosine kinase Txk in T cell activation and thymocyte selection. J. Exp. Med. 190, 1427–1438 (1999).
Testi, R., D'Ambrosio, D., De Maria, R. & Santoni, A. The CD69 receptor: a multipurpose cell-surface trigger for hematopoietic cells. Immunol. Today 15, 479–483 (1994).
Ruegg, C. et al. V7, a novel leukocyte surface protein that participates in T cell activation. II. Molecular cloning and characterization of the V7 gene. J. Immunol. 154, 4434–4443 (1995).
Saeki, H., Moore, A., Brown, M. & Hwang, S. Cutting edge: secondary lymphoid-tissue chemokine (SLC) and CC chemokine receptor 7 (CCR7) participate in the emigration pathway of mature dendritic cells from the skin to regional lymph nodes. J. Immunol. 162, 2472–2475 (1999).
Moller, M. et al. Frequent disruption of the RB1 pathway in diffuse large B cell lymphoma: prognostic significance of E2F-1 and p16NK4A. Leukemia 14, 898–904 (2000).
Brenner, C. & Kroemer, G. Mitochondria—the death signal integrators. Science 289, 1150–1151 (2000).
Youn, H., Sun, L., Prywes, R. & Liu, J. Apoptosis of T cells mediated by Ca2+-induced release of the transcription factor MEF2. Science 286, 790–793 (1999).
Kuang, A., Cado, D. & Winoto, A. Nur77 transcription activity correlates with its apoptotic function in vivo. Eur. J. Immunol. 29, 3722–3728 (1999).
Xue, Y. et al. Positive and negative thymic selection in T cell receptor-transgenic mice correlate with Nur77 mRNA expression. Eur. J. Immunol. 27, 2048–2056 (1997).
Li, H. et al. Cytochrome c release and apoptosis induced by mitochondrial targeting of nuclear orphan receptor TR3. Science 289, 1159–1164 (2000).
Manning, C. et al. Suppression of human inflammatory cell function by subtype-selective PDE4 inhibitors correlates with inhibition of PDE4A and PDE4B. Br. J. Pharmacol. 128, 1393–1398 (1999).
Lerner, A., Kim, D. & Lee, R. The cAMP signaling pathway as a therapeutic target in lymphoid malignancies. Leuk. Lymphoma 37, 39–51 (2000).
Kim, D. & Lerner, A. Type 4 cyclic adenosine monophosphate phosphodiesterase as a therapeutic target in chronic lymphocytic leukemia. Blood 92, 2484–2494 (1998).
Mischak, H. et al. Expression of protein kinase C genes in hemopoietic cells is cell-type- and B cell-differentiation stage specific. J. Immunol. 147, 3981–3987 (1991).
Leitges, M. et al. Immunodeficiency in protein kinase Cb-deficient mice. Science 273, 788–791 (1996).
King, L., Norvell, A. & Monroe, J. Antigen receptor-induced signal transduction imbalances associated with the negative selection of immature B cells. J. Immunol. 162, 2655–2662 (1999).
Teicher, B. et al. Enzymatic rationale and preclinical support for a potent protein kinase C beta inhibitor in cancer therapy. Adv. Enzyme Reg. 39, 313–327 (1999).
Tamayo, P. et al. Interpreting patterns of gene expression with self-organizing maps: Methods and application to hematopoietic differentiation. Proc. Natl. Acad. Sci. USA 96, 2907–2912 (1999).
Huberty, C.J. Applied Discriminant Analysis. (John Wiley and Sons, Inc., 1994).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
We thank members of the Center for Genome Research and members of the Shipp and Aster laboratories for technical assistance and helpful comments. This work was supported in part by grants from Bristol–Myers Squibb, Millennium Pharmaceuticals and Affymetrix (E.S.L.).
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Shipp, M., Ross, K., Tamayo, P. et al. Diffuse large B-cell lymphoma outcome prediction by gene-expression profiling and supervised machine learning. Nat Med 8, 68–74 (2002). https://doi.org/10.1038/nm0102-68
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nm0102-68
This article is cited by
-
Unleashing the power of machine learning in cancer analysis: a novel gene selection and classifier ensemble strategy
Research on Biomedical Engineering (2024)
-
Self-regularized Lasso for selection of most informative features in microarray cancer classification
Multimedia Tools and Applications (2024)
-
Learning from high dimensional data based on weighted feature importance in decision tree ensembles
Computational Statistics (2024)
-
MOBILE pipeline enables identification of context-specific networks and regulatory mechanisms
Nature Communications (2023)
-
Iterated cross validation method for prediction of survival in diffuse large B-cell lymphoma for small size dataset
Scientific Reports (2023)