Abstract
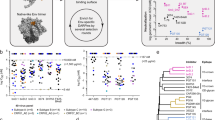
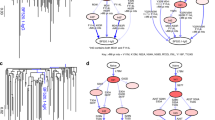
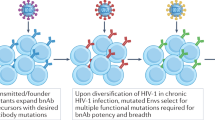
The elicitation of broadly neutralizing antibodies (bNAbs) is likely to be essential for a preventative HIV-1 vaccine, but this has not yet been achieved by immunization. In contrast, some HIV-1-infected individuals naturally mount bNAb responses during chronic infection, suggesting that years of maturation may be required for neutralization breadth1,2,3,4,5,6. Recent studies have shown that viral diversification precedes the emergence of bNAbs, but the significance of this observation is unknown7,8. Here we delineate the key viral events that drove neutralization breadth within the CAP256-VRC26 family of 33 monoclonal antibodies (mAbs) isolated from a superinfected individual. First, we identified minority viral variants, termed bNAb-initiating envelopes, that were distinct from both of the transmitted/founder (T/F) viruses and that efficiently engaged the bNAb precursor. Second, deep sequencing revealed a pool of diverse epitope variants (immunotypes) that were preferentially neutralized by broader members of the antibody lineage. In contrast, a 'dead-end' antibody sublineage unable to neutralize these immunotypes showed limited evolution and failed to develop breadth. Thus, early viral escape at key antibody-virus contact sites selects for antibody sublineages that can tolerate these changes, thereby providing a mechanism for the generation of neutralization breadth within a developing antibody lineage.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Primary accessions
BioProject
NCBI Reference Sequence
Sequence Read Archive
Referenced accessions
Protein Data Bank
References
Gray, E.S. et al. HIV-1 neutralization breadth develops incrementally over 4 years and is associated with CD4+ T cell decline and high viral load during acute infection. J. Virol. 85, 4828–4840 (2011).
Hraber, P. et al. Prevalence of broadly neutralizing antibody responses during chronic HIV-1 infection. AIDS 28, 163–169 (2014).
Piantadosi, A. et al. Breadth of neutralizing antibody response to human immunodeficiency virus type 1 is affected by factors early in infection but does not influence disease progression. J. Virol. 83, 10269–10274 (2009).
Sather, D.N. et al. Factors associated with the development of cross-reactive neutralizing antibodies during human immunodeficiency virus type 1 infection. J. Virol. 83, 757–769 (2009).
Euler, Z. et al. Longitudinal analysis of early HIV-1-specific neutralizing activity in an elite neutralizer and in five patients who developed cross-reactive neutralizing activity. J. Virol. 86, 2045–2055 (2012).
Walker, L.M. et al. A limited number of antibody specificities mediate broad and potent serum neutralization in selected HIV-1 infected individuals. PLoS Pathog. 6, e1001028 (2010).
Doria-Rose, N.A. et al. Developmental pathway for potent V1V2-directed HIV-neutralizing antibodies. Nature 509, 55–62 (2014).
Liao, H.X. et al. Co-evolution of a broadly neutralizing HIV-1 antibody and founder virus. Nature 496, 469–476 (2013).
Mascola, J.R. & Haynes, B.F. HIV-1 neutralizing antibodies: understanding nature's pathways. Immunol. Rev. 254, 225–244 (2013).
Gao, F. et al. Cooperation of B cell lineages in induction of HIV-1-broadly neutralizing antibodies. Cell 158, 481–491 (2014).
Wibmer, C.K. et al. Viral escape from HIV-1 neutralizing antibodies drives increased plasma neutralization breadth through sequential recognition of multiple epitopes and immunotypes. PLoS Pathog. 9, e1003738 (2013).
Moore, P.L. et al. Evolution of an HIV glycan-dependent broadly neutralizing antibody epitope through immune escape. Nat. Med. 18, 1688–1692 (2012).
Moore, P.L. et al. Multiple pathways of escape from HIV broadly cross-neutralizing V2-dependent antibodies. J. Virol. 87, 4882–4894 (2013).
Doria-Rose, N.A. et al. A new member of the V1V2-directed CAP256-VRC26 lineage that shows increased breadth and exceptional potency. J. Virol. (in the press).
Klein, F. et al. Antibodies in HIV-1 vaccine development and therapy. Science 341, 1199–1204 (2013).
Kwong, P.D., Mascola, J.R. & Nabel, G.J. Broadly neutralizing antibodies and the search for an HIV-1 vaccine: the end of the beginning. Nat. Rev. Immunol. 13, 693–701 (2013).
Sok, D. et al. Recombinant HIV envelope trimer selects for quaternary-dependent antibodies targeting the trimer apex. Proc. Natl. Acad. Sci. USA 111, 17624–17629 (2014).
Walker, L.M. et al. Broad neutralization coverage of HIV by multiple highly potent antibodies. Nature 477, 466–470 (2011).
Scheid, J.F. et al. Broad diversity of neutralizing antibodies isolated from memory B cells in HIV-infected individuals. Nature 458, 636–640 (2009).
Mouquet, H. et al. Memory B cell antibodies to HIV-1 gp140 cloned from individuals infected with clade A and B viruses. PLoS ONE 6, e24078 (2011).
Pappas, L. et al. Rapid development of broadly influenza neutralizing antibodies through redundant mutations. Nature 516, 418–422 (2014).
Walker, L.M. et al. Broad and potent neutralizing antibodies from an African donor reveal a new HIV-1 vaccine target. Science 326, 285–289 (2009).
Briney, B.S., Willis, J.R. & Crowe, J.E. Jr. Human peripheral blood antibodies with long HCDR3s are established primarily at original recombination using a limited subset of germline genes. PLoS ONE 7, e36750 (2012).
Wardemann, H. et al. Predominant autoantibody production by early human B cell precursors. Science 301, 1374–1377 (2003).
Jardine, J. et al. Rational HIV immunogen design to target specific germline B cell receptors. Science 340, 711–716 (2013).
McGuire, A.T. et al. Engineering HIV envelope protein to activate germline B cell receptors of broadly neutralizing anti-CD4 binding site antibodies. J. Exp. Med. 210, 655–663 (2013).
van Loggerenberg, F. et al. Establishing a cohort at high risk of HIV infection in South Africa: challenges and experiences of the CAPRISA 002 acute infection study. PLoS ONE 3, e1954 (2008).
Jabara, C.B., Jones, C.D., Roach, J., Anderson, J.A. & Swanstrom, R. Accurate sampling and deep sequencing of the HIV-1 protease gene using a Primer ID. Proc. Natl. Acad. Sci. USA 108, 20166–20171 (2011).
Goecks, J., Nekrutenko, A., Taylor, J. & Galaxy, T. Galaxy: a comprehensive approach for supporting accessible, reproducible, and transparent computational research in the life sciences. Genome Biol. 11, R86 (2010).
Blankenberg, D. et al. Galaxy: a web-based genome analysis tool for experimentalists. Curr. Protoc. Mol. Biol. 89, 19.10.1–19.10.21 (2010).
Giardine, B. et al. Galaxy: a platform for interactive large-scale genome analysis. Genome Res. 15, 1451–1455 (2005).
Blankenberg, D. et al. Manipulation of FASTQ data with Galaxy. Bioinformatics 26, 1783–1785 (2010).
Katoh, K. & Standley, D.M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013).
Edgar, R.C. MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 32, 1792–1797 (2004).
Wright, I.A. & Travers, S.A. RAMICS: trainable, high-speed and biologically relevant alignment of high-throughput sequencing reads to coding DNA. Nucleic Acids Res. 42, e106 (2014).
Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 9, 90–95 (2007).
Salazar-Gonzalez, J.F. et al. Deciphering human immunodeficiency virus type 1 transmission and early envelope diversification by single-genome amplification and sequencing. J. Virol. 82, 3952–3970 (2008).
Gao, F. et al. Molecular cloning and analysis of functional envelope genes from human immunodeficiency virus type 1 sequence subtypes A through G. The WHO and NIAID Networks for HIV Isolation and Characterization. J. Virol. 70, 1651–1667 (1996).
Kraus, M.H. et al. A rev1-vpu polymorphism unique to HIV-1 subtype A and C strains impairs envelope glycoprotein expression from rev-vpu-env cassettes and reduces virion infectivity in pseudotyping assays. Virology 397, 346–357 (2010).
Wei, X. et al. Antibody neutralization and escape by HIV-1. Nature 422, 307–312 (2003).
Hall, T.A. BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucl. Acids. Symp. Ser. 41, 95–98 (1999).
Acknowledgements
We thank the participants in the CAPRISA 002 cohort for their commitment to attending the CAPRISA clinics in South Africa. Thanks to the CAPRISA 002 study team (including N. Naicker, V. Asari, N. Majola, T. Cekwane, N. Samsunder, Z. Mchunu, D. Nkosi, H. Shozi, S. Ndlovu, P. Radebe, K. Leask and M. Upfold) for managing the cohort and providing specimens. We are grateful to C. K. Wibmer for useful discussions. We thank P. Labuschagne, D. Matten and T. York for assistance with viral next-generation sequence analysis, R. Bailer and M. Louder for the CAP256-VRC26 UCA neutralization data against the large virus panel and J. Mulllikin and the US National Institutes of Health (NIH) Intramural Sequencing Center (NISC) Comparative Sequencing Program for antibody NGS data at 34 weeks. We are grateful to J. Baalwa, D. Ellenberger, F. Gao, B. Hahn, K. Hong, J. Kim, F. McCutchan, D. Montefiori, J. Overbaugh, E. Sanders-Buell, G. Shaw, R. Swanstrom, M. Thomson, S. Tovanabutra and L. Zhang for contributing the HIV-1 envelope plasmids used in our neutralization panel. The JC53bl-13 (TZM-bl) and 293T cell lines were obtained through the NIH AIDS Reagent Program, Division of AIDS, NIAID, NIH from J.C. Kappes, X. Wu and Tranzyme, Inc., and A. Rice, respectively. We acknowledge research funding from the Centre for the AIDS Programme of Research (CAPRISA) (S.S.A.K.), the South African Medical Research Council (MRC) SHIP program (P.L.M.), the NIH through a U01 grant (AI116086-01) (P.L.M.), the intramural research programs of the Vaccine Research Center and the National Institute of Allergy and Infectious Diseases (NIAID) (J.R.M.). CAPRISA is funded by the South African HIV/AIDS Research and Innovation Platform of the South African Department of Science and Technology, and was initially supported by the US NIAID, NIH, US Department of Health and Human Services grant U19 AI51794 (S.S.A.K.). Funding was received by J.N.B. and C.A. from the Columbia University-Southern African Fogarty AIDS International Training and Research Program through the Fogarty International Center, NIH grant 5 D43 TW000231 (to Quarraisha Abdool Karim). Additional fellowships included a University of the Witwatersrand Postgraduate Merit Award (J.N.B.), a Poliomyelitis Research Foundation PhD Bursary (J.N.B.), a National Research Foundation of South Africa Innovation PhD Bursary (J.N.B.), a National Research Foundation of South Africa Postdoctoral Fellowship (C.A.), and a Wellcome Trust Intermediate Fellowship in Public Health and Tropical Medicine, grant 089933/Z/09/Z (P.L.M.).
Author information
Authors and Affiliations
Contributions
J.N.B., L.M. and P.L.M. designed the study, analyzed data and wrote the manuscript. C.A. and C.W. designed and performed viral NGS and analysis and edited the manuscript. N.A.D.-R. led the isolation of CAP256 mAbs. J.N.B. and O.K. performed neutralization experiments. C.A.S. and L.S. generated and analyzed antibody NGS data. J.N.B. and T.K. expressed and purified monoclonal antibodies. J.N.B. and D.K. generated single-genome sequences and viral envelope clones. G.B. developed computational tools to analyze NGS data, and assisted in the NGS analysis. J.G. assessed CAP256-VRC26 UCA neutralization of heterologous viruses. N.J.G. and the CAPRISA Study Team managed the CAPRISA cohort and contributed samples and data for CAP256. S.S.A.K., C.W. and L.M. established and led the CAPRISA cohort. C.W., N.A.D.-R., L.S., P.D.K., and J.R.M. contributed to data analysis and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Table 1 (PDF 3563 kb)
Source data
Rights and permissions
About this article
Cite this article
Bhiman, J., Anthony, C., Doria-Rose, N. et al. Viral variants that initiate and drive maturation of V1V2-directed HIV-1 broadly neutralizing antibodies. Nat Med 21, 1332–1336 (2015). https://doi.org/10.1038/nm.3963
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3963
This article is cited by
-
Engineering of HIV-1 neutralizing antibody CAP256V2LS for manufacturability and improved half life
Scientific Reports (2022)
-
Cross-reactive serum and memory B-cell responses to spike protein in SARS-CoV-2 and endemic coronavirus infection
Nature Communications (2021)
-
Broadly neutralizing plasma antibodies effective against autologous circulating viruses in infants with multivariant HIV-1 infection
Nature Communications (2020)
-
Serum alpha-mannosidase as an additional barrier to eliciting oligomannose-specific HIV-1-neutralizing antibodies
Scientific Reports (2020)
-
Induction of neutralizing antibodies against tier 2 human immunodeficiency virus 1 in rhesus macaques infected with tier 1B simian/human immunodeficiency virus
Archives of Virology (2019)