Abstract
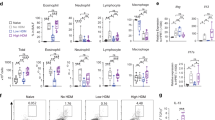
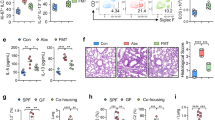
Epidemiological data point toward a critical period in early life during which environmental cues can set an individual on a trajectory toward respiratory health or disease1,2,3,4,5,6,7,8. The neonatal immune system matures during this period9, although little is known about the signals that lead to its maturation. Here we report that the formation of the lung microbiota is a key parameter in this process. Immediately following birth, neonatal mice were prone to develop exaggerated airway eosinophilia, release type 2 helper T cell cytokines and exhibit airway hyper-responsiveness following exposure to house dust mite allergens, even though their lungs harbored high numbers of natural CD4+Foxp3+CD25+Helios+ regulatory T (Treg) cells. During the first 2 weeks after birth, the bacterial load in the lungs increased, and representation of the bacterial phyla shifts from a predominance of Gammaproteobacteria and Firmicutes towards Bacteroidetes. The changes in the microbiota were associated with decreased aeroallergen responsiveness and the emergence of a Helios− Treg cell subset that required interaction with programmed death ligand 1 (PD-L1) for development. Absence of microbial colonization10 or blockade of PD-L1 during the first 2 weeks postpartum maintained exaggerated responsiveness to allergens through to adulthood. Adoptive transfer of Treg cells from adult mice to neonates before aeroallergen exposure ameliorated disease. Thus, formation of the airway microbiota induces regulatory cells early in life, which, when dysregulated, can lead to sustained susceptibility to allergic airway inflammation in adulthood.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
von Mutius, E. & Radon, K. Living on a farm: impact on asthma induction and clinical course. Immunol. Allergy Clin. North Am. 28, 631–647, ix–x (2008).
von Mutius, E. & Vercelli, D. Farm living: effects on childhood asthma and allergy. Nat. Rev. Immunol. 10, 861–868 (2010).
Burke, H. et al. Prenatal and passive smoke exposure and incidence of asthma and wheeze: systematic review and meta-analysis. Pediatrics 129, 735–744 (2012).
Lee, S.L. et al. Foetal exposure to maternal passive smoking is associated with childhood asthma, allergic rhinitis, and eczema. ScientificWorldJournal 2012, 542983 (2012).
Stensballe, L.G., Simonsen, J., Jensen, S.M., Bonnelykke, K. & Bisgaard, H. Use of antibiotics during pregnancy increases the risk of asthma in early childhood. J. Pediatr. 162, 832–838.e3 (2013).
Gern, J.E., Rosenthal, L.A., Sorkness, R.L. & Lemanske, R.F. Jr. Effects of viral respiratory infections on lung development and childhood asthma. J. Allergy Clin. Immunol. 115, 668–674, quiz 675 (2005).
Jackson, D.J. The role of rhinovirus infections in the development of early childhood asthma. Curr. Opin. Allergy Clin. Immunol. 10, 133–138 (2010).
Wu, P. & Hartert, T.V. Evidence for a causal relationship between respiratory syncytial virus infection and asthma. Expert Rev. Anti Infect. Ther. 9, 731–745 (2011).
Holt, P.G. & Jones, C.A. The development of the immune system during pregnancy and early life. Allergy 55, 688–697 (2000).
Herbst, T. et al. Dysregulation of allergic airway inflammation in the absence of microbial colonization. Am. J. Respir. Crit. Care Med. 184, 198–205 (2011).
Hooper, L.V., Littman, D.R. & Macpherson, A.J. Interactions between the microbiota and the immune system. Science 336, 1268–1273 (2012).
Macpherson, A.J. & Harris, N.L. Interactions between commensal intestinal bacteria and the immune system. Nat. Rev. Immunol. 4, 478–485 (2004).
Hilty, M. et al. Disordered microbial communities in asthmatic airways. PLoS ONE 5, e8578 (2010).
Huang, Y.J. et al. Airway microbiota and bronchial hyperresponsiveness in patients with suboptimally controlled asthma. J. Allergy Clin Immunol. 127, 372–381.e1–e3 (2011).
Pragman, A.A., Kim, H.B., Reilly, C.S., Wendt, C. & Isaacson, R.E. The lung microbiome in moderate and severe chronic obstructive pulmonary disease. PLoS ONE 7, e47305 (2012).
Erb-Downward, J.R. et al. Analysis of the lung microbiome in the “healthy” smoker and in COPD. PLoS ONE 6, e16384 (2011).
Marsland, B.J., Yadava, K. & Nicod, L.P. The airway microbiome and disease. Chest 144, 632–637 (2013).
Twigg, H.L. III et al. Use of bronchoalveolar lavage to assess the respiratory microbiome: signal in the noise. Lancet Respir. Med. 1, 354–356 (2013).
Trompette, A. et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat. Med. 20, 159–166 (2014).
Francisco, L.M. et al. PD-L1 regulates the development, maintenance, and function of induced regulatory T cells. J. Exp. Med. 206, 3015–3029 (2009).
Wang, L. et al. Programmed death 1 ligand signaling regulates the generation of adaptive Foxp3+CD4+ regulatory T cells. Proc. Natl. Acad. Sci. USA 105, 9331–9336 (2008).
Geuking, M.B. et al. Intestinal bacterial colonization induces mutualistic regulatory T cell responses. Immunity 34, 794–806 (2011).
Round, J.L. & Mazmanian, S.K. Inducible Foxp3+ regulatory T-cell development by a commensal bacterium of the intestinal microbiota. Proc. Natl. Acad. Sci. USA 107, 12204–12209 (2010).
Curotto de Lafaille, M.A. et al. Adaptive Foxp3+ regulatory T cell-dependent and -independent control of allergic inflammation. Immunity 29, 114–126 (2008).
Curotto de Lafaille, M.A. & Lafaille, J.J. Natural and adaptive foxp3+ regulatory T cells: more of the same or a division of labor? Immunity 30, 626–635 (2009).
Rangel-Moreno, J. et al. The development of inducible bronchus-associated lymphoid tissue depends on IL-17. Nat. Immunol. 12, 639–646 (2011).
Olszak, T. et al. Microbial exposure during early life has persistent effects on natural killer T cell function. Science 336, 489–493 (2012).
Cahenzli, J., Koller, Y., Wyss, M., Geuking, M.B. & McCoy, K.D. Intestinal microbial diversity during early-life colonization shapes long-term IgE levels. Cell Host Microbe 14, 559–570 (2013).
Nembrini, C. et al. Bacterial-induced protection against allergic inflammation through a multicomponent immunoregulatory mechanism. Thorax 66, 755–763 (2011).
Hagner, S. et al. Farm-derived gram-positive bacterium Staphylococcus sciuri W620 prevents asthma phenotype in HDM- and OVA-exposed mice. Allergy 68, 322–329 (2013).
Bacchetti De Gregoris, T., Aldred, N., Clare, A.S. & Burgess, J.G. Improvement of phylum- and class-specific primers for real-time PCR quantification of bacterial taxa. J. Microbiol. Methods 86, 351–356 (2011).
Acknowledgements
This work is supported by the Leenaards Foundation in Lausanne and the Swiss National Science Foundation grant 310030_146983, awarded to B.J.M. B.J.M. is a Cloetta Medical Research Fellow and, by holding this title, receives financial support. B.J.M. and C.M.L. are part of the European Cooperation in Science and Technology Action BM1201, Developmental Origins of Chronic Lung Disease. S.S. and C.M.L. are supported by the Wellcome Trust grants 087618/Z/08/Z and 083586/Z/07/Z. C.M.L. is a Wellcome Senior Fellow in Basic Biomedical Science and, by holding this title, receives financial support. We thank D. Pinschewer and S. Kallert for valuable discussions and critical feedback.
Author information
Authors and Affiliations
Contributions
B.J.M. conceived the study. B.J.M. and E.S.G. designed the study. E.S.G., A.T. and K.Y. performed experiments. S.S. and R.S. performed neonatal lung function experiments. K.D.M. provided germ-free mice. E.S.G., A.T., K.Y., S.S., C.M.L., L.P.N. and B.J.M. provided critical analysis and discussion. E.S.G. and B.J.M. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 (PDF 652 kb)
Rights and permissions
About this article
Cite this article
Gollwitzer, E., Saglani, S., Trompette, A. et al. Lung microbiota promotes tolerance to allergens in neonates via PD-L1. Nat Med 20, 642–647 (2014). https://doi.org/10.1038/nm.3568
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.3568
This article is cited by
-
Pathophysiology of acute lung injury in patients with acute brain injury: the triple-hit hypothesis
Critical Care (2024)
-
Lung microbiome: new insights into the pathogenesis of respiratory diseases
Signal Transduction and Targeted Therapy (2024)
-
Impact of perinatal administration of probiotics on immune cell composition in neonatal mice
Pediatric Research (2024)
-
Allergie und Mikrobiom: Die Mikrobiota-Hypothese
Deutsche Zeitschrift für Akupunktur (2024)
-
Symbiotic microbiome Staphylococcus epidermidis restricts IL-33 production in allergic nasal epithelium via limiting the cellular necroptosis
BMC Microbiology (2023)