Abstract
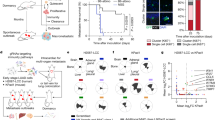
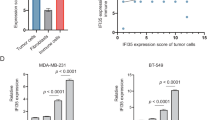
Breast cancer metastasis is a key determinant of long-term patient survival. By comparing the transcriptomes of primary and metastatic tumor cells in a mouse model of spontaneous bone metastasis, we found that a substantial number of genes suppressed in bone metastases are targets of the interferon regulatory factor Irf7. Restoration of Irf7 in tumor cells or administration of interferon led to reduced bone metastases and prolonged survival time. In mice deficient in the interferon (IFN) receptor or in natural killer (NK) and CD8+ T cell responses, metastasis was accelerated, indicating that Irf7-driven suppression of metastasis was reliant on IFN signaling to host immune cells. We confirmed the clinical relevance of these findings in over 800 patients in which high expression of Irf7-regulated genes in primary tumors was associated with prolonged bone metastasis–free survival. This gene signature may identify patients that could benefit from IFN-based therapies. Thus, we have identified an innate immune pathway intrinsic to breast cancer cells, the suppression of which restricts immunosurveillance to enable metastasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
American Cancer Society. Breast Cancer Facts & Figures 2011–2012. <http://www.cancer.org/Research/CancerFactsFigures/BreastCancerFactsFigures/breast-cancer-facts-and-figures-2011-2012> (2012).
Fehm, T. et al. Tumor cell dormancy: implications for the biology and treatment of breast cancer. APMIS 116, 742–753 (2008).
Eckhardt, B.L., Francis, P.A., Parker, B.S. & Anderson, R.L. Strategies for the discovery and development of therapies for metastatic breast cancer. Nat. Rev. Drug Discov. 11, 479–497 (2012).
Dunn, G.P., Bruce, A.T., Ikeda, H., Old, L.J. & Schreiber, R.D. Cancer immunoediting: from immunosurveillance to tumor escape. Nat. Immunol. 3, 991–998 (2002).
Dunn, G.P. et al. A critical function for type I interferons in cancer immunoediting. Nat. Immunol. 6, 722–729 (2005).
Koebel, C.M. et al. Adaptive immunity maintains occult cancer in an equilibrium state. Nature 450, 903–907 (2007).
Schreiber, R.D., Old, L.J. & Smyth, M.J. Cancer immunoediting: integrating immunity's roles in cancer suppression and promotion. Science 331, 1565–1570 (2011).
Vesely, M.D., Kershaw, M.H., Schreiber, R.D. & Smyth, M.J. Natural innate and adaptive immunity to cancer. Annu. Rev. Immunol. 29, 235–271 (2011).
Honda, K. et al. IRF-7 is the master regulator of type-I interferon–dependent immune responses. Nature 434, 772–777 (2005).
Wathelet, M.G. et al. Virus infection induces the assembly of coordinately activated transcription factors on the IFN-β enhancer in vivo. Mol. Cell 1, 507–518 (1998).
Darnell, J.E. Jr., Kerr, I.M. & Stark, G.R. Jak-STAT pathways and transcriptional activation in response to IFNs and other extracellular signaling proteins. Science 264, 1415–1421 (1994).
Samarajiwa, S.A., Forster, S., Auchettl, K. & Hertzog, P.J. INTERFEROME: the database of interferon regulated genes. Nucleic Acids Res. 37, D852–D857 (2009).
Eckhardt, B.L. et al. Genomic analysis of a spontaneous model of breast cancer metastasis to bone reveals a role for the extracellular matrix. Mol. Cancer Res. 3, 1–13 (2005).
Lelekakis, M. et al. A novel orthotopic model of breast cancer metastasis to bone. Clin. Exp. Metastasis 17, 163–170 (1999).
Parker, B.S. et al. Primary tumour expression of the cysteine cathepsin inhibitor Stefin A inhibits distant metastasis in breast cancer. J. Pathol. 214, 337–346 (2008).
Culhane, A.C. et al. GeneSigDB—a curated database of gene expression signatures. Nucleic Acids Res. 38, D716–D725 (2010).
Lu, R., Au, W.C., Yeow, W.S., Hageman, N. & Pitha, P.M. Regulation of the promoter activity of interferon regulatory factor-7 gene. Activation by interferon snd silencing by hypermethylation. J. Biol. Chem. 275, 31805–31812 (2000).
Sheehan, K.C. et al. Blocking monoclonal antibodies specific for mouse IFN-α/β receptor subunit 1 (IFNAR-1) from mice immunized by in vivo hydrodynamic transfection. J. Interferon Cytokine Res. 26, 804–819 (2006).
Hwang, S.Y. et al. A null mutation in the gene encoding a type I interferon receptor component eliminates antiproliferative and antiviral responses to interferons α and β and alters macrophage responses. Proc. Natl. Acad. Sci. USA 92, 11284–11288 (1995).
Yang, L. et al. Abrogation of TGFβ signaling in mammary carcinomas recruits Gr-1+CD11b+ myeloid cells that promote metastasis. Cancer Cell 13, 23–35 (2008).
DuPre', S.A. & Hunter, K.W. Jr. Mouse mammary carcinoma 4T1 induces a leukemoid reaction with splenomegaly: association with tumor-derived growth factors. Exp. Mol. Pathol. 82, 12–24 (2007).
Youn, J.I., Nagaraj, S., Collazo, M. & Gabrilovich, D.I. Subsets of myeloid-derived suppressor cells in tumor-bearing mice. J. Immunol. 181, 5791–5802 (2008).
Waight, J.D., Hu, Q., Miller, A., Liu, S. & Abrams, S.I. Tumor-derived G-CSF facilitates neoplastic growth through a granulocytic myeloid-derived suppressor cell–dependent mechanism. PLoS ONE 6, e27690 (2011).
Adeegbe, D. et al. In vivo induction of myeloid suppressor cells and CD4+Foxp3+ T regulatory cells prolongs skin allograft survival in mice. Cell Transplant. 20, 941–954 (2011).
Hervas-Stubbs, S. et al. Direct effects of type I interferons on cells of the immune system. Clin. Cancer Res. 17, 2619–2627 (2011).
Wu, J.M. et al. Heterogeneity of breast cancer metastases: comparison of therapeutic target expression and promoter methylation between primary tumors and their multifocal metastases. Clin. Cancer Res. 14, 1938–1946 (2008).
Minn, A.J. et al. Genes that mediate breast cancer metastasis to lung. Nature 436, 518–524 (2005).
Harrell, J.C. et al. Genomic analysis identifies unique signatures predictive of brain, lung, and liver relapse. Breast Cancer Res. Treat. 132, 523–535 (2012).
Hayakawa, Y. & Smyth, M.J. CD27 dissects mature NK cells into two subsets with distinct responsiveness and migratory capacity. J. Immunol. 176, 1517–1524 (2006).
Maher, S.G., Romero-Weaver, A.L., Scarzello, A.J. & Gamero, A.M. Interferon: cellular executioner or white knight? Curr. Med. Chem. 14, 1279–1289 (2007).
Noppert, S.J., Fitzgerald, K.A. & Hertzog, P.J. The role of type I interferons in TLR responses. Immunol. Cell Biol. 85, 446–457 (2007).
Swann, J.B. et al. Type I IFN contributes to NK cell homeostasis, activation, and antitumor function. J. Immunol. 178, 7540–7549 (2007).
Dunn, G.P., Koebel, C.M. & Schreiber, R.D. Interferons, immunity and cancer immunoediting. Nat. Rev. Immunol. 6, 836–848 (2006).
Chin, A.I. et al. Toll-like receptor 3–mediated suppression of TRAMP prostate cancer shows the critical role of type I interferons in tumor immune surveillance. Cancer Res. 70, 2595–2603 (2010).
Savitsky, D., Tamura, T., Yanai, H. & Taniguchi, T. Regulation of immunity and oncogenesis by the IRF transcription factor family. Cancer Immunol. Immunother. 59, 489–510 (2010).
Coppin, C., Le, L., Porzsolt, F. & Wilt, T. Targeted therapy for advanced renal cell carcinoma. Cochrane Database Syst. Rev. CD006017 (2008).
Garbe, C. et al. Evidence and interdisciplinary consensus-based German guidelines: surgical treatment and radiotherapy of melanoma. Melanoma Res. 18, 61–67 (2008).
Bi, X. et al. Loss of interferon regulatory factor 5 (IRF5) expression in human ductal carcinoma correlates with disease stage and contributes to metastasis. Breast Cancer Res. 13, R111 (2011).
Peranzoni, E. et al. Myeloid-derived suppressor cell heterogeneity and subset definition. Curr. Opin. Immunol. 22, 238–244 (2010).
Ribechini, E., Greifenberg, V., Sandwick, S. & Lutz, M.B. Subsets, expansion and activation of myeloid-derived suppressor cells. Med. Microbiol. Immunol. 199, 273–281 (2010).
Vuk-Pavlovié, S. et al. Immunosuppressive CD14+HLA-DRlow/− monocytes in prostate cancer. Prostate 70, 443–455 (2010).
Lin, E.Y., Nguyen, A.V., Russell, R.G. & Pollard, J.W. Colony-stimulating factor 1 promotes progression of mammary tumors to malignancy. J. Exp. Med. 193, 727–740 (2001).
Melani, C., Chiodoni, C., Forni, G. & Colombo, M.P. Myeloid cell expansion elicited by the progression of spontaneous mammary carcinomas in c-erbB-2 transgenic BALB/c mice suppresses immune reactivity. Blood 102, 2138–2145 (2003).
Bunt, S.K., Sinha, P., Clements, V.K., Leips, J. & Ostrand-Rosenberg, S. Inflammation induces myeloid-derived suppressor cells that facilitate tumor progression. J. Immunol. 176, 284–290 (2006).
Penn, I. Tumors of the immunocompromised patient. Annu. Rev. Med. 39, 63–73 (1988).
Oruc, M.T., Soran, A., Jain, A.K., Wilson, J.W. & Fung, J. De novo breast cancer in patients with liver transplantation: University of Pittsburgh's experience and review of the literature. Liver Transpl. 10, 1–6 (2004).
DeNardo, D.G. et al. CD4+ T cells regulate pulmonary metastasis of mammary carcinomas by enhancing protumor properties of macrophages. Cancer Cell 16, 91–102 (2009).
Chen, Q., Zhang, X.H. & Massague, J. Macrophage binding to receptor VCAM-1 transmits survival signals in breast cancer cells that invade the lungs. Cancer Cell 20, 538–549 (2011).
Filipazzi, P. et al. Identification of a new subset of myeloid suppressor cells in peripheral blood of melanoma patients with modulation by a granulocyte-macrophage colony-stimulation factor–based antitumor vaccine. J. Clin. Oncol. 25, 2546–2553 (2007).
Diaz-Montero, C.M. et al. Increased circulating myeloid-derived suppressor cells correlate with clinical cancer stage, metastatic tumor burden, and doxorubicin-cyclophosphamide chemotherapy. Cancer Immunol. Immunother. 58, 49–59 (2009).
van 't Veer, L.J. et al. Gene expression profiling predicts clinical outcome of breast cancer. Nature 415, 530–536 (2002).
Weigelt, B. et al. Molecular portraits and 70-gene prognosis signature are preserved throughout the metastatic process of breast cancer. Cancer Res. 65, 9155–9158 (2005).
Coleman, R.E. et al. Breast-cancer adjuvant therapy with zoledronic acid. N. Engl. J. Med. 365, 1396–1405 (2011).
Gnant, M. et al. Adjuvant endocrine therapy plus zoledronic acid in premenopausal women with early-stage breast cancer: 62-month follow-up from the ABCSG-12 randomised trial. Lancet Oncol. 12, 631–641 (2011).
Hu, Z. et al. The molecular portraits of breast tumors are conserved across microarray platforms. BMC Genomics 7, 96 (2006).
Repetto, L. et al. Tamoxifen and interferon-β for the treatment of metastatic breast cancer. Breast Cancer Res. Treat. 39, 235–238 (1996).
Hadden, J.W. The immunology and immunotherapy of breast cancer: an update. Int. J. Immunopharmacol. 21, 79–101 (1999).
Frith, M.C. et al. Detection of functional DNA motifs via statistical over-representation. Nucleic Acids Res. 32, 1372–1381 (2004).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Parker, B.S. et al. Alterations in vascular gene expression in invasive breast carcinoma. Cancer Res. 64, 7857–7866 (2004).
St Croix, B. et al. Genes expressed in human tumor endothelium. Science 289, 1197–1202 (2000).
Haibe-Kains, B. et al. A three-gene model to robustly identify breast cancer molecular subtypes. J. Natl. Cancer Inst. 104, 311–325 (2012).
Cimino, A. et al. Epithelial cell adhesion molecule (EpCAM) is overexpressed in breast cancer metastases. Breast Cancer Res. Treat. 123, 701–708 (2010).
Wu, L., Patten, N., Yamashiro, C.T. & Chui, B. Extraction and amplification of DNA from formalin-fixed, paraffin-embedded tissues. Appl. Immunohistochem. Mol. Morphol. 10, 269–274 (2002).
Mikeska, T., Felsberg, J., Hewitt, C.A. & Dobrovic, A. Analyzing DNA methylation using bisulfite pyrosequencing. Methods Mol. Biol. 791, 33–53 (2011).
Wojdacz, T.K. & Dobrovic, A. Methylation-sensitive high resolution melting (MS-HRM): a new approach for sensitive and high-throughput assessment of methylation. Nucleic Acids Res. 35, e41 (2007).
Acknowledgements
This work was supported by the Cancer Council Victoria (B.S.P.), the Australian National Health and Medical Research Council (NHMRC) (P.J.H.), the Association of International Cancer Research (AICR) (A.M.), the Operational Infrastructure Scheme of the Victorian State government Department of Business and Innovation, the Australian Research Council (ARC) Centre for Structural and Functional Microbial Genomics and fellowship support for B.S.P. (NHMRC), R.L.A. (Australian National Breast Cancer Foundation, NBCF), A.M. (NBCF), C.Y.S. (NBCF, Cure Cancer Australia). We thank P. Hill (St Vincent's Pathology, St Vincent's Hospital, Fitzroy, Australia) for providing archived human breast cancer tissues assistance in the pathological assessment of immunohistochemistry staining and B. Haibe-Kains for assistance with codes in R. We also thank F. Miller (Karmanos Cancer Institute, Detroit, MI) for providing 66cl4 and 4T1 cell lines.
Author information
Authors and Affiliations
Contributions
B.N.B. and C.Y.S. generated the majority of data in the manuscript. N.P.W. and A.M. contributed to mouse experiments, including intramammary fatpad injections and mouse tissue harvests. Y.C., D.A. and N.E.M. carried out the fluorescence-activated cell sorting (FACS) analysis on immune populations. T.M. did the methylation-sensitive high-resolution melting (MS-HRM), S.F. (PhD student and graduate of computer science and genetics graduate, Monash University) and S.A.S. (bioinformatician) carried out the bioinformatics analysis of the mouse microarray experiments and promoter analyses. S.L., a clinician scientist with expertise in bioinformatics and biostatistics, carried out the prognostic analyses. P.A. is a pathologist who provided and scored the matched primary tumor and metastases tissue arrays. N.A.d.W. produced, purified and tested IFN-α1 for in vivo experiments. J.G. performed some microarray experiments. P.J.H. and B.S.P. were responsible for project design, interpretation of data and drafting the manuscript, in collaboration with R.L.A. and M.J.S.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–13 and Supplementary Tables 1–4 (PDF 6245 kb)
Rights and permissions
About this article
Cite this article
Bidwell, B., Slaney, C., Withana, N. et al. Silencing of Irf7 pathways in breast cancer cells promotes bone metastasis through immune escape. Nat Med 18, 1224–1231 (2012). https://doi.org/10.1038/nm.2830
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2830
This article is cited by
-
Cell-intrinsic and microenvironmental determinants of metastatic colonization
Nature Cell Biology (2024)
-
The Role of Breast Cancer Cells in Bone Metastasis: Suitable Seeds for Nourishing Soil
Current Osteoporosis Reports (2024)
-
Tumor necroptosis-mediated shedding of cell surface proteins promotes metastasis of breast cancer by suppressing anti-tumor immunity
Breast Cancer Research (2023)
-
Predictive and prognostic biomarkers of bone metastasis in breast cancer: current status and future directions
Cell & Bioscience (2023)
-
Bone serves as a transfer station for secondary dissemination of breast cancer
Bone Research (2023)