Abstract
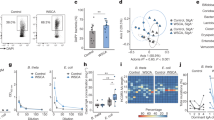
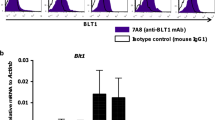
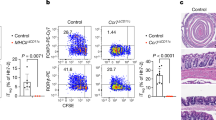
Commensal bacteria that colonize mammalian barrier surfaces are reported to influence T helper type 2 (TH2) cytokine-dependent inflammation and susceptibility to allergic disease, although the mechanisms that underlie these observations are poorly understood. In this report, we find that deliberate alteration of commensal bacterial populations via oral antibiotic treatment resulted in elevated serum IgE concentrations, increased steady-state circulating basophil populations and exaggerated basophil-mediated TH2 cell responses and allergic inflammation. Elevated serum IgE levels correlated with increased circulating basophil populations in mice and subjects with hyperimmunoglobulinemia E syndrome. Furthermore, B cell–intrinsic expression of myeloid differentiation factor 88 (MyD88) was required to limit serum IgE concentrations and circulating basophil populations in mice. Commensal-derived signals were found to influence basophil development by limiting proliferation of bone marrow–resident precursor populations. Collectively, these results identify a previously unrecognized pathway through which commensal-derived signals influence basophil hematopoiesis and susceptibility to TH2 cytokine–dependent inflammation and allergic disease.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Eder, W., Ege, M.J. & von Mutius, E. The asthma epidemic. N. Engl. J. Med. 355, 2226–2235 (2006).
Bahadori, K. et al. Economic burden of asthma: a systematic review. BMC Pulm. Med. 9, 24 (2009).
Mowen, K.A. & Glimcher, L.H. Signaling pathways in TH2 development. Immunol. Rev. 202, 203–222 (2004).
Holgate, S.T. Pathogenesis of asthma. Clin. Exp. Allergy 38, 872–897 (2008).
Vercelli, D. Discovering susceptibility genes for asthma and allergy. Nat. Rev. Immunol. 8, 169–182 (2008).
Zeiger, R.S. Food allergen avoidance in the prevention of food allergy in infants and children. Pediatrics 111, 1662–1671 (2003).
Gilliland, F.D. Outdoor air pollution, genetic susceptibility, and asthma management: opportunities for intervention to reduce the burden of asthma. Pediatrics 123 (suppl. 3), S168–S173 (2009).
Ege, M.J. et al. Exposure to environmental microorganisms and childhood asthma. N. Engl. J. Med. 364, 701–709 (2011).
Eckburg, P.B., Lepp, P.W. & Relman, D.A. Archaea and their potential role in human disease. Infect. Immun. 71, 591–596 (2003).
Whitman, W.B., Coleman, D.C. & Wiebe, W.J. Prokaryotes: the unseen majority. Proc. Natl. Acad. Sci. USA 95, 6578–6583 (1998).
Ley, R.E., Peterson, D.A. & Gordon, J.I. Ecological and evolutionary forces shaping microbial diversity in the human intestine. Cell 124, 837–848 (2006).
Kalliomäki, M. et al. Distinct patterns of neonatal gut microflora in infants in whom atopy was and was not developing. J. Allergy Clin. Immunol. 107, 129–134 (2001).
Kummeling, I. et al. Early life exposure to antibiotics and the subsequent development of eczema, wheeze, and allergic sensitization in the first 2 years of life: the KOALA Birth Cohort Study. Pediatrics 119, e225–e231 (2007).
Marra, F. et al. Antibiotic use in children is associated with increased risk of asthma. Pediatrics 123, 1003–1010 (2009).
Bashir, M.E., Louie, S., Shi, H.N. & Nagler-Anderson, C. Toll-like receptor 4 signaling by intestinal microbes influences susceptibility to food allergy. J. Immunol. 172, 6978–6987 (2004).
Noverr, M.C., Noggle, R.M., Toews, G.B. & Huffnagle, G.B. Role of antibiotics and fungal microbiota in driving pulmonary allergic responses. Infect. Immun. 72, 4996–5003 (2004).
Noverr, M.C., Falkowski, N.R., McDonald, R.A., McKenzie, A.N. & Huffnagle, G.B. Development of allergic airway disease in mice following antibiotic therapy and fungal microbiota increase: role of host genetics, antigen, and interleukin-13. Infect. Immun. 73, 30–38 (2005).
Herbst, T. et al. Dysregulation of allergic airway inflammation in the absence of microbial colonization. Am. J. Respir. Crit. Care Med. 184, 198–205 (2011).
Paul, W.E. & Zhu, J. How are TH2-type immune responses initiated and amplified? Nat. Rev. Immunol. 10, 225–235 (2010).
McCoy, K.D. et al. Natural IgE production in the absence of MHC class II cognate help. Immunity 24, 329–339 (2006).
Sudo, N. et al. The requirement of intestinal bacterial flora for the development of an IgE production system fully susceptible to oral tolerance induction. J. Immunol. 159, 1739–1745 (1997).
Kitaura, J. et al. Evidence that IgE molecules mediate a spectrum of effects on mast cell survival and activation via aggregation of the FcɛRI. Proc. Natl. Acad. Sci. USA 100, 12911–12916 (2003).
Hill, D.A. et al. Metagenomic analyses reveal antibiotic-induced temporal and spatial changes in intestinal microbiota with associated alterations in immune cell homeostasis. Mucosal Immunol. 3, 148–158 (2010).
Smith, K., McCoy, K.D. & Macpherson, A.J. Use of axenic animals in studying the adaptation of mammals to their commensal intestinal microbiota. Semin. Immunol. 19, 59–69 (2007).
Hammad, H. et al. Inflammatory dendritic cells—not basophils—are necessary and sufficient for induction of TH2 immunity to inhaled house dust mite allergen. J. Exp. Med. 207, 2097–2111 (2010).
Mohrs, K., Wakil, A.E., Killeen, N., Locksley, R.M. & Mohrs, M. A two-step process for cytokine production revealed by IL-4 dual-reporter mice. Immunity 23, 419–429 (2005).
Perrigoue, J.G. et al. MHC class II–dependent basophil-CD4+ T cell interactions promote TH2 cytokine-dependent immunity. Nat. Immunol. 10, 697–705 (2009).
Sokol, C.L., Barton, G.M., Farr, A.G. & Medzhitov, R. A mechanism for the initiation of allergen-induced T helper type 2 responses. Nat. Immunol. 9, 310–318 (2008).
Yoshimoto, T. et al. Basophils contribute to TH2-IgE responses in vivo via IL-4 production and presentation of peptide–MHC class II complexes to CD4+ T cells. Nat. Immunol. 10, 706–712 (2009).
Sokol, C.L. et al. Basophils function as antigen-presenting cells for an allergen-induced T helper type 2 response. Nat. Immunol. 10, 713–720 (2009).
Sullivan, B.M. et al. Genetic analysis of basophil function in vivo. Nat. Immunol. 12, 527–535 (2011).
Novey, H.S., Marchioli, L.E., Sokol, W.N. & Wells, I.D. Papain-induced asthma–physiological and immunological features. J. Allergy Clin. Immunol. 63, 98–103 (1979).
Phythian-Adams, A.T. et al. CD11c depletion severely disrupts Th2 induction and development in vivo. J. Exp. Med. 207, 2089–2096 (2010).
Siracusa, M.C. et al. TSLP promotes interleukin-3–independent basophil haematopoiesis and type 2 inflammation. Nature 477, 229–233 (2011).
Xiang, Z., Moller, C. & Nilsson, G. IgE-receptor activation induces survival and Bfl-1 expression in human mast cells but not basophils. Allergy 61, 1040–1046 (2006).
Mombaerts, P. et al. RAG-1–deficient mice have no mature B and T lymphocytes. Cell 68, 869–877 (1992).
Delphin, S. & Stavnezer, J. Characterization of an interleukin 4 (IL-4) responsive region in the immunoglobulin heavy chain germline epsilon promoter: regulation by NF-IL-4, a C/EBP family member and NF-κB/p50. J. Exp. Med. 181, 181–192 (1995).
Engelhardt, K.R. et al. Large deletions and point mutations involving the dedicator of cytokinesis 8 (DOCK8) in the autosomal-recessive form of hyper-IgE syndrome. J. Allergy Clin. Immunol. 124, 1289-302.e4 (2009).
Zhang, Q. et al. Combined immunodeficiency associated with DOCK8 mutations. N. Engl. J. Med. 361, 2046–2055 (2009).
Holgate, S. et al. The use of omalizumab in the treatment of severe allergic asthma: A clinical experience update. Respir. Med. 103, 1098–1113 (2009).
Pace, E. et al. Clinical benefits of 7 years of treatment with omalizumab in severe uncontrolled asthmatics. J. Asthma 48, 387–392 (2011).
Lin, H. et al. Omalizumab rapidly decreases nasal allergic response and FcepsilonRI on basophils. J. Allergy Clin. Immunol. 113, 297–302 (2004).
Shiratori, I. et al. Down-regulation of basophil function by human CD200 and human herpesvirus-8 CD200. J. Immunol. 175, 4441–4449 (2005).
Schnare, M. et al. Toll-like receptors control activation of adaptive immune responses. Nat. Immunol. 2, 947–950 (2001).
Clarke, T.B. et al. Recognition of peptidoglycan from the microbiota by Nod1 enhances systemic innate immunity. Nat. Med. 16, 228–231 (2010).
Hall, J.A. et al. Commensal DNA limits regulatory T cell conversion and is a natural adjuvant of intestinal immune responses. Immunity 29, 637–649 (2008).
Liu, N., Ohnishi, N., Ni, L., Akira, S. & Bacon, K.B. CpG directly induces T-bet expression and inhibits IgG1 and IgE switching in B cells. Nat. Immunol. 4, 687–693 (2003).
Gessner, A., Mohrs, K. & Mohrs, M. Mast cells, basophils, and eosinophils acquire constitutive IL-4 and IL-13 transcripts during lineage differentiation that are sufficient for rapid cytokine production. J. Immunol. 174, 1063–1072 (2005).
Siracusa, M.C., Perrigoue, J.G., Comeau, M.R. & Artis, D. New paradigms in basophil development, regulation and function. Immunol. Cell Biol. 88, 275–284 (2010).
Ohmori, K. et al. IL-3 induces basophil expansion in vivo by directing granulocyte-monocyte progenitors to differentiate into basophil lineage-restricted progenitors in the bone marrow and by increasing the number of basophil/mast cell progenitors in the spleen. J. Immunol. 182, 2835–2841 (2009).
Layland, L.E., Wagner, H. & da Costa, C.U. Lack of antigen-specific TH1 response alters granuloma formation and composition in Schistosoma mansoni-infected MyD88−/− mice. Eur. J. Immunol. 35, 3248–3257 (2005).
Ku, C.L. et al. Selective predisposition to bacterial infections in IRAK-4–deficient children: IRAK-4–dependent TLRs are otherwise redundant in protective immunity. J. Exp. Med. 204, 2407–2422 (2007).
Kim, B.S. et al. Conversion of TH2 memory cells into Foxp3+ regulatory T cells suppressing TH2-mediated allergic asthma. Proc. Natl. Acad. Sci. USA 107, 8742–8747 (2010).
Fink, L.N. et al. Establishment of tolerance to commensal bacteria requires a complex microbiota and is accompanied by decreased intestinal chemokine expression. Am. J. Physiol. Gastrointest. Liver Physiol. 302, G55–G65 (2011).
Redhu, N.S. et al. IgE induces transcriptional regulation of thymic stromal lymphopoietin in human airway smooth muscle cells. J. Allergy Clin. Immunol. 128, 892-896.e2 (2011).
Chen, M.L., Yan, B.S., Bando, Y., Kuchroo, V.K. & Weiner, H.L. Latency-associated peptide identifies a novel CD4+CD25+ regulatory T cell subset with TGFβ-mediated function and enhanced suppression of experimental autoimmune encephalomyelitis. J. Immunol. 180, 7327–7337 (2008).
Acknowledgements
We thank members of the Artis lab for helpful discussions; M.R. Comeau (Tslp−/− mice, Amgen), H. Oettgen (Igh-7−/− mice, Harvard University) and J. Weiser (Nod1−/− mice, University of Pennsylvania) for providing access to mice; L. Shawver, R. Sinha and R. Custers-Allen for assistance with data; J. Sawalle-Belohradsky and B. Hagl for performing DOCK8 sequencing; A. Jansson, G. Notheis, B.H. Belohradsky, M. Albert and the referring physicians for patient care; the Matthew J. Ryan Veterinary Hospital Pathology Lab; the National Institute of Diabetes and Digestive and Kidney Disease Center for the Molecular Studies in Digestive and Liver Disease Molecular Pathology and Imaging Core (DK50306); the Abramson Cancer Center Flow Cytometry and Cell Sorting Resource Laboratory (supported by US National Cancer Institute Comprehensive Cancer Center Support grant (2-P30 CA016520) for technical advice and support; and the University of Pennsylvania Gnotobiotic Mouse Facility for germ-free mice. Research in the Artis lab is supported by the US National Institutes of Health (AI061570, AI087990, AI074878, AI095608, AI083480 and AI095466 to D.A.; T32-AI060516 to D.A.H.; F32-AI085828 to M.C.S.; T32-AI05528 to M.C.A.) the Burroughs Wellcome Fund Investigator in Pathogenesis of Infectious Disease Award (D.A.), the Penn Genome Frontiers Institute (D.A. and F.D.B.), and pilot grants from the University of Pennsylvania Veterinary Center of Infectious Diseases (D.A.). Additional National Institutes of Health support provided by HL107589 and HL111501 to T.K., AI067946 to J.S.O. and UH2DK083981 to F.D.B.
Author information
Authors and Affiliations
Contributions
D.A.H., M.C.S., M.C.A., B.S.K., D.K. and D.A. designed and performed the research; D.F.L., E.D.R., J.S.O., M.K., T.K. and F.D.B. provided reagents; D.A.H., M.C.S., M.C.A., B.S.K. and D.A. analyzed the data; D.A.H., M.C.S. and D.A. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–10 (PDF 1551 kb)
Rights and permissions
About this article
Cite this article
Hill, D., Siracusa, M., Abt, M. et al. Commensal bacteria–derived signals regulate basophil hematopoiesis and allergic inflammation. Nat Med 18, 538–546 (2012). https://doi.org/10.1038/nm.2657
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2657
This article is cited by
-
Lung microbiome: new insights into the pathogenesis of respiratory diseases
Signal Transduction and Targeted Therapy (2024)
-
Combined oral intake of short and long fructans alters the gut microbiota in food allergy model mice and contributes to food allergy prevention
BMC Microbiology (2023)
-
The potential immunomodulatory role of the gut microbiota in the pathogenesis of asthma: an in vitro study
Scientific Reports (2023)
-
Mechanistic and Therapeutic Approaches to Occupational Exposure-Associated Allergic and Non-Allergic Asthmatic Disease
Current Allergy and Asthma Reports (2023)
-
The Airway Microbiome-IL-17 Axis: a Critical Regulator of Chronic Inflammatory Disease
Clinical Reviews in Allergy & Immunology (2022)