Abstract
Precursors to major histocompatibility complex (MHC) class I–presented peptides with extra NH2-terminal residues can be efficiently trimmed to mature epitopes in the endoplasmic reticulum (ER). Here, we purified from liver microsomes a lumenal, soluble aminopeptidase that removes NH2-terminal residues from many antigenic precursors. It was identified as a metallopeptidase named “adipocyte-derived leucine” or “puromycin-insensitive leucine-specific” aminopeptidase. However, because we localized it to the ER, we propose it be renamed ER–aminopeptidase 1 (ERAP1). ERAP1 is inhibited by agents that block precursor trimming in ER vesicles and although it trimmed NH2-extended precursors, it spared presented peptides of 8 amino acid and less. Like other proteins involved in antigen presentation, ERAP1 is induced by interferon-γ. When overexpressed in vivo, we found that ERAP1 stimulates the processing and presentation of an antigenic precursor in the ER.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Rock, K.L. et al. Inhibitors of the proteasome block the degradation of most cell proteins and the generation of peptides presented on MHC class I molecules. Cell 78, 761–771 (1994).
Harding, C.V. et al. Novel dipeptide aldehydes are proteasome inhibitors and block the MHC-I antigen-processing pathway. J. Immunol. 155, 1767–1775 (1995).
Cerundolo, V. et al. The proteasome-specific inhibitor lactacystin blocks presentation of cytotoxic T lymphocyte epitopes in human and murine cells. Eur. J. Immunol. 27, 336–341 (1997).
Craiu, A. et al. Lactacystin and clasto-lactacystin β-lactone modify multiple proteasome β-subunits and inhibit intracellular protein degradation and major histocompatibility complex class I antigen presentation. J. Biol. Chem. 272, 13437–13445 (1997).
Kisselev, A.F., Akopian, T.N., Woo, K.M. & Goldberg, A.L. The sizes of peptides generated from protein by mammalian 26 and 20S proteasomes. Implications for understanding the degradative mechanism and antigen presentation. J. Biol. Chem. 274, 3363–3371 (1999).
Goldberg, A.L., Cascio, P., Saric, T. & Rock, K.L. The importance of the proteasome and subsequent proteolytic steps in the generation of antigenic peptides. Mol. Immunol. 39, 147–164 (2002).
Botbol, V. & Scornik, O.A. Peptide intermediates in the degradation of cellular proteins. Progress Biol. Res. 180, 573–583 (1985).
Rock, K.L. & Goldberg, A.L. Degradation of cell proteins and the generation of MHC class I-presented peptides. Annu. Rev. Immunol. 17, 739–779 (1999).
Craiu, A., Akopian, T., Goldberg, A. & Rock, K.L. Two distinct proteolytic processes in the generation of a major histocompatibility complex class I-presented peptide. Proc. Natl. Acad. Sci. USA 94, 10850–10855 (1997).
Mo, X.Y., Cascio, P., Lemerise, K., Goldberg, A.L. & Rock, K. Distinct proteolytic processes generate the C and N termini of MHC class I-binding peptides. J. Immunol. 163, 5851–5859 (1999).
Lucchiari-Hartz, M. et al. Cytotoxic T lymphocyte epitopes of HIV-1 Nef: Generation of multiple definitive major histocompatibility complex class I ligands by proteasomes. J. Exp. Med. 191, 239–252 (2000).
Cascio, P., Hilton, C., Kisselev, A.F., Rock, K.L. & Goldberg, A.L. 26S proteasomes and immunoproteasomes produce mainly NH2-extended versions of an antigenic peptide. EMBO J. 20, 2357–2366 (2001).
Paz, P., Brouwenstijn, N., Perry, R. & Shastri, N. Discrete proteolytic intermediates in the MHC class I antigen processing pathway and MHC I-dependent peptide trimming in the ER. Immunity 11, 241–251 (1999).
Serwold, T., Gaw, S. & Shastri, N. ER aminopeptidase generate a unique pool of peptides for MHC class I molecules. Nature Immunol. 2, 644–651 (2001).
Knuehl, C. et al. The murine cytomegalovirus pp89 immunodominant H-2Ld epitope is generated and translocated into the endoplasmic reticulum as an 11-mer precursor peptide. J. Immunol. 167, 1515–1521 (2001).
Beninga, J., Rock, K.L. & Goldberg, A.L. Interferon-γ can stimulate post-proteasomal trimming of the N terminus of an antigenic peptide by inducing leucine aminopeptidase. J. Biol. Chem. 273, 18734–18742 (1998).
Stoltze, L. et al. Two new proteases in the MHC class I processing pathway. Nature Immunol. 1, 413–418 (2000).
Harris, C.A., Hunte, B., Krauss, M.R., Taylor, A. & Epstein, L.B. Induction of leucine aminopeptidase by interferon-γ. Identification by protein microsequencing after purification by preparative two-dimensional gel electrophoresis. J. Biol. Chem. 267, 6865–6869 (1992).
Gaczynska, M., Rock, K.L. & Goldberg, A.L. γ-interferon and expression of MHC genes regulate peptide hydrolysis by proteasomes. Nature 365, 264–267 (1993).
Gaczynska, M., Rock, K.L., Spies, T. & Goldberg, A.L. Peptidase activities of proteasomes are differentially regulated by the major histocompatibility complex-encoded genes for LMP2 and LMP7. Proc. Natl. Acad. Sci. USA 91, 9213–9217 (1994).
Gaczynska, M., Goldberg, A.L., Tanaka, K., Hendil, K.B. & Rock, K.L. Proteasome subunits X and Y alter peptidase activities in opposite ways to the interferon-γ-induced subunits LMP2 and LMP7. J. Biol. Chem. 271, 17275–17280 (1996).
Lauvau, G. et al. Human transporters associated with antigen processing (TAPs) select epitope precursor peptides for processing in the endoplasmic reticulum and presentation to T cells. J. Exp. Med. 190, 1227–1239 (1999).
Neisig, A. et al. Major differences in transporter associated with antigen presentation (TAP)-dependent translocation of MHC class I-presentable peptides and the effect of flanking sequences. J. Immunol. 154, 1273–1279 (1995).
Saric, T. et al. Major histocompatibility complex class I-presented antigenic peptides are degraded in cytosolic extracts primarily by thimet oligopeptidase. J. Biol. Chem. 276, 36474–36481 (2001).
Elliott, T., Willis, A., Cerundolo, V. & Townsend, A. Processing of major histocompatibility class I-restricted antigens in the endoplasmic reticulum. J. Exp. Med. 181, 1481–1491 (1995).
Lobigs, M., Chelvanayagam, G. & Mullbacher, A. Proteolytic processing of peptides in the lumen of the endoplasmic reticulum for antigen presentation by major histocompatibility class I. Eur. J. Immunol. 30, 1496–1506 (2000).
Snyder, H.L., Yewdell, J.W. & Bennink, J.R. Trimming of antigenic peptides in an early secretory compartment. J. Exp. Med. 180, 2389–2394 (1994).
Fruci, D., Niedermann, G., Butler, R.H. & van Endert, P.M. Efficient MHC class I-independent amino-terminal trimming of epitope precursor peptides in the endoplasmic reticulum. Immunity 15, 467–476 (2001).
Brouwenstijn, N., Serwold, T. & Shastri, N. MHC class I molecules can direct proteolytic cleavage of antigenic precursors in the endoplasmic reticulum. Immunity 15, 95–104 (2001).
Komlosh, A. et al. A role for a novel luminal endoplasmic reticulum aminopeptidase in final trimming of 26S proteasome-generated major histocompatability complex class I antigenic peptides. J. Biol. Chem. 276, 30050–30056 (2001).
Lyko, F., Martoglio, B., Jungnickel, B., Rapoport, T.A. & Dobberstein, B. Signal sequence processing in rough microsomes. J. Biol. Chem. 270, 19873–19878 (1995).
Snyder, H.L., Bacik, I., Yewdell, J.W., Behrens, T.W. & Bennink, J.R. Promiscuous liberation of MHC-class I-binding peptides from the C termini of membrane and soluble proteins in the secretory pathway. Eur. J. Immunol. 28, 1339–1346 (1998).
Falk, K., Rötzschke, O. & Rammensee, H.-J. Cellular peptide composition governed by major histocompatibility complex class I molecules. Nature 348, 248–250 (1990).
Menoret, A., Li, Z.H., Niswonger, M.L., Altmeyer, A. & Srivastava, P.K. An endoplasmic reticulum protein implicated in chaperoning peptides to major histocompatibility of class I is an aminopeptidase. J. Biol. Chem. 276, 33313–33318 (2001).
Reed, R.C., Zheng, T. & Nicchitta, C.V. Grp94-associated enzymatic activities: Resolution by chromatographic fractionation. J. Biol. Chem. 277, 25082–25089 (2002).
Hattori, A., Matsumoto, H., Mizutani, S. & Tsujimoto, M. Molecular cloning of adipocyte-derived leucine aminopeptidase highly related to placental leucine aminopeptidase/oxytocinase. J. Biochem. (Tokyo) 125, 931–938 (1999).
Hattori, A. et al. Characterization of recombinant human adipocyte-derived leucine aminopeptidase expressed in Chinese hamster ovary cells. J. Biochem. (Tokyo) 128, 755–762 (2000).
Hattori, A., Matsumoto, K., Mizutani, S. & Tsujimoto, M. Genomic organization of the human adipocyte-derived leucine aminopeptidase gene and its relationship to the placental leucine aminopeptidase/oxytocinase gene. J. Biochem. (Tokyo) 130, 235–241 (2001).
Schomburg, L., Kollmus, H., Friedrichsen, S. & Bauer, K. Molecular characterization of a puromycin-insensitive leucyl-specific aminopeptidase, PILS-AP. Eur. J. Biochem. 267, 3198–3207 (2000).
Roelse, J., Gromme, M., Momburg, F., Hämmerling, G. & Neefjes, J. Trimming of TAP-translocated peptides in the endoplasmic reticulum and in the cytosol during recycling. J. Exp. Med. 11, 1591–1597 (1994).
Keller, S.R., Scott, H.M., Mastick, C.C., Aebersold, R. & Lienhard, G.E. Cloning and characterization of a novel insulin-regulated membrane aminopeptidase from Glut4 vesicles. J. Biol. Chem. 270, 23612–23618 (1995).
Rogi, T., Tsujimoto, M., Nakazato, H., Mizutani, S. & Tomoda, Y. Human placental leucine aminopeptidase/oxytocinase. A new member of type II membrane-spanning zinc metallopeptidase family. J. Biol. Chem. 271, 56–61 (1996).
Yamamoto, N. et al. Identification of 33 polymorphisms in the adipocyte-derived leucine aminopeptidase (ALAP) gene and possible association with hypertension. Hum. Mutat. 19, 251–257 (2002).
Miyashita, H. et al. A mouse orthologue of puromycin-insensitive leucyl-specific aminopeptidase is expressed in endothelial cells and plays an important role in angiogenesis. Blood 99, 3241–3249 (2002).
York, I.A. et al. The ER aminopeptidase, ERAP1, enhances or limits antigen presentation by trimming epitopes to 8–9 residues. Nature Immunol. 3, 1177–1184 (2002).
Kohler, A. et al. The axial channel of the proteasome core particle is gated by the Rpt2 ATPase and controls both substrate entry and product release. Mol. Cell 7, 1143–1152 (2001).
Whitby, F.G. et al. Structural basis for the activation of 20S proteasomes by 11S regulators. Nature 408, 115–120 (2000).
Cascio, P., Call, M., Petre, B.M., Walz, T. & Goldberg, A.L. Properties of the hybrid form of the 26S proteasome containing both 19S and PA28 complexes. EMBO J. 21, 2636–2645 (2002).
Walter, P. & Blobel, G. Preparation of microsomal membranes for cotranslational protein translocation. Meth. Enzymol. 96, 84–93 (1983).
Wang, L. & Dobberstein, B. Oligomeric complexes involved in translocation of proteins across the membrane of the endoplasmic reticulum. FEBS Lett. 457, 316–322 (1999).
Porgador, A., Yewdell, J.W., Deng, Y., Bennink, J.R. & Germain, R.N. Localization, quantitation, and in situ detection of specific peptide-MHC class I complexes using a monoclonal antibody. Immunity 6, 715–726 (1997).
Acknowledgements
We thank S. Trombley for assistance in preparing this manuscript, A. Tibebu Kassa in for help with experiments and M. Jedrychowski and S. Gygi for LC-MS/MS analyses. Supported by grants from the NIH (to A. L. G. and K. L. R.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Web Fig. 1.
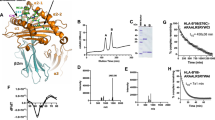
Gel filtration chromatography of ER-lumenal proteins. (a) The ER-lumenal proteins were fractionated on a Sephacryl S-200 HR size exclusion column. The elution positions of aldolase (158 kD), carbonic anhydrase (29 kD) and cytochrome C (12.4 kD) are marked with arrows. Only one peak of aminopeptidase activity was detected with an apparent molecular weight of ~150 kD (Kav = 0.1083). (b) Fractions encompassing the aminopeptidase peak (23–36) were analyzed by electrophoresis on 4–12% NuPAGE gel and the elution profiles of ERAP1 (A-LAP) and Grp94 (gp96) were determined by immunoblotting. (PDF 508 kb)
Web Fig. 2.
Recombinant ERAP1 can remove a variety of residues from NH2-extended antigenic peptides. (a) Reactions contained 150 nmol/ml of QLESIINFEKL and 3 μg/ml of recombinant human enzyme. At the indicated times, an aliquot was removed and fractionated by RP-HPLC. (b) Reactions with indicated peptides were performed and analyzed as in a. The amount of a peptide trimmed was calculated by integration of peptide peaks. (PDF 277 kb)
Rights and permissions
About this article
Cite this article
Saric, T., Chang, SC., Hattori, A. et al. An IFN-γ–induced aminopeptidase in the ER, ERAP1, trims precursors to MHC class I–presented peptides. Nat Immunol 3, 1169–1176 (2002). https://doi.org/10.1038/ni859
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni859
This article is cited by
-
Alanine-based spacers promote an efficient antigen processing and presentation in neoantigen polypeptide vaccines
Cancer Immunology, Immunotherapy (2023)
-
Germline modifiers of the tumor immune microenvironment implicate drivers of cancer risk and immunotherapy response
Nature Communications (2023)
-
ERAP1 is a critical regulator of inflammasome-mediated proinflammatory and ER stress responses
BMC Immunology (2022)
-
A guide to antigen processing and presentation
Nature Reviews Immunology (2022)
-
PA28γ–20S proteasome is a proteolytic complex committed to degrade unfolded proteins
Cellular and Molecular Life Sciences (2022)