Abstract
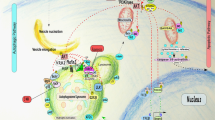
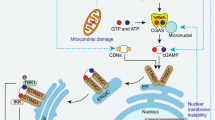
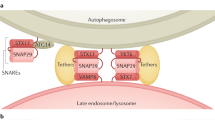
Autophagy is an intracellular bulk degradation system that is highly conserved in eukaryotes. The discovery of autophagy-related ('ATG') proteins in the 1990s greatly advanced the mechanistic understanding of autophagy and clarified the fact that autophagy serves important roles in various biological processes. In addition, studies have revealed other roles for the autophagic machinery beyond autophagy. In this Review, we introduce advances in the knowledge of the roles of autophagy and its components in immunity, including innate immunity, inflammatory responses and adaptive immunity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout

Kim Caesar/Nature Publishing Group

Kim Caesar/Nature Publishing Group

Kim Caesar/Nature Publishing Group

Kim Caesar/Nature Publishing Group
Similar content being viewed by others
References
Mizushima, N. & Komatsu, M. Autophagy: renovation of cells and tissues. Cell 147, 728–741 (2011).
Clark, S.L. Jr. Cellular differentiation in the kidneys of newborn mice studies with the electron microscope. J. Biophys. Biochem. Cytol. 3, 349–362 (1957).
Novikoff, A.B. The proximal tubule cell in experimental hydronephrosis. J. Biophys. Biochem. Cytol. 6, 136–138 (1959).
Ohsumi, Y. Historical landmarks of autophagy research. Cell Res. 24, 9–23 (2014).
Itakura, E. & Mizushima, N. Characterization of autophagosome formation site by a hierarchical analysis of mammalian Atg proteins. Autophagy 6, 764–776 (2010).
Kageyama, S. et al. The LC3 recruitment mechanism is separate from Atg9L1-dependent membrane formation in the autophagic response against Salmonella. Mol. Biol. Cell 22, 2290–2300 (2011). Study demonstrating that early ATG proteins are recruited to intracellular Salmonella.
Matsunaga, K. et al. Autophagy requires endoplasmic reticulum targeting of the PI3-kinase complex via Atg14L. J. Cell Biol. 190, 511–521 (2010).
Hamasaki, M. et al. Autophagosomes form at ER-mitochondria contact sites. Nature 495, 389–393 (2013). Study identifying ER-mitochondria contact sites as the initiation sites for autophagosome formation.
Dooley, H.C. et al. WIPI2 links LC3 conjugation with PI3P, autophagosome formation, and pathogen clearance by recruiting Atg12–5-16L1. Mol. Cell 55, 238–252 (2014).
Mizushima, N. et al. Dissection of autophagosome formation using Apg5-deficient mouse embryonic stem cells. J. Cell Biol. 152, 657–668 (2001).
Nakatogawa, H., Suzuki, K., Kamada, Y. & Ohsumi, Y. Dynamics and diversity in autophagy mechanisms: lessons from yeast. Nat. Rev. Mol. Cell Biol. 10, 458–467 (2009).
Fujita, N. et al. Differential involvement of Atg16L1 in Crohn disease and canonical autophagy: analysis of the organization of the Atg16L1 complex in fibroblasts. J. Biol. Chem. 284, 32602–32609 (2009).
Fujita, N. et al. The Atg16L complex specifies the site of LC3 lipidation for membrane biogenesis in autophagy. Mol. Biol. Cell 19, 2092–2100 (2008).
Kabeya, Y. et al. LC3, a mammalian homologue of yeast Apg8p, is localized in autophagosome membranes after processing. EMBO J. 19, 5720–5728 (2000).
Fujita, N. et al. An Atg4B mutant hampers the lipidation of LC3 paralogues and causes defects in autophagosome closure. Mol. Biol. Cell 19, 4651–4659 (2008).
Sou, Y. et al. The Atg8 conjugation system is indispensable for proper development of autophagic isolation membranes in mice. Mol. Biol. Cell 19, 4762–4775 (2008).
Sanjuan, M.A. et al. Toll-like receptor signalling in macrophages links the autophagy pathway to phagocytosis. Nature 450, 1253–1257 (2007). Identification of LAP in this study.
Flannagan, R.S., Cosío, G. & Grinstein, S. Antimicrobial mechanisms of phagocytes and bacterial evasion strategies. Nat. Rev. Microbiol. 7, 355–366 (2009).
Cossart, P. & Sansonetti, P.J. Bacterial invasion: the paradigms of enteroinvasive pathogens. Science 304, 242–248 (2004).
Nakagawa, I. et al. Autophagy defends cells against invading group A Streptococcus. Science 306, 1037–1040 (2004). Study clarifying, through the use of and LC3 and cells deficient in genes encoding ATG proteins, the fact that autophagy can target intracellular bacteria.
Gutierrez, M.G. et al. Autophagy is a defense mechanism inhibiting BCG and Mycobacterium tuberculosis survival in infected macrophages. Cell 119, 753–766 (2004).
Ogawa, M. et al. Escape of intracellular Shigella from autophagy. Science 307, 727–731 (2005).
Birmingham, C.L., Smith, A.C., Bakowski, M.A., Yoshimori, T. & Brumell, J.H. Autophagy controls Salmonella infection in response to damage to the Salmonella-containing vacuole. J. Biol. Chem. 281, 11374–11383 (2006).
Py, B.F., Lipinski, M.M. & Yuan, J. Autophagy limits Listeria monocytogenes intracellular growth in the early phase of primary infection. Autophagy 3, 117–125 (2007).
Huang, J. & Brumell, J.H. Bacteria-autophagy interplay: a battle for survival. Nat. Rev. Microbiol. 12, 101–114 (2014).
Welchman, R.L., Gordon, C. & Mayer, R.J. Ubiquitin and ubiquitin-like proteins as multifunctional signals. Nat. Rev. Mol. Cell Biol. 6, 599–609 (2005).
Kim, P.K., Hailey, D.W., Mullen, R.T. & Lippincott-Schwartz, J. Ubiquitin signals autophagic degradation of cytosolic proteins and peroxisomes. Proc. Natl. Acad. Sci. USA 105, 20567–20574 (2008).
Fujita, N. & Yoshimori, T. Ubiquitination-mediated autophagy against invading bacteria. Curr. Opin. Cell Biol. 23, 492–497 (2011).
Ng, A.C.Y. et al. Human leucine-rich repeat proteins: a genome-wide bioinformatic categorization and functional analysis in innate immunity. Proc. Natl. Acad. Sci. USA 108, 4631–4638 (2011).
Huett, A. et al. The LRR and RING domain protein LRSAM1 is an E3 ligase crucial for ubiquitin-dependent autophagy of intracellular Salmonella Typhimurium. Cell Host Microbe 12, 778–790 (2012).
Manzanillo, P.S. et al. The ubiquitin ligase parkin mediates resistance to intracellular pathogens. Nature 501, 512–516 (2013).
Narendra, D., Tanaka, A., Suen, D.-F. & Youle, R.J. Parkin is recruited selectively to impaired mitochondria and promotes their autophagy. J. Cell Biol. 183, 795–803 (2008).
Dupont, N. et al. Shigella phagocytic vacuolar membrane remnants participate in the cellular response to pathogen invasion and are regulated by autophagy. Cell Host Microbe 6, 137–149 (2009).
Kobayashi, S. et al. Artificial induction of autophagy around polystyrene beads in nonphagocytic cells. Autophagy 6, 36–45 (2010).
Fujita, N. et al. Recruitment of the autophagic machinery to endosomes during infection is mediated by ubiquitin. J. Cell Biol. 203, 115–128 (2013). Study demonstrating that ubiquitination is required for the recruitment of autophagy-related proteins to ruptured endosomes, and clarifying the mechanism for the recruitment of ATG16L1 to ubiquitinated endosomes.
Hung, Y.-H., Chen, L.M.-W., Yang, J.-Y. & Yuan Yang, W. Spatiotemporally controlled induction of autophagy-mediated lysosome turnover. Nat. Commun. 4, 2111 (2013).
Maejima, I. et al. Autophagy sequesters damaged lysosomes to control lysosomal biogenesis and kidney injury. EMBO J. 32, 2336–2347 (2013).
Thurston, T.L.M., Wandel, M.P., von Muhlinen, N., Foeglein, A. & Randow, F. Galectin 8 targets damaged vesicles for autophagy to defend cells against bacterial invasion. Nature 482, 414–418 (2012). Study identifying galectin-8 as an ubiquitin-independent mediator of the recruitment of NDP52 to ruptured endosomes.
Dreux, M. & Chisari, F.V. Viruses and the autophagy machinery. Cell Cycle 9, 1295–1307 (2010).
Yordy, B., Tal, M.C., Hayashi, K., Arojo, O. & Iwasaki, A. Autophagy and selective deployment of Atg proteins in antiviral defense. Int. Immunol. 25, 1–10 (2013).
Liang, X.H. et al. Protection against fatal Sindbis virus encephalitis by beclin, a novel Bcl-2-interacting protein. J. Virol. 72, 8586–8596 (1998).
Orvedahl, A. et al. Autophagy protects against Sindbis virus infection of the central nervous system. Cell Host Microbe 7, 115–127 (2010).
Orvedahl, A. et al. Image-based genome-wide siRNA screen identifies selective autophagy factors. Nature 480, 113–117 (2011).
Kyei, G.B. et al. Autophagy pathway intersects with HIV-1 biosynthesis and regulates viral yields in macrophages. J. Cell Biol. 186, 255–268 (2009).
Shoji-Kawata, S. et al. Identification of a candidate therapeutic autophagy-inducing peptide. Nature 494, 201–206 (2013).
Martinez, J. et al. Microtubule-associated protein 1 light chain 3 α (LC3)-associated phagocytosis is required for the efficient clearance of dead cells. Proc. Natl. Acad. Sci. USA 108, 17396–17401 (2011).
Martinez, J. et al. Molecular characterization of LC3-associated phagocytosis reveals distinct roles for Rubicon, NOX2 and autophagy proteins. Nat. Cell Biol. 17, 893–906 (2015).
Henault, J. et al. Noncanonical autophagy is required for type I interferon secretion in response to DNA-immune complexes. Immunity 37, 986–997 (2012).
Starr, T. et al. Selective subversion of autophagy complexes facilitates completion of the Brucella intracellular cycle. Cell Host Microbe 11, 33–45 (2012).
Zhao, Z. et al. Autophagosome-independent essential function for the autophagy protein Atg5 in cellular immunity to intracellular pathogens. Cell Host Microbe 4, 458–469 (2008).
Hwang, S. et al. Nondegradative role of Atg5-Atg12/ Atg16L1 autophagy protein complex in antiviral activity of interferon γ. Cell Host Microbe 11, 397–409 (2012).
Taylor, M.P., Burgon, T.B., Kirkegaard, K. & Jackson, W.T. Role of microtubules in extracellular release of poliovirus. J. Virol. 83, 6599–6609 (2009).
Robinson, S.M. et al. Coxsackievirus B exits the host cell in shed microvesicles displaying autophagosomal markers. PLoS Pathog. 10, e1004045 (2014).
Miao, Y., Li, G., Zhang, X., Xu, H. & Abraham, S.N. A TRP channel senses lysosome neutralization by pathogens to trigger their expulsion. Cell 161, 1306–1319 (2015).
Kawai, T. & Akira, S. The roles of TLRs, RLRs and NLRs in pathogen recognition. Int. Immunol. 21, 317–337 (2009).
Deretic, V., Saitoh, T. & Akira, S. Autophagy in infection, inflammation and immunity. Nat. Rev. Immunol. 13, 722–737 (2013).
Latz, E., Xiao, T.S. & Stutz, A. Activation and regulation of the inflammasomes. Nat. Rev. Immunol. 13, 397–411 (2013).
Hampe, J. et al. A genome-wide association scan of nonsynonymous SNPs identifies a susceptibility variant for Crohn disease in ATG16L1. Nat. Genet. 39, 207–211 (2007).
Rioux, J.D. et al. Genome-wide association study identifies new susceptibility loci for Crohn disease and implicates autophagy in disease pathogenesis. Nat. Genet. 39, 596–604 (2007).
Saitoh, T. et al. Loss of the autophagy protein Atg16L1 enhances endotoxin-induced IL-1β production. Nature 456, 264–268 (2008). First study to report that autophagy-deficiency enhances inflammasome activation.
Murthy, A. et al. A Crohn's disease variant in Atg16l1 enhances its degradation by caspase 3. Nature 506, 456–462 (2014). Study providing evidence that the Crohn's disease–associated variant of ATG16L1 enhances inflammation in vivo.
Lassen, K.G. et al. Atg16L1 T300A variant decreases selective autophagy resulting in altered cytokine signaling and decreased antibacterial defense. Proc. Natl. Acad. Sci. USA 111, 7741–7746 (2014).
Plantinga, T.S. et al. Crohn's disease-associated ATG16L1 polymorphism modulates pro-inflammatory cytokine responses selectively upon activation of NOD2. Gut 60, 1229–1235 (2011).
Cadwell, K. et al. Virus-plus-susceptibility gene interaction determines Crohn's disease gene Atg16L1 phenotypes in intestine. Cell 141, 1135–1145 (2010).
Zhou, R., Yazdi, A.S., Menu, P. & Tschopp, J. A role for mitochondria in NLRP3 inflammasome activation. Nature 469, 221–225 (2011). First study to report that autophagy targets old or dysfunctional mitochondria to limit inflammasome activation.
Nakahira, K. et al. Autophagy proteins regulate innate immune responses by inhibiting the release of mitochondrial DNA mediated by the NALP3 inflammasome. Nat. Immunol. 12, 222–230 (2011).
Lupfer, C. et al. Receptor interacting protein kinase 2-mediated mitophagy regulates inflammasome activation during virus infection. Nat. Immunol. 14, 480–488 (2013).
Wen, H. et al. Fatty acid-induced NLRP3-ASC inflammasome activation interferes with insulin signaling. Nat. Immunol. 12, 408–415 (2011).
Razani, B. et al. Autophagy links inflammasomes to atherosclerotic progression. Cell Metab. 15, 534–544 (2012).
Wang, C. et al. Atg16L1 deficiency confers protection from uropathogenic Escherichia coli infection in vivo. Proc. Natl. Acad. Sci. USA 109, 11008–11013 (2012).
Shaw, S.Y. et al. Selective modulation of autophagy, innate immunity, and adaptive immunity by small molecules. ACS Chem. Biol. 8, 2724–2733 (2013).
Guo, W. et al. Small molecule-driven mitophagy-mediated NLRP3 inflammasome inhibition is responsible for the prevention of colitis-associated cancer. Autophagy 10, 972–985 (2014).
Abderrazak, A. et al. Anti-inflammatory and anti-atherogenic effects of the inflammasome NLRP3 inhibitor, arglabin, in ApoE2.Ki mice fed a high fat diet. Circulation 131, 1061–1070 (2015).
Shi, C.-S. et al. Activation of autophagy by inflammatory signals limits IL-1β production by targeting ubiquitinated inflammasomes for destruction. Nat. Immunol. 13, 255–263 (2012).
Harris, J. et al. Autophagy controls IL-1β secretion by targeting pro-IL-1β for degradation. J. Biol. Chem. 286, 9587–9597 (2011).
Dupont, N. et al. Autophagy-based unconventional secretory pathway for extracellular delivery of IL-1β. EMBO J. 30, 4701–4711 (2011).
Lamkanfi, M. & Dixit, V.M. Mechanisms and functions of inflammasomes. Cell 157, 1013–1022 (2014).
Meunier, E. et al. Caspase-11 activation requires lysis of pathogen-containing vacuoles by IFN-induced GTPases. Nature 509, 366–370 (2014).
Netea, M.G., van de Veerdonk, F.L., van der Meer, J.W.M., Dinarello, C.A. & Joosten, L.A.B. Inflammasome-independent regulation of IL-1-family cytokines. Annu. Rev. Immunol. 33, 49–77 (2014).
Castillo, E.F. et al. Autophagy protects against active tuberculosis by suppressing bacterial burden and inflammation. Proc. Natl. Acad. Sci. USA 109, E3168–E3176 (2012).
Kawai, T. & Akira, S. The role of pattern-recognition receptors in innate immunity: update on Toll-like receptors. Nat. Immunol. 11, 373–384 (2010).
Yoneyama, M., Onomoto, K., Jogi, M., Akaboshi, T. & Fujita, T. Viral RNA detection by RIG-I-like receptors. Curr. Opin. Immunol. 32, 48–53 (2015).
Cai, X., Chiu, Y.-H. & Chen, Z.J. The cGAS-cGAMP-STING pathway of cytosolic DNA sensing and signaling. Mol. Cell 54, 289–296 (2014).
Lee, H.K., Lund, J.M., Ramanathan, B., Mizushima, N. & Iwasaki, A. Autophagy-dependent viral recognition by plasmacytoid dendritic cells. Science 315, 1398–1401 (2007). First study to report involvement of ATG proteins in PRR-mediated innate immune response.
Tal, M.C. et al. Absence of autophagy results in reactive oxygen species-dependent amplification of RLR signaling. Proc. Natl. Acad. Sci. USA 106, 2770–2775 (2009).
Jounai, N. et al. The Atg5 Atg12 conjugate associates with innate antiviral immune responses. Proc. Natl. Acad. Sci. USA 104, 14050–14055 (2007).
Lei, Y. et al. The mitochondrial proteins NLRX1 and TUFM form a complex that regulates type I interferon and autophagy. Immunity 36, 933–946 (2012).
Saitoh, T. et al. Atg9a controls dsDNA-driven dynamic translocation of STING and the innate immune response. Proc. Natl. Acad. Sci. USA 106, 20842–20846 (2009). First study to report involvement of ATG proteins in cytosolic dsDNA–induced innate immune response.
Konno, H., Konno, K. & Barber, G.N. Cyclic dinucleotides trigger ULK1 (ATG1) phosphorylation of STING to prevent sustained innate immune signaling. Cell 155, 688–698 (2013).
Liang, Q. et al. Crosstalk between the cGAS DNA sensor and Beclin-1 autophagy protein shapes innate antimicrobial immune responses. Cell Host Microbe 15, 228–238 (2014).
Watson, R.O., Manzanillo, P.S. & Cox, J.S. Extracellular M. tuberculosis DNA targets bacteria for autophagy by activating the host DNA-sensing pathway. Cell 150, 803–815 (2012).
Dey, B. et al. A bacterial cyclic dinucleotide activates the cytosolic surveillance pathway and mediates innate resistance to tuberculosis. Nat. Med. 21, 401–406 (2015).
Collins, A.C. et al. Cyclic GMP-AMP synthase is an innate immune DNA sensor for Mycobacterium tuberculosis. Cell Host Microbe 17, 820–828 (2015).
Blum, J.S., Wearsch, P.A. & Cresswell, P. Pathways of antigen processing. Annu. Rev. Immunol. 31, 443–473 (2013).
Romao, S., Gannage, M. & Münz, C. Checking the garbage bin for problems in the house, or how autophagy assists in antigen presentation to the immune system. Semin. Cancer Biol. 23, 391–396 (2013).
Paludan, C. et al. Endogenous MHC class II processing of a viral nuclear antigen after autophagy. Science 307, 593–596 (2005).
Leung, C.S., Haigh, T.A., Mackay, L.K., Rickinson, A.B. & Taylor, G.S. Nuclear location of an endogenously expressed antigen, EBNA1, restricts access to macroautophagy and the range of CD4 epitope display. Proc. Natl. Acad. Sci. USA 107, 2165–2170 (2010).
Jagannath, C. et al. Autophagy enhances the efficacy of BCG vaccine by increasing peptide presentation in mouse dendritic cells. Nat. Med. 15, 267–276 (2009).
Schmid, D., Pypaert, M. & Münz, C. Antigen-loading compartments for major histocompatibility complex class II molecules continuously receive input from autophagosomes. Immunity 26, 79–92 (2007). Study demonstrating that autophagosomes deliver cytosolic antigen to MHC class II loading compartments.
Jin, Y. et al. Regulation of SIV antigen-specific CD4+ T cellular immunity via autophagosome-mediated MHC II molecule-targeting antigen presentation in mice. PLoS ONE 9, e93143 (2014).
Nedjic, J., Aichinger, M., Emmerich, J., Mizushima, N. & Klein, L. Autophagy in thymic epithelium shapes the T-cell repertoire and is essential for tolerance. Nature 455, 396–400 (2008).
Kasai, M. et al. Autophagic compartments gain access to the MHC class II compartments in thymic epithelium. J. Immunol. 183, 7278–7285 (2009).
Aichinger, M., Wu, C., Nedjic, J. & Klein, L. Macroautophagy substrates are loaded onto MHC class II of medullary thymic epithelial cells for central tolerance. J. Exp. Med. 210, 287–300 (2013). Study demonstrating that autophagic cargo is presented on MHC class II molecules for negative selection of CD4+ T cells in the thymus.
Florey, O., Kim, S.E., Sandoval, C.P., Haynes, C.M. & Overholtzer, M. Autophagy machinery mediates macroendocytic processing and entotic cell death by targeting single membranes. Nat. Cell Biol. 13, 1335–1343 (2011).
Romao, S. et al. Autophagy proteins stabilize pathogen-containing phagosomes for prolonged MHC II antigen processing. J. Cell Biol. 203, 757–766 (2013). Study demonstrating that LAP retains endocytosed antigen for prolonged presentation by MHC class II to CD4+ T cells.
Ma, J., Becker, C., Lowell, C.A. & Underhill, D.M. Dectin-1-triggered recruitment of light chain 3 protein to phagosomes facilitates major histocompatibility complex class II presentation of fungal-derived antigens. J. Biol. Chem. 287, 34149–34156 (2012).
Ma, J., Becker, C., Reyes, C. & Underhill, D.M. Cutting edge: FYCO1 recruitment to dectin-1 phagosomes is accelerated by light chain 3 protein and regulates phagosome maturation and reactive oxygen production. J. Immunol. 192, 1356–1360 (2014).
Lee, H.K. et al. In vivo requirement for Atg5 in antigen presentation by dendritic cells. Immunity 32, 227–239 (2010).
Ireland, J.M. & Unanue, E.R. Autophagy in antigen-presenting cells results in presentation of citrullinated peptides to CD4 T cells. J. Exp. Med. 208, 2625–2632 (2011).
Chaturvedi, A., Dorward, D. & Pierce, S.K. The B cell receptor governs the subcellular location of Toll-like receptor 9 leading to hyperresponses to DNA-containing antigens. Immunity 28, 799–809 (2008).
Pengo, N. et al. Plasma cells require autophagy for sustainable immunoglobulin production. Nat. Immunol. 14, 298–305 (2013). Study demonstrating that plasma cells require autophagy for their antibody-producing function.
Wenger, T. et al. Autophagy inhibition promotes defective neosynthesized proteins storage in ALIS, and induces redirection toward proteasome processing and MHCI-restricted presentation. Autophagy 8, 350–363 (2012).
Kristensen, A.R. et al. Ordered organelle degradation during starvation-induced autophagy. Mol. Cell. Proteomics 7, 2419–2428 (2008).
Cuervo, A.M., Palmer, A., Rivett, A.J. & Knecht, E. Degradation of proteasomes by lysosomes in rat liver. Eur. J. Biochem. 227, 792–800 (1995).
Marshall, R.S., Li, F., Gemperline, D.C., Book, A.J. & Vierstra, R.D. Autophagic degradation of the 26S proteasome is mediated by the dual ATG8/ubiquitin receptor RPN10 in Arabidopsis. Mol. Cell 58, 1053–1066 (2015).
English, L. et al. Autophagy enhances the presentation of endogenous viral antigens on MHC class I molecules during HSV-1 infection. Nat. Immunol. 10, 480–487 (2009).
Tey, S.-K. & Khanna, R. Autophagy mediates transporter associated with antigen processing-independent presentation of viral epitopes through MHC class I pathway. Blood 120, 994–1004 (2012). Study demonstrating that autophagy can contribute to the loading of antigen onto MHC class I molecules under TAP-blocking conditions.
Li, B. et al. Autophagy facilitates major histocompatibility complex class I expression induced by IFN-γ in B16 melanoma cells. Cancer Immunol. Immunother. 59, 313–321 (2010).
Johnstone, C. et al. Exogenous, TAP-independent lysosomal presentation of a respiratory syncytial virus CTL epitope. Immunol. Cell Biol. 90, 978–982 (2012).
Blanchet, F.P. et al. Human immunodeficiency Virus-1 inhibition of immunoamphisomes in dendritic cells impairs early innate and adaptive immune responses. Immunity 32, 654–669 (2010).
Fiegl, D. et al. Amphisomal route of MHC class I cross-presentation in bacteria-infected dendritic cells. J. Immunol. 190, 2791–2806 (2013).
De Luca, A. et al. CD4+ T cell vaccination overcomes defective cross-presentation of fungal antigens in a mouse model of chronic granulomatous disease. J. Clin. Invest. 122, 1816–1831 (2012).
Duran, J.M., Anjard, C., Stefan, C., Loomis, W.F. & Malhotra, V. Unconventional secretion of Acb1 is mediated by autophagosomes. J. Cell Biol. 188, 527–536 (2010).
Manjithaya, R., Anjard, C., Loomis, W.F. & Subramani, S. Unconventional secretion of Pichia pastoris Acb1 is dependent on GRASP protein, peroxisomal functions, and autophagosome formation. J. Cell Biol. 188, 537–546 (2010). Two studies (refs. 123,124) establishing the fact that the autophagic machinery contributes to signal peptide–independent secretion.
Thorburn, J. et al. Autophagy regulates selective HMGB1 release in tumor cells that are destined to die. Cell Death Differ. 16, 175–183 (2009).
Martins, I. et al. Molecular mechanisms of ATP secretion during immunogenic cell death. Cell Death Differ. 21, 79–91 (2014).
DeSelm, C.J. et al. Autophagy proteins regulate the secretory component of osteoclastic bone resorption. Dev. Cell 21, 966–974 (2011).
Jackson, W.T. et al. Subversion of cellular autophagosomal machinery by RNA viruses. PLoS Biol. 3, e156 (2005).
Nowag, H. et al. Macroautophagy proteins assist Epstein Barr virus production and Get incorporated into the virus particles. EBioMedicine 1, 116–125 (2014).
Uhl, M. et al. Autophagy within the antigen donor cell facilitates efficient antigen cross-priming of virus-specific CD8+ T cells. Cell Death Differ. 16, 991–1005 (2009).
Li, Y. et al. Efficient cross-presentation depends on autophagy in tumor cells. Cancer Res. 68, 6889–6895 (2008).
Ye, W. et al. Cross-presentation of viral antigens in dribbles leads to efficient activation of virus-specific human memory T cells. J. Transl. Med. 12, 100 (2014).
Mortensen, M. et al. The autophagy protein Atg7 is essential for hematopoietic stem cell maintenance. J. Exp. Med. 208, 455–467 (2011). Study demonstrating which hematopoietic lineages requires autophagy for their development.
Pua, H.H., Dzhagalov, I., Chuck, M., Mizushima, N. & He, Y.-W. A critical role for the autophagy gene Atg5 in T cell survival and proliferation. J. Exp. Med. 204, 25–31 (2007).
Arsov, I. et al. A role for autophagic protein beclin 1 early in lymphocyte development. J. Immunol. 186, 2201–2209 (2011).
Pua, H.H., Guo, J., Komatsu, M. & He, Y.-W. Autophagy is essential for mitochondrial clearance in mature T lymphocytes. J. Immunol. 182, 4046–4055 (2009).
Farfariello, V., Amantini, C. & Santoni, G. Transient receptor potential vanilloid 1 activation induces autophagy in thymocytes through ROS-regulated AMPK and Atg4C pathways. J. Leukoc. Biol. 92, 421–431 (2012).
Willinger, T. & Flavell, R.A. Canonical autophagy dependent on the class III phosphoinositide-3 kinase Vps34 is required for naive T-cell homeostasis. Proc. Natl. Acad. Sci. USA 109, 8670–8675 (2012).
Henson, S.M. et al. p38 signaling inhibits mTORC1-independent autophagy in senescent human CD8+ T cells. J. Clin. Invest. 124, 4004–4016 (2014).
Jia, W., Pua, H.H., Li, Q.-J. & He, Y.-W. Autophagy regulates endoplasmic reticulum homeostasis and calcium mobilization in T lymphocytes. J. Immunol. 186, 1564–1574 (2011).
Parekh, V.V. et al. Impaired autophagy, defective T cell homeostasis, and a wasting syndrome in mice with a T cell-specific deletion of Vps34. J. Immunol. 190, 5086–5101 (2013).
Kovacs, J.R. et al. Autophagy promotes T-cell survival through degradation of proteins of the cell death machinery. Cell Death Differ. 19, 144–152 (2012).
Puleston, D. J. et al. Autophagy is a critical regulator of memory CD8+ T cell formation. eLife 3, e03706 (2014).
Xu, X. et al. Autophagy is essential for effector CD8+ T cell survival and memory formation. Nat. Immunol. 15, 1152–1161 (2014).
Miller, B.C. et al. The autophagy gene ATG5 plays an essential role in B lymphocyte development. Autophagy 4, 309–314 (2008).
Conway, K.L. et al. ATG5 regulates plasma cell differentiation. Autophagy 9, 528–537 (2013).
Milan, E. et al. A plastic SQSTM1/p62-dependent autophagic reserve maintains proteostasis and determines proteasome inhibitor susceptibility in multiple myeloma cells. Autophagy 11, 1161–1178 (2015).
Chen, M. et al. Essential role for autophagy in the maintenance of immunological memory against influenza infection. Nat. Med. 20, 503–510 (2014). Study demonstrating that the maintenance of memory B cells requires autophagy.
Benjamin, J.L., Sumpter, R. Jr., Levine, B. & Hooper, L.V. Intestinal epithelial autophagy is essential for host defense against invasive bacteria. Cell Host Microbe 13, 723–734 (2013).
Conway, K.L. et al. Atg16l1 is required for autophagy in intestinal epithelial cells and protection of mice from Salmonella infection. Gastroenterology 145, 1347–1357 (2013).
Yano, T. et al. Autophagic control of listeria through intracellular innate immune recognition in Drosophila. Nat. Immunol. 9, 908–916 (2008).
Shelly, S., Lukinova, N., Bambina, S., Berman, A. & Cherry, S. Autophagy is an essential component of Drosophila immunity against vesicular stomatitis virus. Immunity 30, 588–598 (2009).
Jia, K. et al. Autophagy genes protect against Salmonella typhimurium infection and mediate insulin signaling-regulated pathogen resistance. Proc. Natl. Acad. Sci. USA 106, 14564–14569 (2009).
Cadwell, K. et al. A key role for autophagy and the autophagy gene Atg16l1 in mouse and human intestinal Paneth cells. Nature 456, 259–263 (2008). Study demonstrating that ATG16L1 and ATG5 are required for maintenance of the granule-secretion pathway in Paneth cells in the intestine.
Marchiando, A.M. et al. A deficiency in the autophagy gene Atg16L1 enhances resistance to enteric bacterial infection. Cell Host Microbe 14, 216–224 (2013).
Xu, Y. et al. Toll-like receptor 4 is a sensor for autophagy associated with innate immunity. Immunity 27, 135–144 (2007).
Delgado, M.A., Elmaoued, R.A., Davis, A.S., Kyei, G. & Deretic, V. Toll-like receptors control autophagy. EMBO J. 27, 1110–1121 (2008).
Travassos, L.H. et al. Nod1 and Nod2 direct autophagy by recruiting ATG16L1 to the plasma membrane at the site of bacterial entry. Nat. Immunol. 11, 55–62 (2010).
Cooney, R. et al. NOD2 stimulation induces autophagy in dendritic cells influencing bacterial handling and antigen presentation. Nat. Med. 16, 90–97 (2010).
Jounai, N. et al. NLRP4 negatively regulates autophagic processes through an association with Beclin1. J. Immunol. 186, 1646–1655 (2011).
Yanai, H. et al. Conditional ablation of HMGB1 in mice reveals its protective function against endotoxemia and bacterial infection. Proc. Natl. Acad. Sci. USA 110, 20699–20704 (2013).
Tang, D. et al. Endogenous HMGB1 regulates autophagy. J. Cell Biol. 190, 881–892 (2010).
Espert, L. et al. Autophagy is involved in T cell death after binding of HIV-1 envelope proteins to CXCR4. J. Clin. Invest. 116, 2161–2172 (2006).
Singh, S.B., Davis, A.S., Taylor, G.A. & Deretic, V. Human IRGM induces autophagy to eliminate intracellular mycobacteria. Science 313, 1438–1441 (2006).
Keller, C.W. et al. TNF-α induces macroautophagy and regulates MHC class II expression in human skeletal muscle cells. J. Biol. Chem. 286, 3970–3980 (2011).
Mostowy, S. et al. p62 and NDP52 proteins target intracytosolic Shigella and Listeria to different autophagy pathways. J. Biol. Chem. 286, 26987–26995 (2011).
Pilli, M. et al. TBK-1 promotes autophagy-mediated antimicrobial defense by controlling autophagosome maturation. Immunity 37, 223–234 (2012).
Harris, J. et al. T helper 2 cytokines inhibit autophagic control of intracellular Mycobacterium tuberculosis. Immunity 27, 505–517 (2007).
Shen, S. et al. Cytoplasmic STAT3 represses autophagy by inhibiting PKR activity. Mol. Cell 48, 667–680 (2012).
Acknowledgements
Supported by the Ministry of Education, Culture, Sports, Science and Technology of Japan (S.T.S., T.S. and Y.T.), the Japan Agency for Medical Research and Development CREST (S.T.S., T.S. and Y.T.), Cancer Research Switzerland (KFS-3234-08-2013 for H.N. and C.M.), Worldwide Cancer Research (14-1033 for H.N. and C.M.), the clinical research programs KFSPMS and KFSPHHLD of the University of Zurich (H.N. and C.M.), the Baugarten Foundation (H.N. and C.M.), the Sobek Foundation (H.N. and C.M.), Fondation Acteria (H.N. and C.M.), the Swiss Vaccine Research Institute (H.N. and C.M.) and the Swiss National Science Foundation (310030_143979 and CRSII3_136241 for H.N. and C.M.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Shibutani, S., Saitoh, T., Nowag, H. et al. Autophagy and autophagy-related proteins in the immune system. Nat Immunol 16, 1014–1024 (2015). https://doi.org/10.1038/ni.3273
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3273
This article is cited by
-
The application of nanoparticles-based ferroptosis, pyroptosis and autophagy in cancer immunotherapy
Journal of Nanobiotechnology (2024)
-
Transcription factor EB-mediated autophagy affects cell migration and inhibits apoptosis to promote endometriosis
Apoptosis (2024)
-
Autophagy as a potential mechanism underlying the biological effect of 1,25-Dihydroxyvitamin D3 on periodontitis: a narrative review
BMC Oral Health (2023)
-
Orf virus induces complete autophagy to promote viral replication via inhibition of AKT/mTOR and activation of the ERK1/2/mTOR signalling pathway in OFTu cells
Veterinary Research (2023)
-
Role and clinical implication of autophagy in COVID-19
Virology Journal (2023)