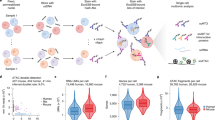
Abstract
Long noncoding RNAs are emerging as important regulators of cellular functions, but little is known of their role in the human immune system. Here we investigated long intergenic noncoding RNAs (lincRNAs) in 13 subsets of T lymphocytes and B lymphocytes by next-generation sequencing–based RNA sequencing (RNA-seq analysis) and de novo transcriptome reconstruction. We identified over 500 previously unknown lincRNAs and described lincRNA signatures. Expression of linc-MAF-4, a chromatin-associated lincRNA specific to the TH1 subset of helper T cells, was inversely correlated with expression of MAF, a TH2-associated transcription factor. Downregulation of linc-MAF-4 skewed T cell differentiation toward the TH2 phenotype. We identified a long-distance interaction between the genomic regions of the gene encoding linc-MAF-4 and MAF, where linc-MAF-4 associated with the chromatin modifiers LSD1 and EZH2; this suggested that linc-MAF-4 regulated MAF transcription through the recruitment of chromatin modifiers. Our results demonstrate a key role for lincRNA in T lymphocyte differentiation.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
Accession codes
References
Zhu, J., Yamane, H. & Paul, W.E. Differentiation of effector CD4 T cell populations. Annu. Rev. Immunol. 28, 445–489 (2010).
Zhou, L., Chong, M.M. & Littman, D.R. Plasticity of CD4+ T cell lineage differentiation. Immunity 30, 646–655 (2009).
O'Shea, J.J. & Paul, W.E. Mechanisms underlying lineage commitment and plasticity of helper CD4+ T cells. Science 327, 1098–1102 (2010).
Kanno, Y., Vahedi, G., Hirahara, K., Singleton, K. & O'Shea, J.J. Transcriptional and epigenetic control of T helper cell specification: molecular mechanisms underlying commitment and plasticity. Annu. Rev. Immunol. 30, 707–731 (2012).
O'Connell, R.M., Rao, D.S., Chaudhuri, A.A. & Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122 (2010).
Pagani, M. et al. Role of microRNAs and long-non-coding RNAs in CD4+ T-cell differentiation. Immunol. Rev. 253, 82–96 (2013).
Cobb, B.S. et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J. Exp. Med. 201, 1367–1373 (2005).
Koralov, S.B. et al. Dicer ablation affects antibody diversity and cell survival in the B lymphocyte lineage. Cell 132, 860–874 (2008).
O'Connell, R.M. et al. MicroRNA-155 promotes autoimmune inflammation by enhancing inflammatory T cell development. Immunity 33, 607–619 (2010).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007).
Rossi, R.L. et al. Distinct microRNA signatures in human lymphocyte subsets and enforcement of the naive state in CD4+ T cells by the microRNA miR-125b. Nat. Immunol. 12, 796–803 (2011).
Cabili, M.N. et al. Integrative annotation of human large intergenic noncoding RNAs reveals global properties and specific subclasses. Genes Dev. 25, 1915–1927 (2011).
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: analysis of their gene structure, evolution, and expression. Genome Res. 22, 1775–1789 (2012).
Hrdlickova, B. et al. Expression profiles of long non-coding RNAs located in autoimmune disease-associated regions reveal immune cell-type specificity. Genome Med 6, 88 (2014).
Fatica, A. & Bozzoni, I. Long non-coding RNAs: new players in cell differentiation and development. Nat. Rev. Genet. 15, 7–21 (2014).
Guttman, M. et al. Chromatin signature reveals over a thousand highly conserved large non-coding RNAs in mammals. Nature 458, 223–227 (2009).
Guttman, M. et al. lincRNAs act in the circuitry controlling pluripotency and differentiation. Nature 477, 295–300 (2011).
Khalil, A.M. et al. Many human large intergenic noncoding RNAs associate with chromatin-modifying complexes and affect gene expression. Proc. Natl. Acad. Sci. USA 106, 11667–11672 (2009).
Yoon, J.H. et al. LincRNA-p21 suppresses target mRNA translation. Mol. Cell 47, 648–655 (2012).
Kretz, M. et al. Control of somatic tissue differentiation by the long non-coding RNA TINCR. Nature 493, 231–235 (2013).
Poliseno, L. et al. A coding-independent function of gene and pseudogene mRNAs regulates tumour biology. Nature 465, 1033–1038 (2010).
Sumazin, P. et al. An extensive microRNA-mediated network of RNA-RNA interactions regulates established oncogenic pathways in glioblastoma. Cell 147, 370–381 (2011).
Cesana, M. et al. A long noncoding RNA controls muscle differentiation by functioning as a competing endogenous RNA. Cell 147, 358–369 (2011).
Pang, K.C. et al. Genome-wide identification of long noncoding RNAs in CD8+ T cells. J. Immunol. 182, 7738–7748 (2009).
Collier, S.P., Collins, P.L., Williams, C.L., Boothby, M.R. & Aune, T.M. Cutting edge: influence of Tmevpg1, a long intergenic noncoding RNA, on the expression of Ifng by Th1 cells. J. Immunol. 189, 2084–2088 (2012).
Gomez, J.A. et al. The NeST long ncRNA controls microbial susceptibility and epigenetic activation of the interferon-γ locus. Cell 152, 743–754 (2013).
Carpenter, S. et al. A long noncoding RNA mediates both activation and repression of immune response genes. Science 341, 789–792 (2013).
Hu, G. et al. Expression and regulation of intergenic long noncoding RNAs during T cell development and differentiation. Nat. Immunol. 14, 1190–1198 (2013).
Haas, B.J. et al. De novo transcript sequence reconstruction from RNA-seq using the Trinity platform for reference generation and analysis. Nat. Protoc. 8, 1494–1512 (2013).
Hart, T., Komori, H.K., LaMere, S., Podshivalova, K. & Salomon, D.R. Finding the active genes in deep RNA-seq gene expression studies. BMC Genomics 14, 778 (2013).
Flicek, P. et al. Ensembl 2013. Nucleic Acids Res. 41, D48–D55 (2013).
Dobin, A. et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Rhind, N. et al. Comparative functional genomics of the fission yeasts. Science 332, 930–936 (2011).
Finn, R.D. et al. The Pfam protein families database. Nucleic Acids Res. 38, D211–D222 (2010).
Lin, M.F., Jungreis, I. & Kellis, M. PhyloCSF: a comparative genomics method to distinguish protein coding and non-coding regions. Bioinformatics 27, i275–i282 (2011).
Guttman, M., Russell, P., Ingolia, N.T., Weissman, J.S. & Lander, E.S. Ribosome profiling provides evidence that large noncoding RNAs do not encode proteins. Cell 154, 240–251 (2013).
Mercer, T.R., Dinger, M.E., Sunkin, S.M., Mehler, M.F. & Mattick, J.S. Specific expression of long noncoding RNAs in the mouse brain. Proc. Natl. Acad. Sci. USA 105, 716–721 (2008).
Ørom, U.A. et al. Long noncoding RNAs with enhancer-like function in human cells. Cell 143, 46–58 (2010).
Al-Shahrour, F., Minguez, P., Vaquerizas, J.M., Conde, L. & Dopazo, J. BABELOMICS: a suite of web tools for functional annotation and analysis of groups of genes in high-throughput experiments. Nucleic Acids Res. 33, W472–W476 (2005).
Volders, P.J. et al. LNCipedia: a database for annotated human lncRNA transcript sequences and structures. Nucleic Acids Res. 41, D246–D251 (2013).
Ho, I.C., Lo, D. & Glimcher, L.H. c-maf promotes T helper cell type 2 (Th2) and attenuates Th1 differentiation by both interleukin 4-dependent and -independent mechanisms. J. Exp. Med. 188, 1859–1866 (1998).
Liu, X., Nurieva, R.I. & Dong, C. Transcriptional regulation of follicular T-helper (Tfh) cells. Immunol. Rev. 252, 139–145 (2013).
Sato, K. et al. Marked induction of c-Maf protein during Th17 cell differentiation and its implication in memory Th cell development. J. Biol. Chem. 286, 14963–14971 (2011).
Mattick, J.S. The genetic signatures of noncoding RNAs. PLoS Genet. 5, e1000459 (2009).
Klattenhoff, C.A. et al. Braveheart, a long noncoding RNA required for cardiovascular lineage commitment. Cell 152, 570–583 (2013).
Cabianca, D.S. et al. A long ncRNA links copy number variation to a polycomb/trithorax epigenetic switch in FSHD muscular dystrophy. Cell 149, 819–831 (2012).
Tsai, M.C. et al. Long noncoding RNA as modular scaffold of histone modification complexes. Science 329, 689–693 (2010).
Kaneko, S. et al. Interactions between JARID2 and noncoding RNAs regulate PRC2 recruitment to chromatin. Mol. Cell 53, 290–300 (2014).
Bonnal, R.J. et al. Biogem: an effective tool-based approach for scaling up open source software development in bioinformatics. Bioinformatics 28, 1035–1037 (2012).
Rousseeuw, P.J. & Leroy, A.M. John Wiley & Sons. in Wiley Series in Probability and Mathematical Statistics Applied Probability and Statistics (Wiley, New York, 1987).
Bodega, B. et al. Remodeling of the chromatin structure of the facioscapulohumeral muscular dystrophy (FSHD) locus and upregulation of FSHD-related gene 1 (FRG1) expression during human myogenic differentiation. BMC Biol. 7, 41 (2009).
Acknowledgements
We thank C. Cheroni for support in statistical analysis; M. Moro and M.C. Crosti for technical assistance with cell sorting; S. Biffo, D. Gabellini, P. Della Bona and A. Lanzavecchia for discussions and critical revision of the manuscript; B.J. Haas and A. Dobin for help with the integration of genome-guided Trinity with STAR aligner; the Istituto Nazionale Genetica Molecolare Bioinformatics Facility for support; and the Google Summer of Code Project for supporting C. Wheeler in the development of a plug-in used here for the open-source bioinformatics library BioRuby that adds support for the multiple-alignment format (https://github.com/csw/bioruby-maf). Supported by Il Consiglio Nazionale delle Ricerche–Il Ministero dell'Istuzione dell'Universita e della Ricerca (EPIGEN), Fondazione Cariplo (2013-0955), the Associazione Italiana per la Ricerca sul Cancro (IG2013-ID14596), the European Research Council (269022 to S.A.; 617978 to M.P.) and Fondazione Romeo ed Enrica Invernizzi.
Author information
Authors and Affiliations
Contributions
V.R., A.A. and R.J.P.B. set up all the bioinformatics pipelines, performed the bioinformatics analyses and contributed to the preparation of the manuscript; G.R. and I.P. designed and performed the main experiments, analyzed the data and contributed to the preparation of the manuscript; S.C., P.G., E.P., E.S. and B.B. performed experiments and analyzed the data; M.M., R.D.F. and J.G. discussed results, provided advice and commented on the manuscript; S.A. and M.P. designed the study, supervised research and wrote the manuscript; and all authors discussed and interpreted the results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Distribution and expression of lincRNAs in primary human lymphocytes subsets.
(a) Bar plots of expressed genes across a panel of 13 lymphocyte subsets. Average expression (± sdev) of at least four samples for each subset is reported
(b) Stacked bar plots of expressed genes percentages according to their biotype (protein coding, lincRNAs, pseudogenes, non-coding genes and other) across the analyzed human lymphocyte subsets
(c) Distribution of novel (striped) and previously annotated (black) lincRNAs in all human chromosomes
(d) Distribution of expressed novel (striped) and previously annotated (black) lincRNAs across the analyzed human lymphocyte subsets.
(e) Boxplots of gene expression values of lincRNA (blue) and protein coding genes (red) on either the whole dataset (global expression) or on a dataset filtered according to the specificity score (specific expression, Maximal JS score > 0.4)
(f) The density distribution of JS score for cell-specific receptor genes (black line) was fitted to a log-normal distribution (dotted red line). In order to derive a threshold for the cell-specificity score, we calculated the JS score value corresponding to one standard deviation away from the mean value of the fitted distribution (0.27). As a reference, the JS density distribution for the metabolic genes is reported (green line)
(g) Density distributions of maximal expression values of lincRNAs (blue area plot) and protein coding genes (red line), divided according to cellular specificity (maximal JS score < 0.4 or JS score > 0.4)
Supplementary Figure 2 Specificity of lincRNAs and protein-coding genes in primary human lymphocytes subsets.
(a) Silhouette scores (y-axis) are reported as a function of K (x-axis), the number of clusters used to partition the gene expression dataset of lincRNA genes. The average Silhouette value was calculated by taking the average of each clusters's average Si. In the graph Si data are reported for lincRNAs genes, for which the highest Si value (implying better clustering of the data) is 15
(b) Specificity of lincRNAs and protein coding genes (FPKM >1) by K-Means clustering across 13 human lymphocyte populations. Colour intensity represents the Z-score log2-normalized raw FPKM counts estimated by Cufflinks
Supplementary Figure 3 LincRNA signatures in a differentiation time course.
CD4+ naïve, TH1, TH2 and TH17 signature lincRNAs trends in CD4+ naïve T cells differentiated in TH0 conditions. RNA was collected at different time points during CD4+ naïve T cells differentiation and RNA-seq experiments were performed. Thin lines represent the trends of each signature lincRNA. Bold lines represent the average trend of all signature lincRNAs for each subset. Data are represented as a log2 normalized ratio between each time point and the relative time 0.
Supplementary Figure 4 Regulation of MAF transcription by linc-MAF-4.
(a) Expression levels (FPKM) of linc-MAF-4 and its neighboring protein coding genes DYNLRB2 and CDYL2 in CD4+ T cell subsets
(b) Expression of TBX21 an GATA3 in activated CD4+ naïve T cells differentiated in TH1 or TH2 polarizing conditions assessed at different time points by RT-qPCR (average of four independent experiments ± SEM)
(c) Expression of linc-MAF-4 and MAF assessed at different time points by RT-qPCR in activated CD4+ naïve T cells differentiated in TH1, TH2 and TH0 polarizing conditions. Bar plot of the percentage of c-Maf positive cells determined by intracellular staining at different time points is also shown (average of four independent experiments ± SEM)
(d) CD4+ naïve T cells differentiated in TH17 polarizing conditions according to Kleinewietfeld et al. (Nature 2013; 496, 518). Upper panels: intracellular staining of IL-17 and CCR6 protein expression at day 8 of differentiation (data are representative of four independent experiments) Lower panels: linc-MAF-4, MAF, RORC and IL17 transcript levels assessed at different time points by RT-qPCR (average of four independent experiments ± SEM)
(e) Test of linc-MAF-4 siRNAs in CD4+ naïve T cells. Four siRNA sequences were transfected independently in activated CD4+ naïve T cells and linc-MAF-4, MAF, GATA3 and IL4 transcript levels were assessed by RT-qPCR at day 3 post-transfection and activation (average of five independent experiments ± SEM)
(f) Intracellular staining of c-Maf and GATA-3 in naive CD4+ T cells stimulated with anti-CD3 and anti-CD28 and transfected with a control siRNA or linc-MAF-4 siRNA assessed at day 4 post-transfection and activation. Data are representative of five independent experiments
Supplementary Figure 5 Chromosome-conformation capture on in vitro–differentiated CD4+ TH1 cells.
(a) 2.5% agarose gel of the experimental triplicate used for 3C followed by BAC controls amplified with different primers that span the region between linc-MAF-4 and MAF
(b) Sequencing results with pertaining electropherograms and BLAST alignments for M1-L7 and M1-L12 amplicons
(c) Validation of anti-LSD1 and EZH2 antibodies used in RIP assay. LSD1 and EZH2 immunoprecipitates specifically retrieve HOTAIR RNA in HeLa cells as shown by Tsai et al. Science 329, 689 (2010). RNU2.1 and a region upstream the TSS of linc-MAF-4 were used as negative controls
(d) ChIP-qPCR analysis of EZH2 and H3K27me3 at MYOD1 locus, of H3K27me3 at a control region within the chromatin loop and of LSD1 at beta-actin locus in activated CD4+ naïve T cells transfected with linc-MAF-4 siRNA (black) or ctrl siRNA (white) (average of at least three independent experiments ± SEM)
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5, Supplementary Tables 1 and 3, and Supplementary Note (PDF 4639 kb)
Supplementary Table 2
GSEA gene lists CD4+ TH1 and TH2 specific genes (XLSX 14 kb)
Rights and permissions
About this article
Cite this article
Ranzani, V., Rossetti, G., Panzeri, I. et al. The long intergenic noncoding RNA landscape of human lymphocytes highlights the regulation of T cell differentiation by linc-MAF-4. Nat Immunol 16, 318–325 (2015). https://doi.org/10.1038/ni.3093
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3093
This article is cited by
-
Multi-omics analysis identifies RFX7 targets involved in tumor suppression and neuronal processes
Cell Death Discovery (2023)
-
Non-Coding RNA-Mediated Gene Regulation in Cardiovascular Disorders: Current Insights and Future Directions
Journal of Cardiovascular Translational Research (2023)
-
Identifying common transcriptome signatures of cancer by interpreting deep learning models
Genome Biology (2022)
-
Long Intergenic Noncoding RNAs Affect Biological Pathways Underlying Autoimmune and Neurodegenerative Disorders
Molecular Neurobiology (2022)
-
Emerging Role of LncRNAs in Autoimmune Lupus
Inflammation (2022)