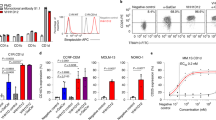
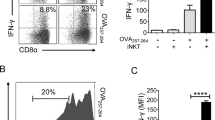
Abstract
Type I natural killer T cells (NKT cells) are characterized by an invariant variable region 14–joining region 18 (Vα14-Jα18) T cell antigen receptor (TCR) α-chain and recognition of the glycolipid α-galactosylceramide (α-GalCer) restricted to the antigen-presenting molecule CD1d. Here we describe a population of α-GalCer-reactive NKT cells that expressed a canonical Vα10-Jα50 TCR α-chain, which showed a preference for α-glucosylceramide (α-GlcCer) and bacterial α-glucuronic acid–containing glycolipid antigens. Structurally, despite very limited TCRα sequence identity, the Vα10 TCR–CD1d–α-GlcCer complex had a docking mode similar to that of type I TCR–CD1d–α-GalCer complexes, although differences at the antigen-binding interface accounted for the altered antigen specificity. Our findings provide new insight into the structural basis and evolution of glycolipid antigen recognition and have notable implications for the scope and immunological role of glycolipid-specific T cell responses.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Godfrey, D.I., MacDonald, H.R., Kronenberg, M., Smyth, M.J. & Van Kaer, L. NKT cells: what's in a name? Nat. Rev. Immunol. 4, 231–237 (2004).
Cardell, S. et al. CD1-restricted CD4+ T cells in major histocompatibility complex class II-deficient mice. J. Exp. Med. 182, 993–1004 (1995).
Park, S.H. et al. The mouse CD1d-restricted repertoire is dominated by a few autoreactive T cell receptor families. J. Exp. Med. 193, 893–904 (2001).
Blomqvist, M. et al. Multiple tissue-specific isoforms of sulfatide activate CD1d-restricted type II NKT cells. Eur. J. Immunol. 39, 1726–1735 (2009).
Cui, J.Q. et al. Requirement for Vα14 NKT cells in Il-12-mediated rejection of tumors. Science 278, 1623–1626 (1997).
Renukaradhya, G.J. et al. Type I NKT cells protect (and type II NKT cells suppress) the host's innate antitumor immune response to a B cell lymphoma. Blood 111, 5637–5645 (2008).
Kim, J.H., Choi, E.Y. & Chung, D.H. Donor bone marrow type II (non-Vα14Jα18 CD1d-restricted) NKT cells suppress graft-versus-host disease by producing IFN-γ and IL-4. J. Immunol. 179, 6579–6587 (2007).
Terabe, M. et al. A nonclassical non- Vα14Jα18 CD1d-restricted (type II) NKT cell is sufficient for down-regulation of tumor immunosurveillance. J. Exp. Med. 202, 1627–1633 (2005).
Berzins, S.P., Smyth, M.J. & Godfrey, D.I. Working with NKT cells—pitfalls and practicalities. Curr. Opin. Immunol. 17, 448–454 (2005).
Gadola, S.D. et al. Structure and binding kinetics of three different human CD1d-α-galactosylceramide-specific T cell receptors. J. Exp. Med. 203, 699–710 (2006).
Gadola, S.D., Dulphy, N., Salio, M. & Cerundolo, V. V alpha 24-J alpha Q-independent, CD1d-restricted recognition of alpha-galactosylceramide by human CD4+ and CD8αβ+ T lymphocytes. J. Immunol. 168, 5514–5520 (2002).
Brigl, M. et al. Conserved and heterogeneous lipid antigen specificities of CD1d-restricted NKT cell receptors. J. Immunol. 176, 3625–3634 (2006).
Godfrey, D.I. et al. Antigen recognition by CD1d-restricted NKT T cell receptors. Semin. Immunol. 22, 61–67 (2010).
Pellicci, D.G. et al. Differential recognition of CD1d-alpha-galactosyl ceramide by the Vβ8.2 and Vβ7 semi-invariant NKT T cell receptors. Immunity 31, 47–59 (2009).
Borg, N.A. et al. CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor. Nature 448, 44–49 (2007).
Li, Y. et al. The Vα14 invariant natural killer T cell TCR forces microbial glycolipids and CD1d into a conserved binding mode. J. Exp. Med. 207, 2383–2393 (2010).
Wun, K.S. et al. A molecular basis for the exquisite CD1d-restricted Ag-specificity and functional responses of NKT cells. Immunity 34, 327–339 (2011).
Mallevaey, T. et al. The molecular basis of NKT cell autoreactivity and recognition of self-CD1d. Immunity 34, 315–326 (2011).
Wun, K.S. et al. A minimal binding footprint on CD1d-glycolipid is a basis for selection of the unique human NKT TCR. J. Exp. Med. 205, 939–949 (2008).
Scott-Browne, J.P. et al. Germline-encoded recognition of diverse glycolipids by natural killer T cells. Nat. Immunol. 8, 1105–1113 (2007).
Mallevaey, T. et al. T cell receptor CDR2β and CDR3β loops collaborate functionally to shape the iNKT cell repertoire. Immunity 31, 60–71 (2009).
Florence, W.C. et al. Adaptability of the semi-invariant natural killer T-cell receptor towards structurally diverse CD1d-restricted ligands. EMBO J. 28, 3579–3590 (2009).
Stenstrom, M., Skold, M., Andersson, A. & Cardell, S.L. Natural killer T-cell populations in C57BL/6 and NK1.1 congenic BALB.NK mice-a novel thymic subset defined in BALB.NK mice. Immunology 114, 336–345 (2005).
Yu, K.O. et al. Modulation of CD1d-restricted NKT cell responses by using N-acyl variants of α-galactosylceramides. Proc. Natl. Acad. Sci. USA 102, 3383–3388 (2005).
Jervis, P.J. et al. Synthesis and biological activity of α-glucosyl C24:0 and C20:2 ceramides. Bioorg. Med. Chem. Lett. 20, 3475–3478 (2010).
Wolucka, B.A., McNeil, M.R., Kalbe, L., Cocito, C. & Brennan, P.J. Isolation and characterization of a novel glucuronosyl diacylglycerol from Mycobacterium smegmatis. Biochim. Biophys. Acta 1170, 131–136 (1993).
Kawano, T. et al. Cd1d-restricted and TCR-mediated activation of Vα14 NKT cells by glycosylceramides. Science 278, 1626–1629 (1997).
Kinjo, Y. et al. Recognition of bacterial glycosphingolipids by natural killer T cells. Nature 434, 520–525 (2005).
Mattner, J. et al. Exogenous and endogenous glycolipid antigens activate NKT cells during microbial infections. Nature 434, 525–529 (2005).
Sriram, V., Du, W., Gervay-Hague, J. & Brutkiewicz, R.R. Cell wall glycosphingolipids of Sphingomonas paucimobilis are CD1d-specific ligands for NKT cells. Eur. J. Immunol. 35, 1692–1701 (2005).
Zhou, D. et al. Lysosomal glycosphingolipid recognition by NKT cells. Science 306, 1786–1789 (2004).
Dias, B.R. et al. Identification of iGb3 and iGb4 in melanoma B16F10-Nex2 cells and the iNKT cell-mediated antitumor effect of dendritic cells primed with iGb3. Mol. Cancer 8, 116–129 (2009).
Kjer-Nielsen, L. et al. A structural basis for selection and cross-species reactivity of the semi-invariant NKT cell receptor in CD1d/glycolipid recognition. J. Exp. Med. 203, 661–673 (2006).
Feng, D., Bond, C.J., Ely, L.K., Maynard, J. & Garcia, K.C. Structural evidence for a germline-encoded T cell receptor-major histocompatibility complex interaction 'codon'. Nat. Immunol. 8, 975–983 (2007).
Dai, S. et al. Crossreactive T Cells spotlight the germline rules for αβ T cell-receptor interactions with MHC molecules. Immunity 28, 324–334 (2008).
Godfrey, D.I., Rossjohn, J. & McCluskey, J. The fidelity, occasional promiscuity, and versatility of T cell receptor recognition. Immunity 28, 304–314 (2008).
Smiley, S.T., Kaplan, M.H., Grusby, M.J. & Immunoglobulin, E. Production in the absence of interleukin-4-secreting Cd1-dependent cells. Science 275, 977–979 (1997).
Sonoda, K.H., Exley, M., Snapper, S., Balk, S.P. & Stein-Streilein, J. CD1-reactive natural killer T cells are required for development of systemic tolerance through an immune-privileged site. J. Exp. Med. 190, 1215–1226 (1999).
Crowe, N.Y. et al. Differential antitumor immunity mediated by NKT cell subsets in vivo. J. Exp. Med. 202, 1279–1288 (2005).
Long, X. et al. Synthesis and evaluation of stimulatory properties of Sphingomonadaceae glycolipids. Nat. Chem. Biol. 3, 559–564 (2007).
Casanova, J.L., Romero, P., Widmann, C., Kourilsky, P. & Maryanski, J.L. T cell receptor genes in a series of class I major histocompatibility complex-restricted cytotoxic T lymphocyte clones specific for a Plasmodium berghei nonapeptide: implications for T cell allelic exclusion and antigen-specific repertoire. J. Exp. Med. 174, 1371–1383 (1991).
Day, E.B. et al. The context of epitope presentation can influence functional quality of recalled influenza A virus-specific memory CD8+ T cells. J. Immunol. 179, 2187–2194 (2007).
Kedzierska, K., Turner, S.J. & Doherty, P.C. Conserved T cell receptor usage in primary and recall responses to an immunodominant influenza virus nucleoprotein epitope. Proc. Natl. Acad. Sci. USA 101, 4942–4947 (2004).
Lefranc, M.P. et al. IMGT, the International ImMunoGeneTics database. Nucleic Acids Res. 26, 297–303 (1998).
Garboczi, D.N. et al. Assembly, specific binding, and crystallization of a human TCR-αβ with an antigenic Tax peptide from human T lymphotropic virus type 1 and the class I MHC molecule HLA-A2. J. Immunol. 157, 5403–5410 (1996).
Holst, J. et al. Generation of T-cell receptor retrogenic mice. Nat. Protoc. 1, 406–417 (2006).
Leslie, A.G.W. Joint CCP4 + ESF-EAMCB Newsletter on Protein Crystallography. Recent changes to the MOSFLM package for processing film and image plate data 26 (1992).
CCP4 (Collaborative Computational Project, 4). The CCP4 suite: Programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Otwinoski, Z. & Minor, W. Processing of x-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Zajonc, D.M. et al. Structure and function of a potent agonist for the semi-invariant natural killer T cell receptor. Nat. Immunol. 6, 810–818 (2005).
Acknowledgements
We thank M. Taniguchi (Chiba University Graduate School of Medicine) for Jα18−/− mice; M. Kronenberg (La Jolla Institute for Allergy and Immunology) for the baculovirus-based CD1d expression system; P. Savage (Brigham Young University) for α-GalCer (C24:1 PBS-44 analog); the Australian Synchrotron staff at the MX1 and MX2 beamlines of the Australian synchrotron for assistance with data collection; S. Mattarollo, S. Doak, S. Berzins and A. Denton for discussions and assistance with some experiments; K. Field, N. Sanders and M. Reitsma for assistance with flow cytometry; and M. Stirling and the staff of the Peter MacCallum Cancer Centre Animal House and D. Maksel from the Protein Crystallography Unit at Monash University for technical assistance. Supported by the Cancer Council of Victoria, the National Health and Medical Research Council of Australia (A.P.U., L.C.S., M.J.S. and D.I.G.), the Australian Research Council (D.I.G., O.P. and J.R.), the Cancer Research Institute (G.C.) and the US National Institutes of Health (AI45889 to S.A.P.).
Author information
Authors and Affiliations
Contributions
A.P.U. identified and carried out cellular and molecular characterization of Vα10 NKT cells and produced protein complexes for crystallographic studies; O.P. solved the crystal structures and did structural analysis; G.C. and K.K. carried out studies of glycolipid specificity and function; L.C.S. did surface plasmon resonance studies; D.G.P., E.B.D., L.K.-N., J.P.V., S.J.T., G.S.B., B.C., A.G.B., S.J.W., P.I., S.A.P., J.M., M.J.S., J.R. and D.I.G. provided intellectual input and key reagents and assisted with experimental design and interpretation and writing of the manuscript; and M.J.S., J.R. and D.I.G. led the investigation together and devised the project and contributed equally to this work.
Corresponding authors
Ethics declarations
Competing interests
S.A.P. has received payment as a consultant for Vaccinex for work related to the development of therapeutics based on glycolipids pretreated with CD1d.
Supplementary information
Supplementary Text and Figures
Supplementary Figure 1 and Supplementary Tables 1–2 (PDF 751 kb)
Rights and permissions
About this article
Cite this article
Uldrich, A., Patel, O., Cameron, G. et al. A semi-invariant Vα10+ T cell antigen receptor defines a population of natural killer T cells with distinct glycolipid antigen–recognition properties. Nat Immunol 12, 616–623 (2011). https://doi.org/10.1038/ni.2051
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2051
This article is cited by
-
Current Developments in the Preclinical and Clinical use of Natural Killer T cells
BioDrugs (2023)
-
Role of NKT cells in cancer immunotherapy—from bench to bed
Medical Oncology (2022)
-
Thymic development of unconventional T cells: how NKT cells, MAIT cells and γδ T cells emerge
Nature Reviews Immunology (2020)
-
Single-cell RNA sequencing identifies shared differentiation paths of mouse thymic innate T cells
Nature Communications (2020)
-
Distinct CD1d docking strategies exhibited by diverse Type II NKT cell receptors
Nature Communications (2019)