Abstract
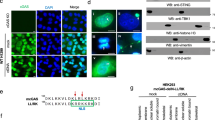
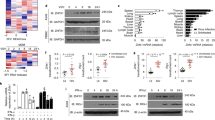
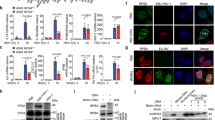
Intracellular nucleic acid sensors detect microbial RNA and DNA and trigger the production of type I interferon. However, the cytosolic nucleic acid–sensing system remains to be fully identified. Here we show that the cytosolic nucleic acid–binding protein LRRFIP1 contributed to the production of interferon-β (IFN-β) induced by vesicular stomatitis virus (VSV) and Listeria monocytogenes in macrophages. LRRFIP1 bound exogenous nucleic acids and increased the expression of IFN-β induced by both double-stranded RNA and double-stranded DNA. LRRFIP1 interacted with β-catenin and promoted the activation of β-catenin, which increased IFN-β expression by binding to the C-terminal domain of the transcription factor IRF3 and recruiting the acetyltransferase p300 to the IFN-β enhanceosome via IRF3. Therefore, LRRFIP1 and its downstream partner β-catenin constitute another coactivator pathway for IRF3-mediated production of type I interferon.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Takeda, K., Kaisho, T. & Akira, S. Toll-like receptors. Annu. Rev. Immunol. 21, 335–376 (2003).
O'Neill, L.A. & Bowie, A.G. The family of five: TIR-domain-containing adaptors in Toll-like receptor signalling. Nat. Rev. Immunol. 7, 353–364 (2007).
Alexopoulou, L., Holt, A.C., Medzhitov, R. & Flavell, R.A. Recognition of double-stranded RNA and activation of NF-κB by Toll-like receptor 3. Nature 413, 732–738 (2001).
Yoneyama, M. et al. The RNA helicase RIG-I has an essential function in double-stranded RNA-induced innate antiviral responses. Nat. Immunol. 5, 730–737 (2004).
Kato, H. et al. Differential roles of MDA5 and RIG-I helicases in the recognition of RNA viruses. Nature 441, 101–105 (2006).
Hiscott, J., Lin, R., Nakhaei, P. & Paz, S. MasterCARD: a priceless link to innate immunity. Trends Mol. Med. 12, 53–56 (2006).
Pichlmair, A. et al. RIG-I-mediated antiviral responses to single-stranded RNA bearing 5′-phosphates. Science 314, 997–1001 (2006).
Kato, H. et al. Length-dependent recognition of double-stranded ribonucleic acids by retinoic acid-inducible gene-I and melanoma differentiation-associated gene 5. J. Exp. Med. 205, 1601–1610 (2008).
Ishii, K.J. et al. A Toll-like receptor-independent antiviral response induced by double-stranded B-form DNA. Nat. Immunol. 7, 40–48 (2006).
Takaoka, A. et al. DAI (DLM-1/ZBP1) is a cytosolic DNA sensor and an activator of innate immune response. Nature 448, 501–505 (2007).
Burckstummer, T. et al. An orthogonal proteomic-genomic screen identifies AIM2 as a cytoplasmic DNA sensor for the inflammasome. Nat. Immunol. 10, 266–272 (2009).
Hornung, V. et al. AIM2 recognizes cytosolic dsDNA and forms a caspase-1-activating inflammasome with ASC. Nature 458, 514–518 (2009).
Fernandes-Alnemri, T., Yu, J.W., Datta, P., Wu, J. & Alnemri, E.S. Aim2 activates the inflammasome and cell death in response to cytoplasmic DNA. Nature 458, 509–513 (2009).
Roberts, T.L. et al. HIN-200 proteins regulate caspase activation in response to foreign cytoplasmic DNA. Science 323, 1057–1060 (2009).
Ishii, K.J. et al. TANK-binding kinase-1 delineates innate and adaptive immune responses to DNA vaccines. Nature 451, 725–729 (2008).
Wang, Z. et al. Regulation of innate immune responses by DAI (DLM-1/ZBP1) and other DNA-sensing molecules. Proc. Natl. Acad. Sci. USA 105, 5477–5482 (2008).
Fernandes-Alnemri, T. et al. The AIM2 inflammasome is critical for innate immunity to Francisella tularensis. Nat. Immunol. (2010).
Rathinam, V.A. et al. The AIM2 inflammasome is essential for host defense against cytosolic bacteria and DNA viruses. Nat. Immunol. (2010).
Chiu, Y.H., Macmillan, J.B. & Chen, Z.J. RNA polymerase III detects cytosolic DNA and induces type I interferons through the RIG-I pathway. Cell 138, 576–591 (2009).
Ablasser, A. et al. RIG-I-dependent sensing of poly(dA:dT) through the induction of an RNA polymerase III-transcribed RNA intermediate. Nat. Immunol. 10, 1065–1072 (2009).
O'Neill, L.A. DNA makes RNA makes innate immunity. Cell 138, 428–430 (2009).
Sun, Q. et al. The specific and essential role of MAVS in antiviral innate immune responses. Immunity 24, 633–642 (2006).
Stetson, D.B. & Medzhitov, R. Recognition of cytosolic DNA activates an IRF3-dependent innate immune response. Immunity 24, 93–103 (2006).
Liu, Y.T. & Yin, H.L. Identification of the binding partners for flightless I, a novel protein bridging the leucine-rich repeat and the gelsolin superfamilies. J. Biol. Chem. 273, 7920–7927 (1998).
Wilson, S.A., Brown, E.C., Kingsman, A.J. & Kingsman, S.M. TRIP: a novel double stranded RNA binding protein which interacts with the leucine rich repeat of flightless I. Nucleic Acids Res. 26, 3460–3467 (1998).
Suriano, A.R. et al. GCF2/LRRFIP1 represses tumor necrosis factor α expression. Mol. Cell. Biol. 25, 9073–9081 (2005).
Thanos, D. & Maniatis, T. Virus induction of human IFN β gene expression requires the assembly of an enhanceosome. Cell 83, 1091–1100 (1995).
Falvo, J.V., Parekh, B.S., Lin, C.H., Fraenkel, E. & Maniatis, T. Assembly of a functional beta interferon enhanceosome is dependent on ATF-2-c-jun heterodimer orientation. Mol. Cell. Biol. 20, 4814–4825 (2000).
Wathelet, M.G. et al. Virus infection induces the assembly of coordinately activated transcription factors on the IFN-β enhancer in vivo. Mol. Cell 1, 507–518 (1998).
Stadeli, R., Hoffmans, R. & Basler, K. Transcription under the control of nuclear Arm/β-catenin. Curr. Biol. 16, R378–R385 (2006).
Lee, Y.H. & Stallcup, M.R. Interplay of Fli-I and FLAP1 for regulation of beta-catenin dependent transcription. Nucleic Acids Res. 34, 5052–5059 (2006).
Duan, Y. et al. β-Catenin activity negatively regulates bacteria-induced inflammation. Lab. Invest. 87, 613–624 (2007).
Mosimann, C., Hausmann, G. & Basler, K. β-catenin hits chromatin: regulation of Wnt target gene activation. Nat. Rev. Mol. Cell Biol. 10, 276–286 (2009).
Taniguchi, T., Ogasawara, K., Takaoka, A. & Tanaka, N. IRF family of transcription factors as regulators of host defense. Annu. Rev. Immunol. 19, 623–655 (2001).
Parekh, B.S. & Maniatis, T. Virus infection leads to localized hyperacetylation of histones H3 and H4 at the IFN-β promoter. Mol. Cell 3, 125–129 (1999).
Merika, M., Williams, A.J., Chen, G., Collins, T. & Thanos, D. Recruitment of CBP/p300 by the IFN-β enhanceosome is required for synergistic activation of transcription. Mol. Cell 1, 277–287 (1998).
Kumar, H. et al. Essential role of IPS-1 in innate immune responses against RNA viruses. J. Exp. Med. 203, 1795–1803 (2006).
Saito, T. et al. Regulation of innate antiviral defenses through a shared repressor domain in RIG-I and LGP2. Proc. Natl. Acad. Sci. USA 104, 582–587 (2007).
Karpova, A.Y., Ronco, L.V. & Howley, P.M. Functional characterization of interferon regulatory factor 3a (IRF-3a), an alternative splice isoform of IRF-3. Mol. Cell. Biol. 21, 4169–4176 (2001).
Kumar, A. et al. Active beta-catenin signaling is an inhibitory pathway for human immunodeficiency virus replication in peripheral blood mononuclear cells. J. Virol. 82, 2813–2820 (2008).
Fang, D. et al. Phosphorylation of β-catenin by AKT promotes β-catenin transcriptional activity. J. Biol. Chem. 282, 11221–11229 (2007).
Shapira, S.D. et al. A physical and regulatory map of host-influenza interactions reveals pathways in H1N1 infection. Cell 139, 1255–1267 (2009).
Cobas, M. et al. β-catenin is dispensable for hematopoiesis and lymphopoiesis. J. Exp. Med. 199, 221–229 (2004).
Wang, C. et al. The E3 ubiquitin ligase Nrdp1 'preferentially' promotes TLR-mediated production of type I interferon. Nat. Immunol. 10, 744–752 (2009).
An, H. et al. SHP-2 phosphatase negatively regulates the TRIF adaptor protein-dependent type I interferon and proinflammatory cytokine production. Immunity 25, 919–928 (2006).
Hou, J. et al. MicroRNA-146a feedback inhibits RIG-I-dependent Type I IFN production in macrophages by targeting TRAF6, IRAK1, and IRAK2. J. Immunol. 183, 2150–2158 (2009).
Yun, M.S., Kim, S.E., Jeon, S.H., Lee, J.S. & Choi, K.Y. Both ERK and Wnt/β-catenin pathways are involved in Wnt3a-induced proliferation. J. Cell Sci. 118, 313–322 (2005).
Leung, T.H., Hoffmann, A. & Baltimore, D. One nucleotide in a κB site can determine cofactor specificity for NF-κB dimers. Cell 118, 453–464 (2004).
An, H. et al. Phosphatase SHP-1 promotes TLR- and RIG-I-activated production of type I interferon by inhibiting the kinase IRAK1. Nat. Immunol. 9, 542–550 (2008).
Mokrani, H., Sharaf el, D.O., Mansuroglu, Z. & Bonnefoy, E. Binding of YY1 to the proximal region of the murine β interferon promoter is essential to allow CBP recruitment and K8H4/K14H3 acetylation on the promoter region after virus infection. Mol. Cell. Biol. 26, 8551–8561 (2006).
Acknowledgements
We thank X. Ma and T. Zuo for technical assistance; W. Pan (Second Military Medical University) for VSV; and H. Shen (University of Pennsylvania School of Medicine) for L. monocytogenes. Supported by the National Natural Science Foundation of China (30825036 and 30721091), the National Key Basic Research Program of China (2007CB512403), the National Grand Program on Key Infectious Disease (2009ZX10004-309) and the Program for New Century Excellent Talents in University of the Ministry of Education of China (NCET-07-0143).
Author information
Authors and Affiliations
Contributions
P.Y., H.A., X.L., M.W., Y.Z. and Y.R. did the experiments; and X.C. and H.A. designed the study and wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11 and Supplementary Table 1 (PDF 504 kb)
Rights and permissions
About this article
Cite this article
Yang, P., An, H., Liu, X. et al. The cytosolic nucleic acid sensor LRRFIP1 mediates the production of type I interferon via a β-catenin-dependent pathway. Nat Immunol 11, 487–494 (2010). https://doi.org/10.1038/ni.1876
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.1876
This article is cited by
-
Chromatin-Associated Molecular Patterns (CAMPs) in sepsis
Cell Death & Disease (2022)
-
Mass spectrometry-based draft of the mouse proteome
Nature Methods (2022)
-
Cytosolic and nuclear recognition of virus and viral evasion
Molecular Biomedicine (2021)
-
Cytosolic DNA sensing by cGAS: regulation, function, and human diseases
Signal Transduction and Targeted Therapy (2021)
-
Targeting nuclear acid-mediated immunity in cancer immune checkpoint inhibitor therapies
Signal Transduction and Targeted Therapy (2020)