Abstract
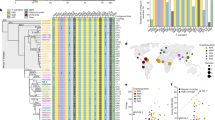
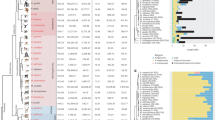
We report the sequences of 1,244 human Y chromosomes randomly ascertained from 26 worldwide populations by the 1000 Genomes Project. We discovered more than 65,000 variants, including single-nucleotide variants, multiple-nucleotide variants, insertions and deletions, short tandem repeats, and copy number variants. Of these, copy number variants contribute the greatest predicted functional impact. We constructed a calibrated phylogenetic tree on the basis of binary single-nucleotide variants and projected the more complex variants onto it, estimating the number of mutations for each class. Our phylogeny shows bursts of extreme expansion in male numbers that have occurred independently among each of the five continental superpopulations examined, at times of known migrations and technological innovations.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Jobling, M.A. & Tyler-Smith, C. The human Y chromosome: an evolutionary marker comes of age. Nat. Rev. Genet. 4, 598–612 (2003).
Wei, W. et al. A calibrated human Y-chromosomal phylogeny based on resequencing. Genome Res. 23, 388–395 (2013).
Poznik, G.D. et al. Sequencing Y chromosomes resolves discrepancy in time to common ancestor of males versus females. Science 341, 562–565 (2013).
Wilson Sayres, M.A., Lohmueller, K.E. & Nielsen, R. Natural selection reduced diversity on human Y chromosomes. PLoS Genet. 10, e1004064 (2014).
Karmin, M. et al. A recent bottleneck of Y chromosome diversity coincides with a global change in culture. Genome Res. 25, 459–466 (2015).
Batini, C. et al. Large-scale recent expansion of European patrilineages shown by population resequencing. Nat. Commun. 6, 7152 (2015).
Sikora, M.J., Colonna, V., Xue, Y. & Tyler-Smith, C. Modeling the contrasting Neolithic male lineage expansions in Europe and Africa. Investig. Genet. 4, 25 (2013).
Helgason, A. et al. The Y-chromosome point mutation rate in humans. Nat. Genet. 47, 453–457 (2015).
Fu, Q. et al. Genome sequence of a 45,000-year-old modern human from western Siberia. Nature 514, 445–449 (2014).
Balanovsky, O. et al. Deep phylogenetic analysis of haplogroup G1 provides estimates of SNP and STR mutation rates on the human Y-chromosome and reveals migrations of Iranic speakers. PLoS One 10, e0122968 (2015).
1000 Genomes Project Consortium. A global reference for human genetic variation. Nature 526, 68–74 (2015).
Handsaker, R.E. et al. Large multiallelic copy number variations in humans. Nat. Genet. 47, 296–303 (2015).
Bellos, E., Johnson, M.R. & Coin, L.J.M. cnvHiTSeq: integrative models for high-resolution copy number variation detection and genotyping using population sequencing data. Genome Biol. 13, R120 (2012).
Hammer, M.F. et al. Out of Africa and back again: nested cladistic analysis of human Y chromosome variation. Mol. Biol. Evol. 15, 427–441 (1998).
Groucutt, H.S. et al. Rethinking the dispersal of Homo sapiens out of Africa. Evol. Anthropol. 24, 149–164 (2015).
Fu, Q. et al. An early modern human from Romania with a recent Neanderthal ancestor. Nature 524, 216–219 (2015).
Zhang, F., Gu, W., Hurles, M.E. & Lupski, J.R. Copy number variation in human health, disease, and evolution. Annu. Rev. Genomics Hum. Genet. 10, 451–481 (2009).
Willems, T. et al. Population-scale sequencing data enables precise estimates of Y-STR mutation rates. Am. J. Hum. Genet. http://dx.doi.org/10.1016/j.ajhg.2016.04.001 (2016).
Kumar, P., Henikoff, S. & Ng, P.C. Predicting the effects of coding non-synonymous variants on protein function using the SIFT algorithm. Nat. Protoc. 4, 1073–1081 (2009).
Sudmant, P.H. et al. Global diversity, population stratification, and selection of human copy-number variation. Science 349, aab3761 (2015).
Raghavan, M. et al. Genomic evidence for the Pleistocene and recent population history of Native Americans. Science 349, aab3884 (2015).
de Filippo, C., Bostoen, K., Stoneking, M. & Pakendorf, B. Bringing together linguistic and genetic evidence to test the Bantu expansion. Proc. R. Soc. B Biol. Sci. 279, 3256–3263 (2012).
Jobling, M.A., Hollox, E., Hurles, M., Kivisild, T. & Tyler-Smith, C. Human Evolutionary Genetics 2nd edn (Garland Science, 2014).
Allentoft, M.E. et al. Population genomics of Bronze Age Eurasia. Nature 522, 167–172 (2015).
Haak, W. et al. Massive migration from the steppe was a source for Indo-European languages in Europe. Nature 522, 207–211 (2015).
Harding, A.F. European Societies in the Bronze Age (Cambridge University Press, 2000).
Bryant, E.F. & Patton, L.L. The Indo-Aryan Controversy: Evidence and Inference in Indian History (Routledge, 2005).
Xue, Y. et al. Human Y chromosome base-substitution mutation rate measured by direct sequencing in a deep-rooting pedigree. Curr. Biol. 19, 1453–1457 (2009).
Betzig, L. Means, variances, and ranges in reproductive success: comparative evidence. Evol. Hum. Behav. 33, 309–317 (2012).
Zerjal, T. et al. The genetic legacy of the Mongols. Am. J. Hum. Genet. 72, 717–721 (2003).
Balaresque, P. et al. Y-chromosome descent clusters and male differential reproductive success: young lineage expansions dominate Asian pastoral nomadic populations. Eur. J. Hum. Genet. 23, 1413–1422 (2015).
Wickham, H. ggplot2: Elegant Graphics for Data Analysis (Springer, 2009).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Garrison, E. & Marth, G. Haplotype-based variant detection from short-read sequencing. arXiv http://arxiv.org/abs/1207.3907 (2012).
Rimmer, A. et al. Integrating mapping-, assembly- and haplotype-based approaches for calling variants in clinical sequencing applications. Nat. Genet. 46, 912–918 (2014).
Iqbal, Z., Caccamo, M., Turner, I., Flicek, P. & McVean, G. De novo assembly and genotyping of variants using colored de Bruijn graphs. Nat. Genet. 44, 226–232 (2012).
McKenna, A. et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 20, 1297–1303 (2010).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Pique-Regi, R. et al. Sparse representation and Bayesian detection of genome copy number alterations from microarray data. Bioinformatics 24, 309–318 (2008).
Pique-Regi, R., Cáceres, A. & González, J.R. R-Gada: a fast and flexible pipeline for copy number analysis in association studies. BMC Bioinformatics 11, 380 (2010).
Conrad, D.F. et al. Origins and functional impact of copy number variation in the human genome. Nature 464, 704–712 (2010).
Perry, G.H. et al. Copy number variation and evolution in humans and chimpanzees. Genome Res. 18, 1698–1710 (2008).
Polley, S. et al. Evolution of the rapidly mutating human salivary agglutinin gene (DMBT1) and population subsistence strategy. Proc. Natl. Acad. Sci. USA 112, 5105–5110 (2015).
Carpenter, D. et al. Obesity, starch digestion and amylase: association between copy number variants at human salivary (AMY1) and pancreatic (AMY2) amylase genes. Hum. Mol. Genet. 24, 3472–3480 (2015).
Verma, R.S. & Babu, A. Human Chromosomes: Principles & Techniques 2nd edn. (McGraw-Hill, 1995).
Gribble, S.M. et al. Massively parallel sequencing reveals the complex structure of an irradiated human chromosome on a mouse background in the Tc1 model of Down syndrome. PLoS One 8, e60482 (2013).
Felsenstein, J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Stamatakis, A. RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30, 1312–1313 (2014).
McLaren, W. et al. Deriving the consequences of genomic variants with the Ensembl API and SNP Effect Predictor. Bioinformatics 26, 2069–2070 (2010).
Kircher, M. et al. A general framework for estimating the relative pathogenicity of human genetic variants. Nat. Genet. 46, 310–315 (2014).
Adzhubei, I.A. et al. A method and server for predicting damaging missense mutations. Nat. Methods 7, 248–249 (2010).
van Oven, M. & Kayser, M. Updated comprehensive phylogenetic tree of global human mitochondrial DNA variation. Hum. Mutat. 30, E386–E394 (2009).
Danecek, P. et al. The variant call format and VCFtools. Bioinformatics 27, 2156–2158 (2011).
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013).
Kloss-Brandstätter, A. et al. HaploGrep: a fast and reliable algorithm for automatic classification of mitochondrial DNA haplogroups. Hum. Mutat. 32, 25–32 (2011).
Hudson, R.R. Generating samples under a Wright–Fisher neutral model of genetic variation. Bioinformatics 18, 337–338 (2002).
Lohmueller, K.E., Bustamante, C.D. & Clark, A.G. Methods for human demographic inference using haplotype patterns from genomewide single-nucleotide polymorphism data. Genetics 182, 217–231 (2009).
Lohmueller, K.E., Bustamante, C.D. & Clark, A.G. The effect of recent admixture on inference of ancient human population history. Genetics 185, 611–622 (2010).
Acknowledgements
We thank the 1000 Genomes Project sample donors for making this work possible, all Project members for their contributions, and A. Martin for ADMIXTURE results. The tree in Figure 2 was drawn using FigTree. G.D.P. was supported by the National Science Foundation (NSF) Graduate Research Fellowship under grant DGE-1147470 and by National Library of Medicine training grant LM-007033. Work at the Wellcome Trust Sanger Institute (Q.A., R.B., M.C., Y.C., S.L., A. Massaia, S.A. McCarthy, C.T.-S., Y.X., and F.Y.) was supported by Wellcome Trust grant 098051. F.L.M. was supported by National Institutes of Health (NIH) grant 1R01GM090087, by NSF grant DMS-1201234, and by a postdoctoral fellowship from the Stanford Center for Computational, Evolutionary and Human Genomics (CEHG). T.F.W. was supported by an AWS Education Grant, and the work of T.F.W., M.G., and Y.E. was supported in part by NIJ award 2014-DN-BX-K089. M.C. is supported by a Fundación Barrié Fellowship. H.S. and L. Coin are supported by Australian Research Council grants DP140103164 and FT110100972, respectively. M.G. was supported by a National Defense Science and Engineering Graduate Fellowship. G.R.S.R. was supported by the European Molecular Biology Laboratory and the Sanger Institute through an EBI–Sanger Postdoctoral Fellowship. X.Z.-B., P.F., D.R.Z., and L. Clarke were supported by Wellcome Trust grants 085532, 095908, and 104947 and by the European Molecular Biology Laboratory. P.A.U. was supported by SAP grant SP0#115016. C.L. was supported in part by NIH grant U41HG007497. Y.E. holds a Career Award at the Scientific Interface from the Burroughs Wellcome Fund. C.D.B. was supported by NIH grant 5R01HG003229-09.
Author information
Authors and Affiliations
Consortia
Contributions
G.D.P., Y.X., C.D.B., and C.T.-S. conceived and designed the project. R.B., S.L., and F.Y. generated FISH data. A. Malhotra, M.R., E.C., C.Z., and C.L. generated aCGH data. G.D.P., Y.X., F.L.M., T.F.W., A. Massaia, M.A.W.S., Q.A., S.A. McCarthy, A.N., S.K., Y.C., J.L.R.-F., M.C., H.S., M.G., R.D., G.R.S.R., T.W.F., E.G., A. Marcketta, D.M., X.Z.-B., G.R.A., S.A. McCarroll, P.F., P.A.U., L. Coin, D.R.Z., L. Clarke, A.A., Y.E., R.E.H., C.D.B., and C.T.-S. analyzed the data. G.D.P., Y.X., F.L.M., T.F.W., A. Massaia, M.A.W.S., Q.A., and C.T.-S. wrote the manuscript. All authors reviewed, revised, and provided feedback on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
G.D.P. and A.A. are employees of 23andMe. P.F. is a member of the Scientific Advisory Board (SAB) for Omicia, Inc. P.A.U. has consulted for and owns stock options of 23andMe. Y.E. is an SAB member of Identify Genomics, BigDataBio, and Solve, Inc. C.D.B. is on the SABs of AncestryDNA, BigDataBio, Etalon DX, Liberty Biosecurity, and Personalis. He is also a founder and SAB chair of IdentifyGenomics. None of these entities had a role in the design, execution, interpretation, or presentation of this study.
Additional information
A list of members and affiliations appears in the Supplementary Note.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–31, Supplementary Tables 1–19 and Supplementary Note. (PDF 16615 kb)
Supplementary Data
Supplementary Data on SNVs, CNVs, STRs, haplogroups, phylogenetic analyses, functional annotations, mtDNA analysis, and expansion analyses. (ZIP 6186 kb)
Rights and permissions
About this article
Cite this article
Poznik, G., Xue, Y., Mendez, F. et al. Punctuated bursts in human male demography inferred from 1,244 worldwide Y-chromosome sequences. Nat Genet 48, 593–599 (2016). https://doi.org/10.1038/ng.3559
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.3559
This article is cited by
-
Differentiated genomic footprints suggest isolation and long-distance migration of Hmong-Mien populations
BMC Biology (2024)
-
Multiple founding paternal lineages inferred from the newly-developed 639-plex Y-SNP panel suggested the complex admixture and migration history of Chinese people
Human Genomics (2023)
-
Parallel signatures of Mycobacterium tuberculosis and human Y-chromosome phylogeography support the Two Layer model of East Asian population history
Communications Biology (2023)
-
Genetic sexing of subadult skeletal remains
Scientific Reports (2023)
-
Genomic history of coastal societies from eastern South America
Nature Ecology & Evolution (2023)