Abstract
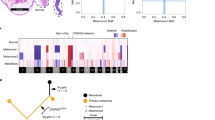
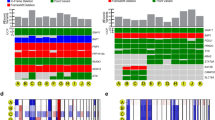
Gene expression profiles and chromosome 3 copy number divide uveal melanomas into two distinct classes correlating with prognosis1,2,3. Using exome sequencing, we identified recurrent somatic mutations in EIF1AX and SF3B1, specifically occurring in uveal melanomas with disomy 3, which rarely metastasize. Targeted resequencing showed that 24 of 31 tumors with disomy 3 (77%) had mutations in either EIF1AX (15; 48%) or SF3B1 (9; 29%). Mutations were infrequent (2/35; 5.7%) in uveal melanomas with monosomy 3, which are associated with poor prognosis2. Resequencing of 13 uveal melanomas with partial monosomy 3 identified 8 tumors with a mutation in either SF3B1 (7; 54%) or EIF1AX (1; 8%). All EIF1AX mutations caused in-frame changes affecting the N terminus of the protein, whereas 17 of 19 SF3B1 mutations encoded an alteration of Arg625. Resequencing of ten uveal melanomas with disomy 3 that developed metastases identified SF3B1 mutations in three tumors, none of which targeted Arg625.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
Primary accessions
European Nucleotide Archive
Ensembl
References
Onken, M.D., Worley, L.A., Ehlers, J.P. & Harbour, J.W. Gene expression profiling in uveal melanoma reveals two molecular classes and predicts metastatic death. Cancer Res. 64, 7205–7209 (2004).
Prescher, G. et al. Prognostic implications of monosomy 3 in uveal melanoma. Lancet 347, 1222–1225 (1996).
Tschentscher, F. et al. Tumor classification based on gene expression profiling shows that uveal melanomas with and without monosomy 3 represent two distinct entities. Cancer Res. 63, 2578–2584 (2003).
Damato, B., Dopierala, J.A. & Coupland, S.E. Genotypic profiling of 452 choroidal melanomas with multiplex ligation-dependent probe amplification. Clin. Cancer Res. 16, 6083–6092 (2010).
Thomas, S. et al. Prognostic significance of chromosome 3 alterations determined by microsatellite analysis in uveal melanoma: a long-term follow-up study. Br. J. Cancer 106, 1171–1176 (2012).
Abdel-Rahman, M.H. et al. Frequency, molecular pathology and potential clinical significance of partial chromosome 3 aberrations in uveal melanoma. Mod. Pathol. 24, 954–962 (2011).
Van Raamsdonk, C.D. et al. Frequent somatic mutations of GNAQ in uveal melanoma and blue naevi. Nature 457, 599–602 (2009).
Van Raamsdonk, C.D. et al. Mutations in GNA11 in uveal melanoma. N. Engl. J. Med. 363, 2191–2199 (2010).
Bauer, J. et al. Oncogenic GNAQ mutations are not correlated with disease-free survival in uveal melanoma. Br. J. Cancer 101, 813–815 (2009).
Harbour, J.W. et al. Frequent mutation of BAP1 in metastasizing uveal melanomas. Science 330, 1410–1413 (2010).
Harbour, J.W. et al. Recurrent mutations at codon 625 of the splicing factor SF3B1 in uveal melanoma. Nat. Genet. 45, 133–135 (2013).
Forbes, S.A. et al. The Catalogue of Somatic Mutations in Cancer (COSMIC). Curr. Protoc. Hum. Genet. Chapter 10 Unit 10.11 (2008).
Rossi, D. et al. Mutations of the SF3B1 splicing factor in chronic lymphocytic leukemia: association with progression and fludarabine-refractoriness. Blood 118, 6904–6908 (2011).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Hahn, C.N. & Scott, H.S. Spliceosome mutations in hematopoietic malignancies. Nat. Genet. 44, 9–10 (2012).
Corrionero, A., Minana, B. & Valcarcel, J. Reduced fidelity of branch point recognition and alternative splicing induced by the anti-tumor drug spliceostatin A. Genes Dev. 25, 445–459 (2011).
Folco, E.G., Coil, K.E. & Reed, R. The anti-tumor drug E7107 reveals an essential role for SF3b in remodeling U2 snRNP to expose the branch point–binding region. Genes Dev. 25, 440–444 (2011).
Quesada, V. et al. Exome sequencing identifies recurrent mutations of the splicing factor SF3B1 gene in chronic lymphocytic leukemia. Nat. Genet. 44, 47–52 (2012).
Battiste, J.L., Pestova, T.V., Hellen, C.U. & Wagner, G. The eIF1A solution structure reveals a large RNA-binding surface important for scanning function. Mol. Cell 5, 109–119 (2000).
Chaudhuri, J., Si, K. & Maitra, U. Function of eukaryotic translation initiation factor 1A (eIF1A) (formerly called eIF-4C) in initiation of protein synthesis. J. Biol. Chem. 272, 7883–7891 (1997).
Fekete, C.A. et al. The eIF1A C-terminal domain promotes initiation complex assembly, scanning and AUG selection in vivo. EMBO J. 24, 3588–3601 (2005).
Maag, D. & Lorsch, J.R. Communication between eukaryotic translation initiation factors 1 and 1A on the yeast small ribosomal subunit. J. Mol. Biol. 330, 917–924 (2003).
Fekete, C.A. et al. N- and C-terminal residues of eIF1A have opposing effects on the fidelity of start codon selection. EMBO J. 26, 1602–1614 (2007).
Olsen, D.S. et al. Domains of eIF1A that mediate binding to eIF2, eIF3 and eIF5B and promote ternary complex recruitment in vivo. EMBO J. 22, 193–204 (2003).
Hinnebusch, A.G. Translational regulation of GCN4 and the general amino acid control of yeast. Annu. Rev. Microbiol. 59, 407–450 (2005).
Baird, T.D. & Wek, R.C. Eukaryotic initiation factor 2 phosphorylation and translational control in metabolism. Adv. Nutr. 3, 307–321 (2012).
Ingolia, N.T., Lareau, L.F. & Weissman, J.S. Ribosome profiling of mouse embryonic stem cells reveals the complexity and dynamics of mammalian proteomes. Cell 147, 789–802 (2011).
Lee, S., Liu, B., Huang, S.X., Shen, B. & Qian, S.B. Global mapping of translation initiation sites in mammalian cells at single-nucleotide resolution. Proc. Natl. Acad. Sci. USA 109, E2424–E2432 (2012).
McLean, I.W., Foster, W.D. & Zimmerman, L.E. Uveal melanoma: location, size, cell type, and enucleation as risk factors in metastasis. Hum. Pathol. 13, 123–132 (1982).
Tschentscher, F., Prescher, G., Zeschnigk, M., Horsthemke, B. & Lohmann, D.R. Identification of chromosomes 3, 6, and 8 aberrations in uveal melanoma by microsatellite analysis in comparison to comparative genomic hybridization. Cancer Genet. Cytogenet. 122, 13–17 (2000).
Church, D.M. et al. Modernizing reference genome assemblies. PLoS Biol. 9, e1001091 (2011).
Li, H. & Durbin, R. Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25, 1754–1760 (2009).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Thorvaldsdóttir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Acknowledgements
We thank M. Klutz and D. Falkenstein for technical assistance and H.-J. Lüdecke for elaborate discussion. This work was supported by the Deutsche Krebshilfe (DKF, 108612), the Mercator Research Center Ruhr (Pr-2010-0016) and the Wilhelm Sander Stiftung (2012.006.1) and by the Intramural Research Program of the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
M.Z. designed and supervised the study, and wrote the manuscript. C.M. and N.B. provided the samples and clinical data of the affected individuals. J.v.d.N. performed immunohistochemistry. L.K.-H. performed next-generation sequencing. M.M. and S.R. performed bioinformatic analysis of exome sequencing data. L.M. performed exome capture of tumor and blood samples. C.M. and P.T. performed Sanger resequencing and analysis of the sequencing data. B.H. and D.R.L. participated in the design of the study and revised the manuscript. D.R.L. performed statistical analysis. A.G.H. provided background on eIF1A function.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–3 and Supplementary Tables 1–5 (PDF 964 kb)
Rights and permissions
About this article
Cite this article
Martin, M., Maßhöfer, L., Temming, P. et al. Exome sequencing identifies recurrent somatic mutations in EIF1AX and SF3B1 in uveal melanoma with disomy 3. Nat Genet 45, 933–936 (2013). https://doi.org/10.1038/ng.2674
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2674
This article is cited by
-
Genomic alterations in thyroid cancer: biological and clinical insights
Nature Reviews Endocrinology (2024)
-
The journey from melanocytes to melanoma
Nature Reviews Cancer (2023)
-
RNA splicing dysregulation and the hallmarks of cancer
Nature Reviews Cancer (2023)
-
Analysis of uveal melanomas and paired constitutional DNA for exclusion of a BAP1-tumor predisposition syndrome
Familial Cancer (2023)
-
Genome-wide aberrant methylation in primary metastatic UM and their matched metastases
Scientific Reports (2022)