Abstract
H2 turnover at the [FeFe]-hydrogenase cofactor (H-cluster) is assumed to follow a reversible heterolytic mechanism, first yielding a proton and a hydrido-species which again is double-oxidized to release another proton. Three of the four presumed catalytic intermediates (Hox, Hred/Hred and Hsred) were characterized, using various spectroscopic techniques. However, in catalytically active enzyme, the state containing the hydrido-species, which is eponymous for the proposed heterolytic mechanism, has yet only been speculated about. We use different strategies to trap and spectroscopically characterize this transient hydride state (Hhyd) for three wild-type [FeFe]-hydrogenases. Applying a novel set-up for real-time attenuated total-reflection Fourier-transform infrared spectroscopy, we monitor compositional changes in the state-specific infrared signatures of [FeFe]-hydrogenases, varying buffer pH and gas composition. We selectively enrich the equilibrium concentration of Hhyd, applying Le Chatelier’s principle by simultaneously increasing substrate and product concentrations (H2/H+). Site-directed manipulation, targeting either the proton-transfer pathway or the adt ligand, significantly enhances Hhyd accumulation independent of pH.
Similar content being viewed by others
Introduction
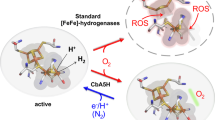
Hydrogen (H2) turnover in [FeFe]-hydrogenases is proposed to occur at the distal iron site (Fed) of the [2Fe2S] moiety (2FeH), which is part of the H-cluster1. The distal iron ion exchanges protons with a conserved proton-transfer pathway (PTP)2,3 via an azadithiolate ligand (adt) that bridges both iron sites (Fig. 1)4,5. Different redox states of the H-cluster can be distinguished by the unique infrared vibrational signatures of the two CN− and three CO ligands at the 2FeH moiety. To tailor [FeFe]-hydrogenases for specific applications or translate their catalytic principle into productive and durable synthetic mimics, detailed knowledge of the succession of turnover steps occurring at the H-cluster is required. The binuclear metal cofactors of [FeFe]- and [NiFe]-hydrogenases have been suggested to oxidize H2 by enhancing the acidity of H2 in the presence of a base, thus facilitating its heterolytic cleavage into a proton (H+) and hydride (H−) as a first intermediate step (Fig. 1)6,7,8,9.
The electron-transfer path is shown in yellow, the PTP is depicted in blue. H2 is released from, or reaches the 2FeH cluster through hydrophobic gas channels (green). At the catalytically pivotal 2FeH site, the presumptive first step in the catalysis of H2 oxidation (H2 heterolysis) is depicted, resulting in the unequal intermediates H+, binding at the adt-bridge (Lewis base) and H−, binding at Fed (Lewis acid).
For [NiFe]-hydrogenases, electron paramagnetic resonance spectroscopy showed that a bridging metal hydride was present in the paramagnetic Ni-C state6,7, while an ultra-high 0.89 Å resolution X-ray crystallography structure enabled the assignment of both, the proton and bridging hydride, in the first state following H2-binding (Ni-R)8. For algal-type [FeFe]-hydrogenases, a comparative density functional theory (DFT)/X-ray absorption spectroscopy analysis indicated a bridging hydride (μ-H−) in the double-reduced Hsred state10, albeit the importance of a μ-H− state for fast catalytic turnover has been questioned. In fact, the terminal hydride (t-H−) is thermodynamically less stable and more reactive compared to μ-H− and would match the high turnover kinetics of [FeFe]-hydrogenases far better11,12,13,14. According to the current working model for the catalytic mechanism of [FeFe]-hydrogenases1, binding of H2 to the oxidized active ready state (Hox) results in the heterolytic cleavage of H2, with Hhyd as the first intermediate. On deprotonation, Hhyd is converted into the rather stable Hsred state before two successive oxidation steps recycle Hox. A direct conversion of Hhyd into Hred has been proposed as well15. However, a terminal hydride has not been assigned to any of the known redox states10, nor has another catalytic wild-type state with the postulated t-H− been reported. This implies that the kinetically relevant hydride state is a transient one and difficult to trap under steady-state conditions. Owing to the rules governing steady-state kinetics, which favour thermodynamically stable intermediates in catalysis, the direct characterization of transient states relies on the utilization of time-resolved approaches with stopped-flow or single-turnover set-ups16.
We take a different approach, exploiting the properties of steady-state kinetics and using Le Chatelier’s principle to selectively enrich the transient t-H− state in three different [FeFe]-hydrogenases: HydA1; DdH; and CpI, which cover the complete range of increasingly complex structured monomeric hydrogenase subtypes (M1–M3)17,18,19,20. In our experimental set-up, attenuated total reflection (ATR) Fourier-transform infrared (FTIR) spectroscopy is used to follow the population of different catalytic states. Compositional changes are monitored in the state-specific infrared signatures as a function of buffer/gas composition.
Results
Accumulation of Hhyd in wild-type [FeFe]-hydrogenases
Figure 2a depicts FTIR difference spectra of HydA1 film at pH 8 recorded while changing the atmosphere from N2 to H2 (for details about the experimental setup see Supplementary Fig. 1). Negative bands mainly comprised the frequency pattern of Hox (1,964/1,940/1,802 cm−1). Positive bands included marker frequencies 1,915/1,891 cm−1 (Hred) and 1,953/1,918/1,882 cm−1 (Hsred), corresponding to earlier measurements with H2-purged HydA1 carried out in transmission FTIR systems21. On steady-state supply of substrate (that is, H2), the enzyme accumulates the apparently rather stable reduced intermediates Hred and Hsred. However, bands at 1,978/1,960/1,860 cm−1 could be identified as minor contributions, indicating the presence of another state, which resembles the band pattern of Htrans in DdH (Htrans-like state)22 and was previously observed as a weak fraction of a complex band pattern in a sodium dithionite (NaDT)-treated HydA1 sample23. According to our working model, deprotonation should lead from the presupposed Hhyd to the reduced states (Fig. 2b), and a plausible way to selectively enrich Hhyd would be a simultaneous increase in substrate (H2) and product (H+) pressure (Fig. 2d). We therefore titrated HydA1 to pH 4 before H2 exposure, enhancing the proton pressure by four orders of magnitude (Fig. 2c). The presence of a band at 1,891 cm−1 in the resulting difference spectrum hinted at a minor contribution of Hred, while no Hsred could be detected. Instead, the Htrans-like state represented the main fraction of our protein sample. No loss of cofactor was observed in the time frame of hours (Supplementary Fig. 2a,b), and the back-titration of HydA1 sample buffer from pH 4 to pH 8 clearly demonstrates a pH-dependent reversibility of the indicated state transitions (Supplementary Fig. 2b,c). The same applies for H2-dependency at pH 4 (Supplementary Fig. 2d). This bi-directionality is not compatible with the supposed Htrans-like nature of the novel pattern22. Therefore, we will refer to this redox state in the following as Hhyd. Similar results were obtained when repeating this experiment with samples of native DdH and CpI as representatives of the other two major [FeFe]-hydrogenase subtypes, implying this state and behaviour to be general features of the [FeFe]-hydrogenase family (Supplementary Fig. 3).
(a) ATR FTIR difference spectrum (N2/pH8→H2/pH8), depicting the accumulation of the reduced states (Hred/Hred and Hsred) in the catalytic cycle of HydA1 under H2 at pH 8. (c) ATR FTIR difference spectrum (N2/pH8→H2/pH4) depicting the accumulation of Hhyd in the catalytic cycle of HydA1 under H2 at pH 4. Catalytic cycles (b,d) beside the corresponding spectra illustrate the shift in steady-state equilibrium depending on applied substrate and product pressures in the working model of the catalytic cycle, which comprises the already-characterized states Hox, Hred/Hred and Hsred, and the missing H2-activated state (Hhyd). Peak labels for Hox are presented in black, for Hhyd in blue, for Hred in green and for Hsred in magenta (red asterisks: Hred). For the complete state-specific CN−-/CO-vibrational spectra of HydA1 observed during sample analysis, see Supplementary Table 1. At pH 4, the vibrational signals of Hox were slightly shifted to higher frequencies (for more information see Supplementary Table 1, Hox-blue).
The influence of proton-transfer efficiency
As the H2/H+ balance appears to affect the electronic configuration of the H-cluster, alternative approaches were designed to enrich Hhyd. HydA1 sample was once more purged with H2 aerosol to first enrich the reduced states. When humidity was removed from the H2 gas stream, sample films reacted by rapid dehydration. As predicted, this as well leads to a loss of Hred/Hsred in favour of the 1,978/1,960/1,860 cm−1 pattern (Supplementary Fig. 4a) indicating that the loss of bulk water has a similar effect to lowering the buffer pH, likely due to subsiding proton-transfer efficiency24 (Supplementary Fig. 4).
To verify that an interference with the PTP enhances the population of the Hhyd state under H2 exposure, we blocked the PTP in HydA1 and CpI via site-directed mutagenesis at two strictly conserved positions for each protein. Position 1 (C169 of HydA1/C299 of CpI; Fig. 1), which was shown in earlier experimental data to be involved in catalytic proton transfer3,5, marks the most proximal position of the PTP to the 2FeH sub-cluster, while position 2 (E141 of HydA1/E279 of CpI; Fig. 1) is situated more distant from the H-cluster. An exchange to alanine almost completely abolished H2-release activity for all variants (Supplementary Fig. 6). To probe the structural integrity and unequivocally connect the monitored effects to an interrupted PTP, we solved the crystal structure of CpI E279A (Fig. 3). The X-ray structure illustrates a fully intact H-cluster and overall identical structural features compared to wild-type CpI (refs 19, 25), with the exception of the clearly diminished electron density at position 279, which is in full accordance with an exchange from glutamate to alanine. As a consequence of this non-conservative exchange, the path of polar side chains is interrupted between S319 and the harnessed H2O molecule. Figure 4a,b shows the reaction of enzyme variants C169A of HydA1 and E279A of CpI towards H2 (black graph) at pH 8. Similar to the effect of acidification or sample drying, the oxidized state is lost in favour of bands at 1,978/1,962/1,863 cm−1 or 1,985/1,970/1,858 cm−1, respectively.
Stick models and simulated annealing omit |Fo–Fc| electron density maps (blue) of the 2FeH cluster, and the amino-acid residues and the H2O molecules involved in proton transfer embedded in the 2Fo–Fc electron density maps (grey) of the X-ray structures of CpI wild-type25 (a) and variant E279A (b). The presumed H+-transfer pathway is indicated in yellow. Crystal structure data for wild-type CpI correspond to pdb database entry 4XDC25. Details of the crystallographic data of E279A are summarized in Supplementary Table 2. For stereo view presentations see Supplementary Fig. 9.
Left: ATR FTIR difference spectra (N2/pH8→H2 or D2/pH8) depicting the accumulation of Hhyd for variants of the PTP E279A (CpI) (a) and C169A (HydA1) (b), as well as HydA1odt (c) under H2 or D2 at pH 8. The grey spectrum in a depicts the absolute absorption bands of Hhyd. Negative bands are assigned to Hox (Supplementary Table 1). Right: location of the corresponding PTP manipulation illustrated in a crystal structure model of CpI (pdb ID: 4XDC). Accumulation of Hhyd under D2 (red spectra) does not affect the tCO-frequencies but induces a selective redshift for the μCO signal of 6–8 cm−1 in all analysed variants. For the complete state-specific CN−/CO-vibrational spectra observed during sample analysis, see Supplementary Table 1.
We further exploited the effect of Hhyd accumulation as a consequence of PTP obstruction to elucidate the proposed role of the azadithiolate ligand in the heterolytic cleavage of H2. The earlier-described procedure of in vitro maturation allows for the replacement of the native 2FeH site with derivatives such as odt-2FeH, which contains an oxodithiolate group ((SCH2)2O,odt) instead of the native azadithiolate ((SCH2)2NH, adt)4,25,26. After in vitro maturation with odt-2FeH in presence of H2, HydA1odt exhibits an infrared band pattern that strongly resembles Hhyd (refs 23, 26). The oxidized state of HydA1odt was obtained via extensive purging with humidified N2. Subsequent exposure to H2 induced a decrease in the Hox pattern in favour of bands 1,980, 1,962 and 1868, cm−1 (Fig. 4c, black graph), however at a significantly lower rate compared to the corresponding experiment with wild-type HydA1.
A terminal hydride in wild-type [FeFe]-hydrogenases
To confirm the presence of a terminal hydride (t-H−) in Hhyd for wild-type hydrogenase and all examined variants, we probed the effect of an H/D isotope exchange on the FTIR band position of the bridging CO ligand, μCO (see Fig. 4, red graph, and Supplementary Figs 7 and 8 including Supplementary Note 1). Terminal hydride species on metal carbonyl complexes are vibrationally coupled with the ligand positioned in trans due to resonant frequencies27. Previously published DFT calculations on the H-cluster model of HydA1 variant C169S predict a selective ‘redshift’ of 6 cm−1 for the μCO frequency as a consequence of substituting a hydride for a deuteride species located trans to μCO (ref. 23). When switching the purging gas from H2 to D2, CpI E279A (Fig. 4a) and HydA1 C169A (Fig. 4b) showed a selective shift to lower energies by 5–6 cm−1 of the band at 1,860 cm−1, which corresponds to the stretching vibration of μCO. A similar ‘redshift’ was observed for HydA1odt when the oxidized enzyme was extensively treated with D2 (Fig. 4c and Supplementary Fig. 7b). In Supplementary Fig. 8 we show spectra that document the H/D effect on wild-type HydA1 in presence of pH 4 and H2/D2. This confirms the corresponding behaviour of the Hhyd state for all protein variants examined here.
Discussion
Le Chatelier’s principle was applied to enrich the highly transient H-cluster intermediate Hhyd, which according to the current working model of catalytic H2 turnover is predicted to carry a terminal hydride species. By simultaneously enhancing substrate and product pressure, we accumulated Hhyd for three different wild-type [FeFe]-hydrogenases. Interestingly, a nearly identical FTIR band pattern was observed in our earlier spectroscopic analysis of the largely inactive HydA1 PTP variant C169S (ref. 28). Using DFT calculations, Mulder et al.15,23 assigned this band pattern and redox species to a model of C169S that carried a terminal hydride at Fed (Supplementary Note 1 and Supplementary Fig. 5).
Accumulation of Hhyd was also observed on decreasing the humidity level of the protein sample. Recently, we demonstrated that freeze-drying renders [FeFe]-hydrogenase samples insensitive to O2 exposure24. Although X-ray absorption spectroscopy data showed the formation of a stable O2 adduct at the 2FeH site, H-cluster degradation was not initiated, as opposed to hydrated enzyme in presence of O2 (ref. 29). DFT calculations suggested a protonation step mandatory for generating destructive reactive oxygen species30. Therefore, a lack of H+-transfer activity in the freeze-dried state appeared to be a plausible explanation for this effect and dehydration should have a similar effect as lowering the pH. It can be speculated that the loss of bulk water, being the acceptor pool for exported protons, leads to an accumulation of protons in the PTP from the inside. Accordingly, low pH and dehydration both provide a counter pressure that prevents the H2-activated state from being deprotonated and proceeding to the reduced states (Supplementary Fig. 4b,c).
The N2/H2 difference spectra of site-directed exchange variants of HydA1 and CpI targeting positions C169/C299 and E141/E279 clearly demonstrate how the disrupted proton transport stabilizes the Hhyd state regardless of buffer pH. This highlights that the transient character of Hhyd essentially depends on a functional catalytic proton transfer, just as predicted from the working model (Figs 1 and 2). In return, the analogous behaviour of cysteine and glutamate variants provides experimental proof for the involvement of the median glutamic acid residue (E141HydA1/E279CpI) in the proton-transfer mechanism of [FeFe]-hydrogenases. In future mutagenesis studies on the PTP of [FeFe]-hydrogenases, accumulation of Hhyd can be regarded as an experimental indicator for the involvement of peptide positions in the catalytic H+ transfer.
The relevance of the bridgehead position of the 2FeH subcluster for catalytic proton transfer was clearly demonstrated by the fact that cofactor variant HydA1odt could be trapped in Hhyd under H2 independent of the pH, thus confirming the results of earlier studies, which pointed out the essential role of the native adt ligand as a proton relay to the substrate-binding site at Fed. The fact that H2 exposure of HydA1odt results in Hhyd accumulation suggests that the ether bridgehead of odt is capable to serve as a Lewis base and supports heterolytic H2 cleavage. This reflects earlier investigations of Barton et al.,31 which demonstrated in a comparative study, including different diiron complexes that the weakly basic ether group of odt can assist in proton relay during H2 evolution31. However, in contrast to C169A and E279A, the Hhyd spectrum of HydA1odt could only be regained very slowly in presence of H2. While all examined enzyme variants exhibited an impaired proton transfer, for C169A and E279A the defect was restricted to the H+ transfer across the protein shell (Fig. 4). In case of HydA1odt, the impairment already affected proton abstraction during H2 heterolysis. This finding experimentally verifies the hypothesis that the pending amine group in the native adt ligand significantly enhances the rate of heterolytic H2 cleavage31. The selective shift of the μCO band observed in Hhyd on H/D exchange suggests the presence of a terminal hydride ligand for all PTP variants and catalytically competent wild-type enzymes examined here.
In addition, two recently published studies independently demonstrated a terminal hydride for inactive HydA1 variants assigned to the presumptive Hhyd state. Reijerse et al.32 reported the direct detection of Fed-H bending vibrations via nuclear resonance vibrational spectroscopy of a terminal hydride species in HydA1odt after isolation under H2. This is in good agreement with our real-time analysis of HydA1odt. Furthermore, Mulder et al.15 applied electron paramagnetic resonance and Mössbauer spectroscopy to provide experimental support for the presence of a metal hydride species in HydA1 variant C169S, which largely adopted Hhyd on supplementation with NaDT. Well in agreement with electrochemistry studies on HydA1 under turnover conditions14, the authors determined a Hhyd transition potential close to the H2/H+ redox couple. This argues in favour of the relevance of Hhyd in the catalytic cycle.
Both HydA1 C169S and HydA1odt have been shown to be largely inactive4,15,23,25,26,28,32, and a definite proof for the catalytic relevance of Hhyd can only be given by monitoring the succession of states during catalysis at sub-turnover time resolution as previously done for [NiFe]-hydrogenase16. However, this work strongly suggests that Hhyd, as identified by Mössbauer and nuclear resonance vibrational spectroscopy, corresponds to the first transient state in the catalytic mechanism of H2 oxidation and is shared by all major subtypes of [FeFe]-hydrogenases. It is therefore plausible to assume that we successfully accumulated a state carrying a terminal hydride ligand in native [FeFe]-hydrogenases. In response to modulating substrate and product concentrations (H+/H2), accumulation of Hhyd was found to be fully reversible showing that Hhyd is no artificial dead-end state but part of the dynamic redox equilibrium typically occurring in the catalytically competent enzyme. All available data strongly suggest that Hhyd is indeed the missing link in the catalytic mechanism of [FeFe]-hydrogenases.
Methods
Site-directed mutagenesis of HydA1 and CpI
Plasmids pET21b-HydA1Cr and pET21b-CpI were used as templates for site-directed mutagenesis, which was performed according to the QuikChange-PCR protocol published by Zheng et al.33 For generating the constructs encoding HydA1 variants C169A, C169S and E141A, the 5′-overlapping mismatch primer pairs 5′-CAGCGCGTGTCCGGGCTGGATTGC-3′/5′-CCGGACACGCGCTGGTAAACATCGG-3′, 5′-CAGCTCATGTCCGGGCTGGATTGC-3′/5′-CCGGACATGAGCTGGTAAACATC-GG-3′ and 5′-CATCATGGAAGCGGGCAGCGAACTGCTGCATCGTC-3′/5′-GCAGTTC-GCTGCCCGCTTCCATGATGGTCAGATCC-3′ were used, respectively.
For substitutions C299A and E279A within the polypeptide of CpI, primer pairs 5′-CCTCTGCGTGCCCAGGTTGGGTACGTC-3′/5′-CCTGGGCACGCAGAGGTAAACATT-GGGAAAGGG-3′ and 5′-CCAGTTCGGTAGCCGCTTCCATAATGGTCATATC-3′/5′-TG-GAAGCGGCTACCGAACTGGTTCAACG were utilized.
Heterologous expression of HydA1 and CpI proteins
The original plasmids pET21b-HydA1Cr and pET21b-CpI and their mutagenesis constructs were used for the heterologous expression of wild-type enzymes and variants of HydA1 and CpI (refs 4, 25, 34, 35). Apo-forms of CpI and HydA1 (lacking 2FeH) were expressed anaerobically in Escherichia coli strain BL21 ΔiscR (ref. 36). Cells first grew aerobically in lysogeny broth (LB) medium pH 7.4 (0.1 M morpholineopropanesulfonic acid (Mops)-NaOH) supplemented with 5 g l−1 glucose and 2 mM ammonium iron citrate until an OD600 of 0.35–0.6 was reached. Before switching to anaerobic cultivation, the culture was flush with N2 for 30 min to remove residual O2 and supplemented with 25 mM sodium fumarate. Induction of gene expression was initiated by adding 5 mM cysteine and 0.5 mM β-D-1-thiogalactopyranoside. Cells were collected after 16–24 h expression via centrifugation.
Purification of [FeFe]-hydrogenase apo-proteins
The apo-proteins of wild-type and mutant forms of HydA1 and CpI, containing the 4FeH subcluster but lacking the 2FeH moiety, were expressed under anaerobic conditions in E. coli strain BL21 ΔiscR as described earlier4,34,36. After cell disruption via ultrasonication and the separation of the soluble proteins from the cell debris by ultracentrifugation and subsequent filtering (pore size 0.2 μm), we exploited the C-terminally fused Strep-tag-II sequence (WSHPQFEK) for protein purification, performing affinity chromatography with Strep-Tactin Superflow resin (IBA GmbH) in 100 mM Tris-HCl buffer (pH 8) supplemented with 2 mM NaDT. Protein purity was verified by SDS–PAGE and protein concentration determined via Bradford assay (BioRad).
Synthesis of 2FeH cofactor complexes
The native-like 2FeH-cofactor-mimic complex [Fe2[μ-(SCH2)2NH](CN)2(CO)4][Et4N]2 (adt-2FeH) and its odt derivative [Fe2[μ-(SCH2)2O](CN)2(CO)4][Et4N]2 (odt-2FeH) were synthesized according to literature procedures25.
In vitro maturation
To obtain the fully equipped holoprotein, heterologously expressed wild-type and mutant apo-proteins of HydA1 and CpI were maturated in vitro by adding adt-2FeH (odt-2FeH) to a 10-fold excess in 0.1 M K2HPO4/KH2PO4 buffer (pH 6.8), supplemented with 2 mM NaDT. To ensure complete sample maturation, the reaction mix was incubated for 1 h at room temperature. The resulting holoproteins were cleaned from surplus 2FeH complex by size-exclusion chromatography using NAP 5 columns of GE Healthcare and concentrated in Amicon Ultra centrifugal filters 30K (Millipore). Afterwards, methyl viologen-specific H2 production activity was determined to assess catalytic competence.
In vitro assay to determine H2 production activity
H2 production activities have been determined in vitro for wild-type and mutant proteins using 0.2 μg ml−1 of in vitro maturated [FeFe]-hydrogenase in 2 ml of 100 mM postassium phosphate buffer, pH 6.8, supplemented with 100 mM NaDT and 10 mM methyl viologen. Suba-seal vessels were sealed with stoppers, degassed for 5 min with 100% Argon and incubated for 20 min in a shaking water bath adjusted to 37 °C. For product quantification, 400 μl of sample head-space was analysed via gas chromatography (Shimadzu GC 2010). When enzyme activity was below the detection limit, the assay was repeated with a 10-fold increased enzyme concentration (2 μg ml−1).
Crystallization and structure analysis of CpI variant E279A
Using a 1:1 mix of E279A holoprotein (10 mg ml−1) and reservoir solution (0.1 M MES, pH 6, supplemented with 0.4 M MgCl2, 24% polyethylene glycol and 16% glycerol) in a hanging drop vapour diffusion set-up, single cuboid-shaped crystals of brownish colour were identified after 2–4 days at 277 K under anaerobic conditions. The selected protein crystal was mounted into a CryoLoop (Hampton Research) and flash-frozen in liquid N2. Diffraction data were collected at 100 K at beamline ID23-2 of the ESRF in Grenoble, France and processed using the software package XDS37. Molecular replacement and structure optimization were achieved with the software packages PHENIX38 and Coot39. For details on crystallographic data of CpI E279A, see Supplementary Table 2.
Infrared spectroscopy
All FTIR spectroscopy was conducted on a rapid-scan Tensor 27 spectrometer (Bruker Optik, Germany) equipped with a three-reflection ZnSe/silicon crystal ATR cell (Smith Detection, USA). The spectrometer was situated in an anaerobic gas chamber (Coy Laboratories, USA) in a water-free atmosphere of typically 99% N2 and 1% H2. Supplementary Fig. 1 depicts the process flow diagram of the experimental set-up. Approximately 5 bar nitrogen carrier gas of ultra-high purity (5.0) was provided by a PN1450 nitrogen generator (Inmatec, Germany). The exact amount of N2 was adjusted with a digital SmartTrak mass flow controller (MFC, Sierra, USA). Molecular H2 (H2 5.0, Linde, Germany) was injected separately via a second flow controller to create a well-defined mixture. The gas passed a 200 mbar check valve to protect the flow controllers from humidity. Afterwards, two sequential wash bottles (H2O and miscellaneous content, for example, buffer) could be switched into the gas stream. A bypass loop allowed for an adjustable proportion of the gas to bypass the wash bottles entirely or to run through the liquid volume to create a mix of carrier gas, water vapour and microscopic drops of liquid water (aerosol). The aerosol stream was fed to a customized polychlorotrifluoroethylene (PCTFE) gas cell, screwed gas-tight onto the ATR crystal plate. An exit line eventually guided the aerosol to a gas dump (fume). The PCTFE gas cell was equipped with three inlets and a manometer (not shown in Supplementary Fig. 1). Directly over the silicon crystal of the ATR unit, a cylindrical cavity joined all inlets. A gas-tight lid was screwed on top of the gas cell and allowed irradiation via an acrylic glass window. At this position, an infrared- or ultraviolet/visible-transparent window with protein sample could be adjusted into the aerosol stream. Thus, transmission samples for, for example, ultraviolet/visible or Raman spectroscopy could be prepared under the same conditions as monitored via the ATR silicon crystal. All spectra were recorded with 80 kHz scanning velocity, at a spectral resolution of 1 cm−1, and varying extent of co-additions. All pH titrations were performed with citrate buffer (SSC).
Data availability
The atomic coordinates and factors for the reported crystal structure for CpI variant E279A have been deposited with the Protein Data Bank (PDB) under accession code 5LA3. Further data supporting the findings of this study are available within the article and its Supplementary Information file and from the corresponding authors on reasonable request.
Additional information
How to cite this article: Winkler, M. et al. Accumulating the hydride state in the catalytic cycle of [FeFe]-hydrogenases. Nat. Commun. 8, 16115 doi: 10.1038/ncomms16115 (2017).
Publisher’s note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Change history
26 July 2017
The HTML version of this Article was updated shortly after publication, following a technical error that resulted in a significant loss of image quality in the Supplementary Information file. The HTML has now been updated to include the correct version of the Supplementary Information; the PDF version of the Article was correct from the time of publication.
References
Lubitz, W., Ogata, H., Rudiger, O. & Reijerse, E. Hydrogenases. Chem. Rev. 114, 4081–4148 (2014).
Cornish, A. J., Gartner, K., Yang, H., Peters, J. W. & Hegg, E. L. Mechanism of proton transfer in [FeFe]-hydrogenase from Clostridium pasteurianum. J. Biol. Chem. 286, 38341–38347 (2011).
Morra, S. et al. Site saturation mutagenesis demonstrates a central role for cysteine 298 as proton donor to the catalytic site in CaHydA [FeFe]-hydrogenase. PLoS ONE 7, e48400 (2012).
Berggren, G. et al. Biomimetic assembly and activation of [FeFe]-hydrogenases. Nature 499, 66–69 (2013).
Silakov, A., Wenk, B., Reijerse, E. & Lubitz, W. (14)N HYSCORE investigation of the H-cluster of [FeFe] hydrogenase: evidence for a nitrogen in the dithiol bridge. Phys. Chem. Chem. Phys. 11, 6592–6599 (2009).
Brecht, M., van Gastel, M., Buhrke, T., Friedrich, B. & Lubitz, W. Direct detection of a hydrogen ligand in the [NiFe] center of the regulatory H2-sensing hydrogenase from Ralstonia eutropha in its reduced state by HYSCORE and ENDOR spectroscopy. J. Am. Chem. Soc. 125, 13075–13083 (2003).
Foerster, S. et al. Single crystal EPR studies of the reduced active site of [NiFe] hydrogenase from Desulfovibrio vulgaris Miyazaki F. J. Am. Chem. Soc. 125, 83–93 (2003).
Ogata, H., Nishikawa, K. & Lubitz, W. Hydrogens detected by subatomic resolution protein crystallography in a [NiFe] hydrogenase. Nature 520, 571–574 (2015).
Schilter, D., Camara, J. M., Huynh, M. T., Hammes-Schiffer, S. & Rauchfuss, T. B. Hydrogenase enzymes and their synthetic models: the role of metal hydrides. Chem. Rev. 116, 8693–8749 (2016).
Chernev, P. et al. Hydride binding to the active site of [FeFe]-hydrogenase. Inorg. Chem. 53, 12164–12177 (2014).
Bruschi, M. et al. Influence of the [2Fe]H subcluster environment on the properties of key intermediates in the catalytic cycle of [FeFe] hydrogenases: hints for the rational design of synthetic catalysts. Angew. Chem. Int. Ed. 48, 3503–3506 (2009).
Carroll, M. E., Barton, B. E., Rauchfuss, T. B. & Carroll, P. J. Synthetic models for the active site of the [FeFe]-hydrogenase: catalytic proton reduction and the structure of the doubly protonated intermediate. J. Am. Chem. Soc. 134, 18843–18852 (2012).
Finkelmann, A. R., Stiebritz, M. T. & Reiher, M. Inaccessibility of the μ-hydride species in [FeFe] hydrogenases. Chem. Sci. 5, 215–221 (2014).
Hexter, S. V., Grey, F., Happe, T., Climent, V. & Armstrong, F. A. Electrocatalytic mechanism of reversible hydrogen cycling by enzymes and distinctions between the major classes of hydrogenases. Proc. Natl Acad. Sci. USA 109, 11516–11521 (2012).
Mulder, D. W., Guo, Y., Ratzloff, M. W. & King, P. W. Identification of a catalytic iron-hydride at the H-cluster of [FeFe]-hydrogenase. J. Am. Chem. Soc. 139, 83–86 (2017).
Greene, B. L., Wu, C. H., McTernan, P. M., Adams, M. W. & Dyer, R. B. Proton-coupled electron transfer dynamics in the catalytic mechanism of a [NiFe]-hydrogenase. J. Am. Chem. Soc. 137, 4558–4566 (2015).
Mulder, D. W. et al. Stepwise [FeFe]-hydrogenase H-cluster assembly revealed in the structure of HydA(DeltaEFG). Nature 465, 248–251 (2010).
Nicolet, Y., Piras, C., Legrand, P., Hatchikian, C. E. & Fontecilla-Camps, J. C. Desulfovibrio desulfuricans iron hydrogenase: the structure shows unusual coordination to an active site Fe binuclear center. Structure 7, 13–23 (1999).
Peters, J. W., Lanzilotta, W. N., Lemon, B. J. & Seefeldt, L. C. X-ray crystal structure of the Fe-only hydrogenase (CpI) from Clostridium pasteurianum to 1.8 angstrom resolution. Science 282, 1853–1858 (1998).
Winkler, M., Esselborn, J. & Happe, T. Molecular basis of [FeFe]-hydrogenase function: an insight into the complex interplay between protein and catalytic cofactor. Biochim. Biophys. Acta 1827, 974–985 (2013).
Adamska, A. et al. Identification and characterization of the ‘super-reduced’ state of the H-cluster in [FeFe] hydrogenase: a new building block for the catalytic cycle? Angew. Chem. Int. Ed. 51, 11458–11462 (2012).
Roseboom, W., De Lacey, A. L., Fernandez, V. M., Hatchikian, E. C. & Albracht, S. P. The active site of the [FeFe]-hydrogenase from Desulfovibrio desulfuricans. II. Redox properties, light sensitivity and CO-ligand exchange as observed by infrared spectroscopy. J. Biol. Inorg. Chem. 11, 102–118 (2006).
Mulder, D. W. et al. Investigations on the role of proton-coupled electron transfer in hydrogen activation by [FeFe]-hydrogenase. J. Am. Chem. Soc. 136, 15394–15402 (2014).
Noth, J. et al. Lyophilization protects [FeFe]-hydrogenases against O2-induced H-cluster degradation. Sci. Rep. 5, 13978 (2015).
Esselborn, J. et al. A structural view of synthetic cofactor integration into [FeFe]-hydrogenases. Chem. Sci. 7, 959–968 (2016).
Siebel, J. F. et al. Hybrid [FeFe]-hydrogenases with modified active sites show remarkable residual enzymatic activity. Biochemistry 54, 1474–1483 (2015).
Kaesz, H. D. & Saillant, R. B. Hydride complexes of the transition metals. Chem. Rev. 72, 231–281 (1972).
Knorzer, P. et al. Importance of the protein framework for catalytic activity of [FeFe]-hydrogenases. J. Biol. Chem. 287, 1489–1499 (2012).
Stripp, S. T. et al. How oxygen attacks [FeFe] hydrogenases from photosynthetic organisms. Proc. Natl Acad. Sci. USA 106, 17331–17336 (2009).
Bruska, M. K., Stiebritz, M. T. & Reiher, M. Regioselectivity of H cluster oxidation. J. Am. Chem. Soc. 133, 20588–20603 (2011).
Barton, B. E., Olsen, M. T. & Rauchfuss, T. B. Aza- and oxadithiolates are probable proton relays in functional models for the [FeFe]-hydrogenases. J. Am. Chem. Soc. 130, 16834–16835 (2008).
Reijerse, E. J. et al. Direct observation of an iron-bound terminal hydride in [FeFe]-hydrogenase by nuclear resonance vibrational spectroscopy. J. Am. Chem. Soc. 139, 4306–4309 (2017).
Zheng, L., Baumann, U. & Reymond, J. L. An efficient one-step site-directed and site-saturation mutagenesis protocol. Nucleic Acids Res. 32, e115 (2004).
Esselborn, J. et al. Spontaneous activation of [FeFe]-hydrogenases by an inorganic [2Fe] active site mimic. Nat. Chem. Biol. 9, 607–609 (2013).
Kuchenreuther, J. M. et al. High-yield expression of heterologous [FeFe] hydrogenases in Escherichia coli. PLoS ONE 5, e15491 (2010).
Akhtar, M. K. & Jones, P. R. Deletion of iscR stimulates recombinant clostridial Fe-Fe hydrogenase activity and H2-accumulation in Escherichia coli BL21(DE3). Appl. Microbiol. Biotechnol. 78, 853–862 (2008).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Acknowledgements
T.H. and M.W. gratefully acknowledge financial support from the Deutsche Forschungsgemeinschaft Cluster of Excellence RESOLV (EXC1069), the Volkswagen Foundation (LigH2t) and the EU (Sun2Chem). M.S. and S.T.S. thank the International Max Plank Research School (IMPRS) on Multiscale Biosystems and the Focus Area NanoScale (Freie Universität Berlin) for financial support. J.D. acknowledges support by the China Scholarship Council (CSC). U.-P.A. and F.W. are grateful for financial support by the Funds of the Chemical Industry (Liebig grant to U.-P.A.) and the Deutsche Forschungsgemeinschaft (Emmy Noether grant to U.-P.A., AP242/2-1). We further thank J. Fontecilla-Camps for providing a sample of DdH protein, and the staff at beam line ID23-2 at the ESRF, France, for their help during data collection. We gratefully acknowledge the support of Dr Anne Sawyer during the text revision process.
Author information
Authors and Affiliations
Contributions
M.W., S.T.S. and T.H. designed research; M.W., S.T.S. and T.H. wrote the manuscript; U.-P.A. and F.W. synthesized the 2FeH-adt/-odt complexes for in vitro maturation; J.D. and M.W. produced, purified and maturated the wild-type and mutant proteins of HydA1 and CpI; S.T.S. and M.S. performed and designed ATR FTIR experiments; J.D., E.H. and J.E. performed or contributed to protein crystallization and X-ray structure analysis.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Winkler, M., Senger, M., Duan, J. et al. Accumulating the hydride state in the catalytic cycle of [FeFe]-hydrogenases. Nat Commun 8, 16115 (2017). https://doi.org/10.1038/ncomms16115
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms16115
This article is cited by
-
A personal account on 25 years of scientific literature on [FeFe]-hydrogenase
JBIC Journal of Biological Inorganic Chemistry (2023)
-
A safety cap protects hydrogenase from oxygen attack
Nature Communications (2021)
-
Computational study of [(phenanthroline)2FeII/III–(terephthalate)–CoIII/II(phenanthroline)2]3+ binuclear complex
Structural Chemistry (2020)
-
Crystallographic and spectroscopic assignment of the proton transfer pathway in [FeFe]-hydrogenases
Nature Communications (2018)
-
The plasticity of redox cofactors: from metalloenzymes to redox-active DNA
Nature Reviews Chemistry (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.