Abstract
Mammals have nine different homologues (ALKBH1–9) of the Escherichia coli DNA repair demethylase AlkB. ALKBH2 is a genuine DNA repair enzyme, but the in vivo function of the other ALKBH proteins has remained elusive. It was recently shown that ALKBH8 contains an additional transfer RNA (tRNA) methyltransferase domain, which generates the wobble nucleoside 5-methoxycarbonylmethyluridine (mcm5U) from its precursor 5-carboxymethyluridine (cm5U). In this study, we report that (R)- and (S)-5-methoxycarbonylhydroxymethyluridine (mchm5U), hydroxylated forms of mcm5U, are present in mammalian  , and
, and  , respectively, representing the first example of a diastereomeric pair of modified RNA nucleosides. Through in vitro and in vivo studies, we show that both diastereomers of mchm5U are generated from mcm5U, and that the AlkB domain of ALKBH8 specifically hydroxylates mcm5U into (S)-mchm5U in
, respectively, representing the first example of a diastereomeric pair of modified RNA nucleosides. Through in vitro and in vivo studies, we show that both diastereomers of mchm5U are generated from mcm5U, and that the AlkB domain of ALKBH8 specifically hydroxylates mcm5U into (S)-mchm5U in  . These findings expand the function of the ALKBH oxygenases beyond nucleic acid repair and increase the current knowledge on mammalian wobble uridine modifications and their biogenesis.
. These findings expand the function of the ALKBH oxygenases beyond nucleic acid repair and increase the current knowledge on mammalian wobble uridine modifications and their biogenesis.
Similar content being viewed by others
Introduction
Transfer RNA (tRNA) modification is important both for correct folding and for optimal interaction with components of the protein synthesis machinery, such as aminoacyl transferases, messenger RNA (mRNA) and the ribosome. The nucleotide at the wobble position in the anticodon loop of tRNA interacts with the third nucleotide of the codon in mRNA, and wobble uridines are usually modified, both in bacteria and eukaryotes1,2. Modification of wobble uridines ensures efficient decoding of both cognate and non-cognate codons3,4, and prevents misreading of codons from 'split' boxes, where purine- and pyrimidine-ending codons encode different amino acids1.
In eukaryotes, most wobble uridines carry the modification 5-methoxycarbonylmethyluridine (mcm5U), 5-carbamoylmethyluridine (ncm5U) or derivatives thereof (Fig. 1a). These modifications and their biogenesis have been most closely studied in the yeast Saccharomyces cerevisiae. Here, tRNA isoacceptors that contain ncm5U decode in the 'family' codon boxes, in which all four codons encode the same amino acid, whereas tRNAs containing mcm5U usually decode in the split codon boxes. The Elongator complex has been shown to mediate an early step in mcm5U/ncm5U formation5, whereas the final methyl esterification reaction leading to the formation of mcm5U is catalysed by the methyltransferase (MT) Trm9 (ref. 6).
(a) Wobble uridine modifications described in the present work. mcm5U, 5-methoxycarbonylmethyluridine; mchm5U, 5-methoxycarbonylhydroxymethyluridine; chm5U, 5-carboxyhydroxymethyluridine; cm5U, 5-carboxymethyluridine; ncm5U, 5-carbamoylmethyluridine; nchm5U, 5-carbamoylhydroxymethyluridine. (b) Domain structure and proposed function of ALKBH8. RRM, RNA recognition motif. Together with the accessory protein TRM112, the MT domain of ALKBH8 converts cm5U into mcm5U. In the present work, the ALKBH8 oxygenase has been investigated for its ability to hydroxylate mcm5U into mchm5U. (c) Chemical structure of the two mchm5U diastereomers. (d) The anticodon stem-loop region of silkworm  and calf
and calf  . Anticodon (red line) containing products (blue) generated by RNase T1 cleavage (arrows), and analysed by MALDI-TOF MS, are indicated. (e) LC–MS/MS analysis of mchm5U nucleosides in calf liver total tRNA and isolated isoacceptors. (f) MALDI-quadrupole TOF tandem mass spectrometry of the RNase T1 product harbouring the anticodon of
. Anticodon (red line) containing products (blue) generated by RNase T1 cleavage (arrows), and analysed by MALDI-TOF MS, are indicated. (e) LC–MS/MS analysis of mchm5U nucleosides in calf liver total tRNA and isolated isoacceptors. (f) MALDI-quadrupole TOF tandem mass spectrometry of the RNase T1 product harbouring the anticodon of  . Fragmentation pattern suggests the sequence A-Cm-U-mchm5U-C-G>p as indicated. c-Ions contain the original 5′-end after cleavage of the phophodiester bond between the phosphorous and the 5′-oxygen; y-ions contain the original 3′-end after cleavage of the phophodiester bond between the phosphorous and the 5′-oxygen. w-Ions contain the original 3′-end after cleavage of the phophodiester bond between the 3′-carbon and the 3′-oxygen. The associated number gives the size of the fragment ion in nucleotide residues. >p denotes 2′–3′ cyclic phosphate.
. Fragmentation pattern suggests the sequence A-Cm-U-mchm5U-C-G>p as indicated. c-Ions contain the original 5′-end after cleavage of the phophodiester bond between the phosphorous and the 5′-oxygen; y-ions contain the original 3′-end after cleavage of the phophodiester bond between the phosphorous and the 5′-oxygen. w-Ions contain the original 3′-end after cleavage of the phophodiester bond between the 3′-carbon and the 3′-oxygen. The associated number gives the size of the fragment ion in nucleotide residues. >p denotes 2′–3′ cyclic phosphate.
The 2OG/Fe(II) (2-oxoglutarate- and Fe2+-dependent) oxygenase superfamily encompasses enzymes involved in important processes, for example, generation of hydroxyproline in collagen, regulation of hypoxia-responsive genes, demethylation of histone proteins, nucleic acid repair and, as discovered recently, introduction of the epigenetic modification 5-hydroxymethylcytosine (5hmC) in DNA7,8,9. The 2OG/Fe(II) oxygenase AlkB from Escherichia coli is a repair enzyme that can demethylate lesions in DNA and RNA, such as 1-methyladenine and 3-methylcytosine10,11,12. Bioinformatics analysis has identified a family of nine different mammalian AlkB homologues (ALKBH), including the obesity-associated protein FTO13,14. Although in vitro repair activities have been observed for some ALKBH proteins10,13,15,16, only ALKBH2 has been firmly established as a repair enzyme17, and non-repair functions have also been proposed18,19,20. We and others recently showed that ALKBH8 contains a MT domain, which is the functional mammalian Trm9 homologue, and requires a small accessory protein, TRM112, for activity (Fig. 1b)21,22.
As hydroxylation is the primary reaction catalysed by 2OG/Fe(II) oxygenases in animals, and as the MT domain of ALKBH8 is involved in mcm5U biogenesis, we were intrigued by a report documenting the presence of 5-methoxycarbonylhydroxymethyluridine ((S)-mchm5U), a hydroxylated form of mcm5U, in  from the silkworm Bombyx mori23 (Fig. 1c,d). Mammalian and worm
from the silkworm Bombyx mori23 (Fig. 1c,d). Mammalian and worm  are highly similar, and the primary sequence of the anticodon loop is identical (Supplementary Fig. S1). This suggested that (S)-mchm5U may be present also in mammalian tRNA, and that the ALKBH8 oxygenase could be responsible for its formation (Fig. 1b). Indeed, whilst our manuscript was under preparation, Fu et al. independently reported that the AlkB domain of murine ALKBH8 can hydroxylate wobble mcm5U into (S)-mchm5U in a synthetic substrate resembling the anticodon stem-loop of
are highly similar, and the primary sequence of the anticodon loop is identical (Supplementary Fig. S1). This suggested that (S)-mchm5U may be present also in mammalian tRNA, and that the ALKBH8 oxygenase could be responsible for its formation (Fig. 1b). Indeed, whilst our manuscript was under preparation, Fu et al. independently reported that the AlkB domain of murine ALKBH8 can hydroxylate wobble mcm5U into (S)-mchm5U in a synthetic substrate resembling the anticodon stem-loop of  24.
24.
In the present work, we demonstrate that (S)-mchm5U is found in the wobble position of mammalian  , and that its diastereomer, (R)-mchm5U, is found in
, and that its diastereomer, (R)-mchm5U, is found in  . Moreover, we show through studies of gene-targeted mice and recombinant enzymes that the MT domain of ALKBH8 provides the mcm5U precursor required for formation of mchm5U in
. Moreover, we show through studies of gene-targeted mice and recombinant enzymes that the MT domain of ALKBH8 provides the mcm5U precursor required for formation of mchm5U in  and
and  , whereas its AlkB domain represents an evolutionary conserved oxygenase that catalyses the hydroxylation of wobble mcm5U to (S)-mchm5U in
, whereas its AlkB domain represents an evolutionary conserved oxygenase that catalyses the hydroxylation of wobble mcm5U to (S)-mchm5U in  .
.
Results
Analysis of mchm5U modifications in mammalian tRNA
To investigate the possible presence of mchm5U in mammalian tRNA, we analysed the modification status of nucleosides from bovine tRNA by liquid chromatography coupled to tandem mass spectrometry (LC–MS/MS). In this analysis, a triple-quadrupole mass spectrometer was used to specifically select the relevant nucleoside ions, induce fragmentation and detect the resulting nucleobase fragment ions. Additional specificity was obtained by comparing chromatographic retention times with those of synthetic nucleoside standards. This allows highly specific, sensitive and simultaneous quantification of several nucleosides. Intriguingly, substantial amounts of both the S and R diastereomers of mchm5U were detected in total bovine tRNA nucleosides (Fig. 1e). We next isolated individual, wobble uridine-containing isoacceptors by hybridization to complementary, immobilized DNA oligonucleotides. We found, as expected, (S)-mchm5U to be present in  , whereas
, whereas  contained (R)-mchm5U (Fig. 1e). We were unable to identify additional wobble uridine-containing tRNA isoacceptors carrying a mchm5U modification (data not shown). The presence of wobble (R)-mchm5U in
contained (R)-mchm5U (Fig. 1e). We were unable to identify additional wobble uridine-containing tRNA isoacceptors carrying a mchm5U modification (data not shown). The presence of wobble (R)-mchm5U in  was supported by matrix-assisted laser desorption/ionization (MALDI)-time-of-flight (TOF) mass spectrometry of an RNase T1 digest of the purified isoacceptor. Sequencing by MALDI-quadrupole-TOF tandem mass spectrometry confirmed the presence of mchm5U at the wobble position, and the fragmentation pattern also revealed ribose methylation of C32 (ref. 25), a very common tRNA modification (Fig. 1d,f)2.
was supported by matrix-assisted laser desorption/ionization (MALDI)-time-of-flight (TOF) mass spectrometry of an RNase T1 digest of the purified isoacceptor. Sequencing by MALDI-quadrupole-TOF tandem mass spectrometry confirmed the presence of mchm5U at the wobble position, and the fragmentation pattern also revealed ribose methylation of C32 (ref. 25), a very common tRNA modification (Fig. 1d,f)2.
Analysis of wobble uridines in Alkbh8-targeted mice
To address the possible role of the ALKBH8 oxygenase in mchm5U biogenesis, we used a mouse model, in which essential exons of the Alkbh8 gene had been deleted (Alkbh8−/−; ref. 22). Moreover, we developed two novel 'knock-in' (KI) mice in which an oxygenase-deficient Alkbh8 transgene (KI(MT+)) or a MT-deficient Alkbh8 transgene (KI(Ox+)) was expressed from the Rosa26 or Hprt loci, respectively (Supplementary Figs S2–S5). These transgenes, or the combination of the two (KI(MT+/Ox+)), were introduced into the Alkbh8−/− background (Fig. 2a). The expression of the transgenes was verified by reverse transcription and PCR (data not shown). All the mouse models were devoid of any obvious phenotype. First, we analysed by LC–MS/MS, total tRNA nucleosides from the different mice for the presence of the two mchm5U diastereomers. As expected, wild-type (WT) tRNA contained both (R)- and (S)-mchm5U (Fig. 2b). tRNA from the Alkbh8−/− and KI(Ox+) mice, which lack functional MT activity, were devoid of both (R)- and (S)-mchm5U, whereas the KI(MT+) tRNA contained exclusively (R)-mchm5U. These data indicate that the formation of either mchm5U diastereomer requires the MT function, whereas the oxygenase is required for the formation of (S)-mchm5U only. This implies that a second unknown oxygenase, in the following referred to as OxX, is responsible for the hydroxylation of mcm5U to (R)-mchm5U. Furthermore, KI(MT+/Ox+) tRNA contained, like WT tRNA, both mchm5U stereoisomers (Fig. 2b), indicating that the MT and oxygenase activities of ALKBH8 can be uncoupled and provided by two individual proteins.
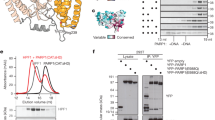
(a) Schematic representation of the ALKBH8 proteins expressed in the different mouse models. An asterisk indicates that the domain has been inactivated by point mutations. (b) Analysis of R and S diastereomers of mchm5U in total tRNA from gene-targeted mice. Murine tRNA was enzymatically degraded to nucleosides, which were analysed by LC–MS/MS. (c) Quantification of uridine modifications in total murine tRNA by LC–MS/MS. The level of each nucleoside is expressed as the molar percentage of the total amount of the modifications indicated in the figure. (d) Recombinant, E. coli-expressed ALKBH8 proteins, or individual domains, used for incubation with murine tRNA. (e) Modification status of murine tRNA treated with human ALKBH8 or its individual domains. ALKBH8 and ALKBH8-MT were associated with co-expressed, co-purified TRM112. (f) Modification status of murine  measured by LC–MS/MS. (g) MALDI-TOF analysis of the anticodon-containing RNase T1 product of
measured by LC–MS/MS. (g) MALDI-TOF analysis of the anticodon-containing RNase T1 product of  . The theoretical monoisotopic mass (green) as well as the measured mass (blue) of relevant modifications are indicated. Because of the natural isotope distribution, each modification gives rise to a cluster of peaks with 1.0 Da spacing where the leftmost peak corresponds to the monoisotopic mass. Overlapping isotope clusters from ncm5U and cm5U with clearly distorted peak patterns occur in the Alkbh8−/− and KI(Ox+) samples. Error bars represent range between duplicate samples in a typical experiment.
. The theoretical monoisotopic mass (green) as well as the measured mass (blue) of relevant modifications are indicated. Because of the natural isotope distribution, each modification gives rise to a cluster of peaks with 1.0 Da spacing where the leftmost peak corresponds to the monoisotopic mass. Overlapping isotope clusters from ncm5U and cm5U with clearly distorted peak patterns occur in the Alkbh8−/− and KI(Ox+) samples. Error bars represent range between duplicate samples in a typical experiment.
To study how manipulation of ALKBH8 activity affects the formation of modified uridines in vivo, we also measured the levels of various other modifications (Fig. 2c, Supplementary Figs S6 and S7). tRNA isolated from KI(MT+) mice displayed a strong accumulation of mcm5U, consistent with mcm5U being a direct substrate for the ALKBH8 oxygenase. In agreement with previous results22, we detected substantial amounts of 5-carboxymethyluridine (cm5U) and elevated levels of ncm5U in tRNA from the MT-deficient Alkbh8−/− and KI(Ox+) mice. Notably, we also detected 5-carbamoylhydroxymethyluridine (nchm5U) and trace amounts of 5-carboxyhydroxymethyluridine (chm5U), the hydroxylated forms of ncm5U and cm5U, in tRNA from the MT-deficient mice (note that the R and S diastereomers of these nucleosides could not be separated chromatographically). This indicates that OxX and possibly also the ALKBH8 oxygenase will hydroxylate ncm5U, and to a lesser extent, cm5U, when mcm5U is absent.
Enzymatic activity of recombinant ALKBH8 on murine tRNA
To investigate whether the observed hypomodification of tRNA from gene-targeted mice could be reversed by recombinant enzymes, total tRNA from mouse liver was incubated with E. coli-expressed human ALKBH8, or combinations of its individual oxygenase (ALKBH8-Ox) and MT (ALKBH8-MT) domains (Fig. 2d,e). The results demonstrated that ALKBH8-Ox hydroxylates mcm5U into (S)-mchm5U, and corroborated the notion that the two ALKBH8 enzymatic domains can function independently (Fig. 2e; see Fig. 2c for samples not treated with enzyme; Supplementary Fig. S8). The latter was further substantiated by the observation that the oxygenase and MT functions of ALKBH8 could be selectively inhibited by the Fe(II) chelator EDTA and the MT inhibitor S-adenosylhomocysteine, respectively (Supplementary Fig. S8). Moreover, ALKBH8-Ox, but not a mutant with putative Fe(II) coordinating residues mutated, displayed the expected Fe(II)-dependent ability to convert 2OG to succinate (Supplementary Fig. S9), supporting the notion that ALKBH8 is a 2OG/Fe(II) oxygenase.
Modification status of  from the Alkbh8-targeted mice
from the Alkbh8-targeted mice
As our results strongly indicated that  is the substrate of the ALKBH8 oxygenase, we also specifically investigated the modification status of this isoacceptor from the different gene-targeted mice. WT
is the substrate of the ALKBH8 oxygenase, we also specifically investigated the modification status of this isoacceptor from the different gene-targeted mice. WT  contained not only the expected (S)-mchm5U modification, but also some chm5U (Fig. 2f, Supplementary Fig. S10), possibly due to non-enzymatic hydrolysis of the labile ester bond in (S)-mchm5U, as previously reported23. In agreement with previous findings22, Alkbh8−/−
contained not only the expected (S)-mchm5U modification, but also some chm5U (Fig. 2f, Supplementary Fig. S10), possibly due to non-enzymatic hydrolysis of the labile ester bond in (S)-mchm5U, as previously reported23. In agreement with previous findings22, Alkbh8−/−  contained ncm5U and cm5U, whereas KI(Ox+)
contained ncm5U and cm5U, whereas KI(Ox+)  contained nchm5U and chm5U in addition, indicating that ALKBH8-Ox, in the absence of its cognate mcm5U substrate can hydroxylate cm5U and ncm5U, albeit inefficiently. Moreover, KI(MT+)
contained nchm5U and chm5U in addition, indicating that ALKBH8-Ox, in the absence of its cognate mcm5U substrate can hydroxylate cm5U and ncm5U, albeit inefficiently. Moreover, KI(MT+)  displayed the expected accumulation of mcm5U. We performed MALDI-TOF mass spectrometry of an RNase T1 digest of the isolated
displayed the expected accumulation of mcm5U. We performed MALDI-TOF mass spectrometry of an RNase T1 digest of the isolated  , to verify its identity and to confirm the presence of mchm5U in its anticodon loop. Clearly, WT
, to verify its identity and to confirm the presence of mchm5U in its anticodon loop. Clearly, WT  contained the expected ~2924.3 Da product corresponding to a (S)-mchm5U-modified anticodon loop (Figs 1d and 2g). Moreover,
contained the expected ~2924.3 Da product corresponding to a (S)-mchm5U-modified anticodon loop (Figs 1d and 2g). Moreover,  from the gene-targeted mice gave rise to alternative additional peaks, agreeing with the altered modification pattern observed by LC–MS/MS.
from the gene-targeted mice gave rise to alternative additional peaks, agreeing with the altered modification pattern observed by LC–MS/MS.
Evolutionary distribution of ALKBH8 and mchm5U modifications
On the basis of sequence homology searches, the ALKBH8 oxygenase appears to be present in most multicellular eukaryotes, such as mammals, bee (Apis mellifera), worm (Caenorhabditis elegans) and plant (Arabidopsis thaliana; Supplementary Fig. S11). Accordingly, we detected (S)-mchm5U in all these organisms (Fig. 3a), but neither in the budding yeast S. cerevisiae nor in the fission yeast Schizosaccharomyces pombe, which both have mcm5U-modified tRNAs, but lack ALKBH8. Notably, (R)-mchm5U was detected only in the mammalian tissues tested (calf and mouse), and here at levels which represented ~350x the detection limit (0.003 (R)-mchm5U per 10,000 unmodified nucleosides). This indicates that if (R)-mchm5U is present in any of the other organisms, the level is at least two orders of magnitude lower than that in mammals. As was observed with the mcm5U-containing KI(MT+) tRNA, the majority of mcm5U in S. cerevisiae or S. pombe tRNA could be converted to (S)-mchm5U by incubation with ALKBH8-Ox (Fig. 3b, Supplementary Fig. S12). Taken together, the observed activity of human ALKBH8-Ox on tRNA from distantly related organisms and the co-occurrence of (S)-mchm5U and ALKBH8 suggest that hydroxylation of mcm5U into (S)-mchm5U is performed by ALKBH8 orthologues in a wide range of eukaryotes.
(a) mchm5U modifications in tRNA from various organisms, investigated by LC–MS/MS. (b) In vitro enzymatic activity of human ALKBH8-Ox on tRNA from the yeast S. cerevisiae (upper panel), and on tRNA from the KI(MT+) mice (lower panel). tRNA was incubated with different amounts of recombinant ALKBH8-Ox, and the levels of mcm5U (closed circles) and (S)-mchm5U (open circles) were analysed by LC–MS/MS.
Wobble uridine modifications and protein translation
In yeast, inactivation of wobble uridine modification enzymes can impair translational decoding, as measured by decreased translation of relevant 'codon-runs'26,27. To investigate the possible effects of the extensive wobble U mis-modification observed in the Alkbh8−/− mice, we generated reporter plasmids containing both Renilla (Rluc; internal reference) and firefly (Fluc) luciferase genes, in which Fluc, but not Rluc, translation depends on in-frame translation with a codon-run of ten identical codons (Fig. 4a). We generated runs of A-ending codons matching the anticodons of several tRNAs containing mcm5U or derivatives, and corresponding non-cognate, G-ending codon-runs as controls. We then determined the Fluc/Rluc ratio on transfection of the constructs into primary fibroblasts from Alkbh8−/− or WT mice (Fig. 4b). Clearly, different A-ending codon-runs affected translation to varying degrees, but the differences between Alkbh8−/− and WT fibroblasts were generally small, showing that the altered wobble U modification pattern of Alkbh8−/− tRNA has a negligible effect on translation efficiency in this experimental system.
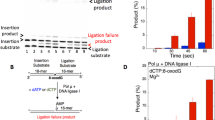
(a) Outline of the reporter construct used. The cytomegalovirus (CMV) promoter drives the expression of an mRNA (blue) encoding the protein elements indicated by boxes (drawing not to scale). (XXA)10 and (XXG)10 indicate runs of ten identical codons that end with A or G, respectively, and that are translated in frame with the firefly luciferase (Fluc) reporter. A Renilla luciferase (Rluc) reporter, which is translated from an internal ribosomal entry site (IRES) and independently of the codon run, serves as an internal reference. (b) Reporter constructs containing the indicated codon runs, encoding the indicated amino acids, were transfected into WT (white bars) and Alkbh8−/− (grey bars) fibroblasts. The activity of the two reporter genes was analysed and used to calculate the efficiency of translation of the codon run. Error bars indicate the standard deviation of triplicate samples.
Discussion
In this study, we have unravelled the biochemical function of the evolutionary conserved ALKBH8 oxygenase. Thus, tRNA modification is included in the broad range of reactions catalysed by 2OG/Fe(II) oxygenases, and the function of the ALKBH proteins is expanded beyond nucleic acid repair. Our data demonstrate how the MT and oxygenase activities of ALKBH8 act in a sequential manner, where the MT generates mcm5U, which in  is further hydroxylated to (S)-mchm5U by the oxygenase. Furthermore, it is demonstrated how selective inactivation of the individual domains of ALKBH8 in vivo severely distorts the modification pattern of wobble uridines, and how the ALKBH8 oxygenase, in the absence of its cognate substrate mcm5U, will hydroxylate ncm5U and cm5U, albeit inefficiently (Fig. 5).
is further hydroxylated to (S)-mchm5U by the oxygenase. Furthermore, it is demonstrated how selective inactivation of the individual domains of ALKBH8 in vivo severely distorts the modification pattern of wobble uridines, and how the ALKBH8 oxygenase, in the absence of its cognate substrate mcm5U, will hydroxylate ncm5U and cm5U, albeit inefficiently (Fig. 5).
 and
and  .
.The model depicts the roles of ALKBH8 and the putative oxygenase OxX in the formation of (S)-mchm5U and (R)-mchm5U in  and
and  , respectively (indicated by solid lines in arrows and borders). Dashed lines in arrows and borders indicate reactions and modifications, respectively, that occur in
, respectively (indicated by solid lines in arrows and borders). Dashed lines in arrows and borders indicate reactions and modifications, respectively, that occur in  and
and  when the ALKBH8 function is perturbed. The colour coding of borders corresponds to that of bar diagrams in Figure 2.
when the ALKBH8 function is perturbed. The colour coding of borders corresponds to that of bar diagrams in Figure 2.
The present study reports the existence of two novel mammalian wobble uridine modifications, (R)- and (S)-mchm5U, and indicates that the evolution of more advanced eukaryotes has been accompanied by an increased complexity of such modifications. Most mcm5U-containing wobble nucleosides contain an additional, wobble restricting 2-thiolation (mcm5s2U), and the corresponding tRNAs decode in the split codon boxes. In contrast,  and
and  , which contain mchm5U, decode in family codon boxes. Therefore, one may speculate that the hydroxylation of mcm5U into mchm5U improves the reading of non-cognate codons; a typical feature of wobble uridine modifications found on family box tRNAs3,4. This ability may be of particular importance in higher eukaryotes, for example, mammals, which consist of a wide diversity of specialized cells, such as elongated neurons. Here, protein translation often occurs at discrete locations28, where low local concentrations of certain isoacceptors could necessitate the extensive use of non-cognate tRNAs in decoding.
, which contain mchm5U, decode in family codon boxes. Therefore, one may speculate that the hydroxylation of mcm5U into mchm5U improves the reading of non-cognate codons; a typical feature of wobble uridine modifications found on family box tRNAs3,4. This ability may be of particular importance in higher eukaryotes, for example, mammals, which consist of a wide diversity of specialized cells, such as elongated neurons. Here, protein translation often occurs at discrete locations28, where low local concentrations of certain isoacceptors could necessitate the extensive use of non-cognate tRNAs in decoding.
Stereoisomers are frequently formed by epimerization, and one could envision (R)-mchm5U being formed by epimerization of (S)-mchm5U, rather than by hydroxylation of mcm5U. However, our presented data strongly argue against this notion, and predict an unknown oxygenase, OxX, for generation of (R)-mchm5U from mcm5U. First, the observation that (R)-mchm5U, but not (S)-mchm5U, is present in the KI(MT+) mice indicates that the former is generated independently of the latter. Second, in the Alkbh8−/− mice, in which the ALKBH8-Ox and ALKBH8-MT functions are both absent, total tRNA contained nchm5U (which is absent from WT tRNA). This presence of nchm5U may be explained by hydroxylation of accumulated ncm5U by the enzyme OxX, which normally catalyses the2 hydroxylation of mcm5U into (R)-mchm5U. Future studies will investigate various oxygenases, including ALKBH proteins of unknown function, as candidates for the OxX function.
Recently, the mammalian Tet proteins (Tet1, Tet2 and Tet3) were shown to be DNA hydroxylases converting epigenetic 5-methylcytosine (5mC) modifications into 5hmC8,29. Interesting analogies exist between ALKBH8 and the Tet proteins, as they all are 2OG/Fe(II) oxygenases acting on nucleic acids, and as they hydroxylate corresponding positions in mcm5U and 5mC, respectively. Conceivably, other ALKBH proteins may also catalyse reactions resulting in hydroxylation (rather than demethylation) of macromolecules.
Methods
Total tRNA and isoacceptor isolation
Total tRNA from S. cerevisiae and calf liver was purchased from Roche and Novagen, respectively. Total tRNA was purified from Mus musculus (liver), A. mellifera (head and thorax of worker bees), C. elegans, A. thaliana and S. pombe by using a RNA/DNA Maxi Kit (QIAGEN). Individual  and
and  isoacceptors were purified from total tRNA using 3′-biotinylated oligonucleotides (Supplementary Table S1), as previously described22.
isoacceptors were purified from total tRNA using 3′-biotinylated oligonucleotides (Supplementary Table S1), as previously described22.
Synthesis of mchm5U nucleoside standards
(R)- and (S)-mchm5U were obtained by a previously reported three-step procedure30. The structure and homogeneity of these nucleosides derivatives were confirmed by nuclear magnetic resonance and mass spectrometry.
LC–MS/MS of nucleosides
The analysis was performed essentially as described22. Briefly, tRNA was enzymatically digested to nucleosides31, which were separated by reverse phase high-performance liquid chromatography on a Zorbax SB-C18 column, using a mobile phase consisting of 0.1% formic acid and a gradient of 5–50% methanol. Online mass spectrometry detection was performed using an Applied Biosystems/MDS Sciex 5000 triple-quadrupole mass spectrometer (Applied Biosystems Sciex) with TurboIonSpray probe operating in positive electrospray ionization mode. The nucleosides were monitored by multiple reaction monitoring using the nucleoside to base ion mass transitions 317.2→185.1 (mcm5U), 333.2→201.1 (mchm5U), 303.2→171.1 (cm5U), 319.2→187.1 (chm5U), 302.2→170.1 (ncm5U), 318.2→186.1 (nchm5U), 268.2→136.1 (A), 244.2→112.1 (C), 284.2→152.2 (G) and 245.2→113.1 (U). Quantification was performed by comparison with pure nucleoside standards run in between the samples. Nucleoside standards for ncm5U and nchm5U were generated from mcm5U and mchm5U as previously described32, whereas cm5U and chm5U were generated by mild alkaline hydrolysis of mcm5U and mchm5U (ref. 22). The fragmentation pattern of the nucleoside standards was in accordance with theoretical predictions, and has been included in Supplementary Figure S13.
MALDI-TOF mass spectrometry
tRNA isoacceptors were digested with RNase T1 (Ambion) and samples prepared for mass spectrometry as previously described22. MALDI (matrix-assisted laser desorption/ionization) mass spectrometry was performed on a Perseptive Voyager STR instrument detecting positive ions in reflector TOF geometry. Spectrum processing was done with the MoverZ software (Genomic Solutions). MALDI tandem mass spectrometry was performed on a Waters QTOF Ultra MALDI instrument (Waters) with sample preparation as described above. The spectra were recorded in positive ion mode and processed with the supplied MassLynx software; additional details have been published previously33.
Gene-targeted mice
Gene-targeting strategy and screening method to verify homologous recombination are presented in Supplementary Figures S2–S5.
Expression of recombinant proteins
Human ALKBH8 (aa 1–664), or its MT domain (ALKBH8-MT; aa 352–664), both in complex with human TRM112, and its AlkB domain (ALKBH8-Ox; aa 1–354) were purified as amino-terminal 6xHis-tagged protein from E. coli as previously described22.
Enzymatic treatment of tRNA
tRNA (5–10 μg) was incubated with recombinant protein (100 pmol, if not indicated otherwise) for 30 min at 37 °C in a 50 μl reaction mixture containing 25 μM S-adenosyl-L-methionine, 50 mM Tris-HCl pH 7.5, 25 mM KCl, 25 mM NH4Ac, 0.5 mM MgCl2, 2 mM ascorbic acid, 100 μM 2-oxoglutarate, 40 μM FeSO4, 10 U RNase inhibitor and then precipitated with 1 volume of isopropanol in the presence of 1 M NH4Ac and 10 μg of glycogen. Incubations with ALKBH8 and ALKBH8-MT were performed in the presence of the co-expressed and co-purified accessory protein TRM112. Pellets were washed with 70% EtOH, dried and dissolved in H2O. The samples were subjected to nucleoside analysis by LC–MS/MS.
Codon-run reporter constructs
Short dsDNA fragments containing NcoI-specific DNA overhangs at both ends and harbouring ten successive, identical Lys, Gln, Arg or Gly codons were generated by annealing oligo pairs AAA10-s/AAA10-as, AAG10-s/AAG10-as, CAA10-s/CAA10-as, CAG10-s/CAG10-as, AGA10-s/AGA10-as, AGG10-s/AGG10-as, GGA10-s/GGA10-as or GGG10-s/GGG10-as (Supplementary Table S1), respectively, and inserted downstream of the translation initiation codon of the firefly luciferase gene in pDualLuc-IRES3 using its NcoI site34. Subsequently, NotI-HindIII fragments from pDualLuc-IRES3 or derivatives containing a codon-run insertion were transferred to pCMV-Script (Stratagene). Following this strategy, pDualLuc-LysAAA10, pDualLuc-LysAAG10, pDualLuc-GlnCAA10, pDualLuc-GlnCAG10, pDualLuc-ArgAGA10, pDualLuc-ArgAGG10, pDualLuc-GlyGGA10 and pDualLuc-GlyGGG10 were constructed (Fig. 4a).
In vivo translation efficiency assay
Primary mouse embryonic fibroblasts were maintained in DMEM (BioWhittaker) supplemented with 15% fetal bovine serum (Saveen & Werner) and 100 U per ml penicillin and streptomycin. Cells were grown to 90% confluency and transfected using a MicroPorator MP-100 (Digital Bio Technology, NanoEnTek). Cells (5×105) were transfected with pDualLuc reporter plasmids using a pulse voltage of 1,350 V, 30 ms pulse width and a 100 μl Gold Tip according to the manufacturer's instructions, and immediately after electroporation transferred to pre-warmed 12-well dishes containing DMEM without antibiotics. Cells were collected at 24 h post-transfection and the intracellular Fluc and Rluc activities were subsequently measured using a Luminometer (Turner Designs TD-20/20), using the Dual-luciferase Reporter Assay according to the manufacturer's protocol (Promega).
Additional information
How to cite this article: van den Born, E. et al. ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA. Nat. Commun. 2:172 doi: 10.1038/ncomms1173 (2011).
References
Agris, P. F. Decoding the genome: a modified view. Nucleic Acids Res. 32, 223–238 (2004).
Sprinzl, M. & Vassilenko, K. S. Compilation of tRNA sequences and sequences of tRNA genes. Nucleic Acids Res. 33, D139–D140 (2005).
Johansson, M. J., Esberg, A., Huang, B., Björk, G. R. & Byström, A. S. Eukaryotic wobble uridine modifications promote a functionally redundant decoding system. Mol. Cell Biol. 28, 3301–3312 (2008).
Nasvall, S. J., Chen, P. & Björk, G. R. The modified wobble nucleoside uridine-5-oxyacetic acid in tRNAPro(cmo5UGG) promotes reading of all four proline codons in vivo. RNA 10, 1662–1673 (2004).
Huang, B., Johansson, M. J. & Byström, A. S. An early step in wobble uridine tRNA modification requires the Elongator complex. RNA 11, 424–436 (2005).
Kalhor, H. R. & Clarke, S. Novel methyltransferase for modified uridine residues at the wobble position of tRNA. Mol. Cell Biol. 23, 9283–9292 (2003).
Loenarz, C. & Schofield, C. J. Expanding chemical biology of 2-oxoglutarate oxygenases. Nat. Chem. Biol. 4, 152–156 (2008).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
Tsukada, Y. et al. Histone demethylation by a family of JmjC domain-containing proteins. Nature 439, 811–816 (2006).
Aas, P. A. et al. Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 421, 859–863 (2003).
Falnes, P. O., Johansen, R. F. & Seeberg, E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419, 178–182 (2002).
Trewick, S. C., Henshaw, T. F., Hausinger, R. P., Lindahl, T. & Sedgwick, B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419, 174–178 (2002).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–1472 (2007).
Kurowski, M. A., Bhagwat, A. S., Papaj, G. & Bujnicki, J. M. Phylogenomic identification of five new human homologs of the DNA repair enzyme AlkB. BMC Genomics 4, 48 (2003).
Duncan, T. et al. Reversal of DNA alkylation damage by two human dioxygenases. Proc. Natl Acad. Sci. USA 99, 16660–16665 (2002).
Westbye, M. P. et al. Human AlkB homolog 1 is a mitochondrial protein that demethylates 3-methylcytosine in DNA and RNA. J. Biol. Chem. 283, 25046–25056 (2008).
Ringvoll, J. et al. Repair deficient mice reveal mABH2 as the primary oxidative demethylase for repairing 1meA and 3meC lesions in DNA. EMBO J. 25, 2189–2198 (2006).
Nordstrand, L. M. et al. Mice lacking Alkbh1 display sex-ratio distortion and unilateral eye defects. PLoS ONE 5, e13827 (2010).
Pan, Z. et al. Impaired placental trophoblast lineage differentiation in Alkbh1(−/−) mice. Dev. Dyn. 237, 316–327 (2008).
Sedgwick, B., Bates, P. A., Paik, J., Jacobs, S. C. & Lindahl, T. Repair of alkylated DNA: recent advances. DNA Repair (Amst.) 6, 429–442 (2007).
Fu, D. et al. Human AlkB homolog ABH8 Is a tRNA methyltransferase required for wobble uridine modification and DNA damage survival. Mol. Cell Biol. 30, 2449–2459 (2010).
Songe-Moller, L. et al. Mammalian ALKBH8 possesses tRNA methyltransferase activity required for the biogenesis of multiple wobble uridine modifications implicated in translational decoding. Mol. Cell Biol. 30, 1814–1827 (2010).
Kawakami, M., Takemura, S., Kondo, T., Fukami, T. & Goto, T. Chemical structure of a new modified nucleoside located in the anticodon of Bombyx mori glycine tRNA2. J. Biochem. 104, 108–111 (1988).
Fu, Y. et al. The AlkB domain of mammalian ABH8 catalyzes hydroxylation of 5-methoxycarbonylmethyluridine at the Wobble position of tRNA. Angew. Chem. Int. Ed. Engl. 49, 8885–8888 (2010).
Andersen, T. E., Kirpekar, F. & Haselmann, K. F. RNA fragmentation in MALDI mass spectrometry studied by H/D-exchange: mechanisms of general applicability to nucleic acids. J. Am. Soc. Mass Spectrom. 17, 1353–1368 (2006).
Begley, U. et al. Trm9-catalyzed tRNA modifications link translation to the DNA damage response. Mol. Cell 28, 860–870 (2007).
Dewez, M. et al. The conserved Wobble uridine tRNA thiolase Ctu1-Ctu2 is required to maintain genome integrity. Proc. Natl Acad. Sci. USA 105, 5459–5464 (2008).
Holt, C. E. & Bullock, S. L. Subcellular mRNA localization in animal cells and why it matters. Science 326, 1212–1216 (2009).
Ito, S. et al. Role of Tet proteins in 5mC to 5hmC conversion, ES-cell self-renewal and inner cell mass specification. Nature 466, 1129–1133 (2010).
Nawrot, B. & Malkiewicz, A. The tRNA 'wobble position' uridines. III. The synthesis of 5-[S-methoxycarbonyl(hydroxy)metyl]uridine and its 2-thio analogue. Nucleosides Nucleotides 8, 1499–1512 (1989).
Crain, P. F. Preparation and enzymatic hydrolysis of DNA and RNA for mass spectrometry. Methods Enzymol. 193, 782–790 (1990).
Fissekis, J. D. & Sweet, F. Synthesis of 5-carboxymethyluridine. A nucleoside from transfer ribonucleic acid. Biochemistry 9, 3136–3142 (1970).
Mengel-Jorgensen, J. & Kirpekar, F. Detection of pseudouridine and other modifications in tRNA by cyanoethylation and MALDI mass spectrometry. Nucleic Acids Res. 30, e135 (2002).
van den Born, E., Posthuma, C. C., Gultyaev, A. P. & Snijder, E. J. Discontinuous subgenomic RNA synthesis in arteriviruses is guided by an RNA hairpin structure located in the genomic leader region. J. Virol. 79, 6312–6324 (2005).
Acknowledgements
We thank H. Nilsen, R. Aamodt and I. Alseth for supplying biological material for tRNA isolation from C. elegans, A. mellifera and S. pombe, respectively. We are grateful to Cesilie G. Castellanos and Hege Wiksen for excellent technical assistance and animal care. We thank GenOway, Lyon, France, The Norwegian Transgenic Center (NTS) and the Centre for Comparative Medicine at Oslo University Hospital HF for the excellent services they provided. We thank Stefan Kernstock and Angela Ho for critical reading of the manuscript. This work was supported by grants from the FRIBIO and FUGE programs in the Research Council of Norway, as well as Grant PNRF-143-AI-1/07 from the Polish–Norwegian Research Fund (to A.K. and P.Ø.F.).
Author information
Authors and Affiliations
Contributions
E.v.d.B, C.B.V., L.S.-M., V.L., G.F.L. and F.K. designed and performed experiments, and analysed data. G.L. synthesized nucleoside standards. H.E.K. and A.M. analysed data. A.K. and P.Ø.F. initiated the study, designed experiments, analysed data and wrote the manuscript. All authors discussed the results and commented on the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S13 and Supplementary Table S1 (PDF 2458 kb)
Rights and permissions
About this article
Cite this article
van den Born, E., Vågbø, C., Songe-Møller, L. et al. ALKBH8-mediated formation of a novel diastereomeric pair of wobble nucleosides in mammalian tRNA. Nat Commun 2, 172 (2011). https://doi.org/10.1038/ncomms1173
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms1173
This article is cited by
-
ALKBH8B, a Putative RNA Demethylase, Plays a Role in the Response of Arabidopsis to Salt Stress and Abscisic Acid
Journal of Plant Biology (2022)
-
Insight into ALKBH8-related intellectual developmental disability based on the first pathogenic missense variant
Human Genetics (2022)
-
The methyltransferase METTL9 mediates pervasive 1-methylhistidine modification in mammalian proteomes
Nature Communications (2021)
-
Reversal of nucleobase methylation by dioxygenases
Nature Chemical Biology (2020)
-
“Too much guts and not enough brains”: (epi)genetic mechanisms and future therapies of Hirschsprung disease — a review
Clinical Epigenetics (2019)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.




 from the Alkbh8-targeted mice
from the Alkbh8-targeted mice

