Abstract
Parasite host switches may trigger disease emergence, but prehistoric host ranges are often unknowable. Lymphatic filariasis and loiasis are major human diseases caused by the insect-borne filarial nematodes Brugia, Wuchereria and Loa. Here we show that the genomes of these nematodes and seven tropical bird lineages exclusively share a novel retrotransposon, AviRTE, resulting from horizontal transfer (HT). AviRTE subfamilies exhibit 83–99% nucleotide identity between genomes, and their phylogenetic distribution, paleobiogeography and invasion times suggest that HTs involved filarial nematodes. The HTs between bird and nematode genomes took place in two pantropical waves, >25–22 million years ago (Myr ago) involving the Brugia/Wuchereria lineage and >20–17 Myr ago involving the Loa lineage. Contrary to the expectation from the mammal-dominated host range of filarial nematodes, we hypothesize that these major human pathogens may have independently evolved from bird endoparasites that formerly infected the global breadth of avian biodiversity.
Similar content being viewed by others
Introduction
Horizontal transfer (HT) of genetic material has widely shaped eukaryotic genomes1 and may often coincide with endosymbiosis2 or parasitism3. A growing body of evidence suggests that host–parasite relationships are gateways for the HT of transposable elements (TEs), genomic parasites that are unable to leave the cell by their own means. Such exchange of TEs across cellular and organismal boundaries appears to be particularly common in DNA transposons4,5,6, whereas it is rather rare in retrotransposons, which exhibit RNA intermediates7,8,9. The only known case of widespread HT of non-retroviral retrotransposons is that of BovB, a family of retrotransposon-like elements (RTEs) present in the genomes of various mammals and lizards/snakes10,11, as well as ticks that parasitize them11. Further evidence exists for a single HT of CR1 retrotransposons between distant groups of butterflies12. However, despite the recent sequencing of many genomes of birds13 and filarial nematodes14,15,16,17, no evidence of HT exists in the evolutionary history of these species-rich taxa.
Here we report widespread retrotransposon HT between endoparasitic nematodes and the breadth of avian biodiversity in the tropics. Surprisingly, these filarial nematodes are extant endoparasites of humans, and cause lymphatic filariasis and loiasis, which affect ∼170 million people14,16. We reconstruct the timing and biogeography of these prehistoric host–parasite associations as witnessed by HT events, and thereby propose a novel scenario for the deep origins of two major human diseases.
Results
A previously undetected transposon from birds and nematodes
We describe AviRTE, a novel family of long interspersed elements (LINEs), from bird and nematode genomes. AviRTE belongs to the RTE superfamily, is distantly related to BovB (Fig. 1), and even more distantly related to known nematode RTEs (Supplementary Fig. 1). Instead, AviRTE groups within a diverse set of RTEs from aquatic or semi-aquatic animals (Fig. 1c, Supplementary Fig. 1). Many of these related and recognizable RTE families are from crocodilians and turtles, a pattern that may reflect the low evolutionary rate of these genomes, which are rich in ancient repeats18. We initially detected fragments of AviRTE in restriction site-associated DNA (RAD) sequences of a Zimmerius flycatcher (Tyrannidae) genome and in BLASTN searches of sequences from other Tyrannidae in GenBank (Supplementary Data 1). These hits are unlikely to be the result of contamination and instead constitute actual TE insertions, because we were able to ascertain orthologous genomic loci of AviRTE presence/absence among multiple species of birds. For example, the ornithine decarboxylase gene exhibits an intronic AviRTE insertion (nested within a 13-bp target site duplication) in some suboscine birds, and an empty insertion site in others (Supplementary Fig. 2). We then examined by BLASTN 48 recently published avian genomes13, a wide range of nematode genomes14,15,16,17, VectorBase’s insect and tick genomes19, and GenBank’s nucleotide and genome collection (including mammalian genomes). We also survey-sequenced the genomes of three hummingbirds and two additional suboscine passerines. In addition to these screenings, we complemented our taxon sampling by targeted PCR of genomic DNA from various bird species (Supplementary Fig. 3; Supplementary Data 1).
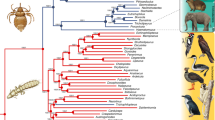
(a,b) Schematic illustration of structural diversity of (a) autonomous AviRTE and (b) non-autonomous SINEs mobilized by AviRTE. The SINEs consist of a promoter-bearing head (yellow) and a bipartite tail that is derived from part of the 5′ UTR (blue) and the full 3′ UTR (red). Typical RNA polymerase III-transcribed SINEs21 were identified within suboscine passerines, where the manakin and tyrant flycatcher lineages share ManaSINE1 with tRNA-Glu gene-derived promoters, whereas the antbird lineage exhibits GymnSINE with 5S-rRNA gene-derived promoters. Furthermore, we detected two emergences of potential SINEs with a GC-rich head including a 5′-GGCCCCGG-3′ motif as a potential protein-binding site62; one in the hornbill lineage (BuceSINE) and one in the manakin lineage (ManaSINE2). Another peculiar SINE, MeloSINE, emerged in the budgerigar lineage and exhibits a head derived from the 3′ portion of the 28S-rRNA gene, a configuration similar to a novel SINE recently discovered in mammals63. (c) Phylogeny (RAxML, GTRCAT model, 1,000 bootstrap replicates, bootstrap values ⩾50% shown) of 370 nucleotide consensus sequences from superfamily RTE (incl. AviRTE and additional BLASTN hits) suggests that AviRTE (red) is distantly related to BovB (orange) and more closely related to RTEs from aquatic animals. RTE subfamilies are in green letters for crocodilians and turtles, in light blue letters for other aquatic vertebrates and in dark blue for aquatic invertebrates. A colour-coded distribution of host taxa across the entire RTE phylogeny is shown in Supplementary Fig. 1.
Horizontal transposon transfer between birds and nematodes
We detected autonomous copies of AviRTE in seven monophyletic clades of birds and two clades of nematodes, but not their respective sister groups (Supplementary Data 1), providing the first evidence for HT in birds and filarial nematodes20. Copy numbers range from 141 to 8,306 copies in avian and 273 to 859 copies in nematode genomes (Supplementary Data 1–2). Wherever possible, we reconstructed the respective consensus sequence, revealing a mean overall nucleotide distance of 0.101 substitutions per site between the full-length consensus sequences derived from bird and nematode genomes (Table 1). The high sequence similarity is not restricted to the ∼3.2-kb-long open reading frame, but is also present across the 5′ and 3′ untranslated regions (UTRs) which are ∼800 bp and ∼40 bp in size, respectively. Notably, we also detected evidence for parallel evolution (Supplementary Fig. 4) of non-autonomous, short interspersed elements (SINEs) that are mobilized by the enzymatic machinery of AviRTE LINEs (Fig. 1b, Supplementary Data 1). All these SINEs share a bipartite tail consisting of fragments of the 5′ and 3′ UTRs of AviRTE (Fig. 1b), yet have different promoter-bearing heads (see legend of Fig. 1b). Altogether, the diversity of non-autonomous elements mobilized by AviRTE is the result of lineage-specific SINE emergences (Supplementary Fig. 4) and surpasses the known diversity of SINEs mobilized by the distantly related BovB family21.
The distribution and timing of transposon invasions
We then studied the phylogenetic distribution and temporal activity of AviRTE retrotransposition across a dated genome-scale phylogeny of birds22. Among the 48 key bird representatives sampled in this phylogeny, AviRTE is present in 7 lineages that span the breadth of avian biodiversity (Fig. 2a). The relatively low sequence divergence between copies (Fig. 2b) and the absence from outgroup genomes of any sequence with even the slightest resemblance to AviRTE (Supplementary Data 1) suggest that this TE family was acquired via HTs long after the Neoaves radiation at the Cretaceous–Paleogene boundary22 (Fig. 2a). Given the evidence for very recent AviRTE retrotransposition in some birds (for example, hornbill; Fig. 2b), we reanalysed all genomes for the presence of full-length AviRTE copies and identified zero to six of such elements per genome (Supplementary Data 1). However, all of these copies exhibit multiple frameshifts and premature stop codons (Supplementary Data 3), suggesting that there are no intact ‘master genes’ of AviRTE in the sampled genome assemblies.
(a) Phylogenetic distribution of AviRTE mapped on simplified chronograms of all major avian clades (high-ranking taxon names in grey letters)22 and a subtree of filarial nematodes27. Together with our dates of AviRTE retrotranspositional activities, this reveals that HT events (red or purple dashed circles denoting minimum and maximum estimates) occurred long after the respective early diversifications of Neoaves and filarial nematodes, putatively in one Oligocene wave (orange) and one Miocene wave (yellow). Dates for AviRTE retrotransposition are either based on a lineage-specific substitution rate of fourfold degenerate sites from the respective bird13 (red colour), or a neutral substitution rate from the nematode Pristionchus pacificus23 (purple colour). For comparison, HT dates based on the mean neutral substitution rate from Caenorhabditis spp.26 are also shown (grey dashed circles). Minimum estimates for genome invasions and extinctions are the respective start and end points of red or purple lines, respectively, and correspond to the 95% interval of AviRTE retrotranspositional activity measured on the scale of pairwise divergence to consensus (Supplementary Data 2). Also shown are the 99% intervals (light red or light purple lines) as maximum estimates for genome invasions and extinctions, and the mean of activity (tick mark). (b,c), Landscape plots of AviRTE divergence (red; co-mobilized SINEs in blue) in (b) avian and (c) nematode genomes illustrate per-genome retrotranspositional activity on a relative time axis. The avian plots are in the same order as the corresponding taxa in the avian phylogeny of panel a. We note that all nematode AviRTE plots except the one of Loa loa are highly similar in terms of copy numbers (Supplementary Data 1) and mean divergences (Supplementary Data 2), possibly resulting from a single TE invasion of the germline genome of the Brugia/Wuchereria ancestor.
We then conservatively inferred minimum times of HTs by applying lineage-specific substitution rates of seven bird species13 (derived from dated branch lengths of the corresponding phylogeny22) to the upper boundary of the 95% interval of divergences between copies of AviRTE (see Methods). These minimum estimates suggest two temporally distinct bursts of invasions (t-test, P=0.0006), the first wave >25.0 to >23.6 Myr ago among hummingbirds, psittacid parrots and hornbills (Fig. 2a), and the second wave >20.2 to >17.7 Myr ago in tinamous, suboscine passerines, mesites and trogons. Consistent with the hypothesis of HT involving nematodes, the two waves of HT in birds are temporally compatible with the dates inferred when considering the per-genome divergences of AviRTE copies in nematode genomes (Fig. 2c) under a neutral substitution rate23 and a generation time of 90 days24. Accordingly, genome invasions are inferred to have taken place >21.2 Myr ago in the ancestor of Brugia spp./Wuchereria bancrofti, the causative agents for lymphatic filariasis, and >16.8 Myr ago in the ancestor of Loa loa, the causative agent for loiasis. In contrast to the aforementioned lineage-specific substitution rates of birds, the nematode HT dates are based on the neutral substitution rate of a mutation accumulation line from a different nematode, Pristionchus pacificus23, because such rates are unavailable for filarial nematodes. Although Weller et al.23 suggested the rate in Pristionchus to be representative for nematodes, we emphasize that molecular dating of nematodes is notoriously difficult due to differences in life style and a virtually non-existent fossil record25, and our nematode dates of AviRTE transfers should therefore be treated with caution. Nevertheless, these nematode HT dates, and dates derived from the slightly lower neutral substitution rates of Caenorhabditis species26 both suggest temporally distinct invasions in the Brugia/Wuchereria lineage and the Loa lineage (Fig. 2a, Supplementary Data 2). We also note that the Pristionchus-based minimum dates are temporally compatible with the split of the Brugia/Wuchereria/Loa lineage from Acanthocheilonema viteae and other AviRTE-free outgroups (Fig. 2a, Supplementary Data 1) in an independently dated nematode phylogeny27. Furthermore, grouping the HT dates from the Brugia/Wuchereria lineage in the first and L. loa in the second of the aforementioned waves of HT again suggests that AviRTE transfers occurred in two distinct bursts (t-test, P=0.0002). The two independent genome invasions of nematodes are surprising given that these two lineages are closely related within filarial nematodes27,28, but in agreement with differences between the shapes of the AviRTE divergence landscapes of the Loa lineage and the Brugia/Wuchereria lineage (Fig. 2c, Supplementary Data 1–2). Finally, the very short internodes in the AviRTE phylogeny indicate a rapid succession of HTs in birds and nematodes (Fig. 3c), and there is phylogenetic evidence that these two bursts of HT are discrete. More precisely, AviRTE subfamilies of the second wave form a monophyletic group nested within the first wave, and the nematode AviRTEs group with avian AviRTEs of similar minimum invasion dates (Fig. 3b).
(a) Diverse geographic origins of seven representatives of AviRTE-bearing birds illustrated on a paleogeographic map of the Earth 20 Myr ago. The Mollweide projection map is copyright to Ronald Blakey (used with permission). (b) Paleobiogeography of the AviRTE phylogeny (see panel c) inferred via statistical dispersal-vicariance (S-DIVA) analysis. We used avifaunal regions sensu Ericson35. For nodes marked with an asterisk, only those areas which received >33% of the ancestral area distribution on this node are shown. Strikingly, the two waves of HT events (cf., Fig. 2a) are not only separated temporally (genome invasion dates of the first and second waves of HT are shown in orange and yellow letters, respectively), but also phylogenetically and biogeographically. (c) The species tree of avian22 and nematode27 hosts (left) is highly incongruent with the AviRTE phylogeny (right; RAxML, GTRCAT model, 1,000 bootstrap replicates, bootstrap values ⩾50% shown) and the assumption of nine HT events (red asterisks) is required to reconcile the two topologies. The AviRTE phylogeny is rooted to several more or less closely related outgroups (Supplementary Fig. 4; cf., Fig. 1c); however, only the closest outgroup is shown for comparability of ingroup internode lengths. Green branches are topologically identical between both trees, implying vertical transmission in species-rich avian lineages. Orange and yellow colours indicate the Oligocene and Miocene waves of HT events, respectively.
The paleobiogeography of transposon invasions
Although the AviRTE-bearing lineages of birds span the entirety of the avian Tree of Life, it is striking that all of these mainly occur in tropical regions29. They include typical Neotropical avifauna such as hummingbirds and tinamous, the Madagascan endemic mesites and members of more widespread tropical bird assemblages (Fig. 3a,b). It was recently noted that genome-scale dating of birds yields much lower divergence estimates than other molecular studies22,30,31, which explains why our inferred avian HT dates postdate, for example, previous dates for the diversifications of suboscine passerines32 and psittacid parrots33. However, irrespective of this discrepancy between absolute dates, our dense taxon sampling suggests that HTs occurred in the respective ancestors of Psittacidae (psittacid parrots), Suboscines (suboscine passerines) and Trochilidae (hummingbirds); and potentially Bucerotidae (hornbills), Mesitornithidae (mesites), Tinamidae (tinamous) and Trogonidae (trogons) (Supplementary Data 1, Fig. 3c). These relative dates predate the onsets of lineage-specific diversification and thus permit paleobiogeographic inferences. We reconstructed the ancestral areas of HTs by assuming a pantropical distribution of filarial nematodes and considering existing evidence for the paleobiogeography of the respective avian clades. Passerines and parrots are more closely related than previously thought22,34 and diversified in Australasia33,35 (but see ref. 30). However, suboscine passerines likely originated in the Neotropics30,32, while the biogeographic origins of hornbills and trogons are unknown35. We find a Neotropical origin of AviRTE is likely given the deep branching of hummingbirds in the AviRTE phylogeny (Fig. 3b), and that the remainder of the first wave of HT (Fig. 2a) occurred across all tropical regions except Madagascar (see Methods and Supplementary Fig. 5 for geographically less constrained analyses). In contrast, the second wave of HT (Fig. 2a) includes Madagascar and took place predominantly in the Neotropics. Altogether, these results suggest that the paleobiogeography of AviRTE transfers occurred on a global, pantropical scale. It is further worth noting that the respective phylogenetic positions of AviRTEs from Brugia spp./W. bancrofti and L. loa coincide with the aforementioned temporal similarities in genome invasion dates. We therefore propose that the ancestor of Brugia/Wuchereria was involved in the first burst of AviRTE transfers, while the Loa ancestor was part of the second wave of HT. In addition, direct comparison of the host species tree with the AviRTE tree (Fig. 3c) suggests that the differences in phylogenetic topology are most parsimoniously explained by nine HT events. However, the TE relationships within hummingbirds, psittacid parrots and suboscine passerines appear to be the result of vertical inheritance, respectively, suggesting that AviRTE shaped the genome evolution of these species-rich lineages by persistence of retrotranspositional activity across their diversifications (Supplementary Data 2).
Possible vectors for HT
This study is the first to report HT of TEs involving the genomes of birds or filarial nematodes. Our results provide phylogenetic, paleobiogeographic and temporal evidence that the endoparasitic Brugia/Wuchereria and Loa lineages each were involved in AviRTE transfer. These nematodes presently have a near-pantropical distribution and are transmitted by mosquitoes14,27 and deerflies16, respectively. Although most birds are capable of flight, their dispersal has been historically rather limited across avifaunal boundaries, especially in the tropics29,35. We therefore suggest that the two bursts of pantropical exchanges of AviRTE between five avifaunal regions were catalysed by the pantropical dispersal potential of filarial nematodes via their dipteran vectors. While it is conceivable that AviRTE transfer occurred directly from blood-sucking dipterans to birds, we find that AviRTE is absent from available dipteran genomes sequences19 and endoparasitic interaction between insect-borne nematodes and birds may be a more plausible platform for such rampant HT. Irrespective of the macroscopic vectors for HT, however, it remains mysterious as to how exactly TEs move from one germline genome into another, with potential candidates being naked RNA or viruses8. Alternatively, intracellular Wolbachia bacteria are plausible cell-penetrating vectors8 and infect many filarial nematodes, yet they are absent in L. loa16. It is further worth noting that successful HT is much more complex than the mere infiltration of a new host cell by transposon DNA or retrotransposon RNA. For ancestrally transferred AviRTEs to be visible in extant genomes, a full-length retrotransposon RNA has to colonize a new germline genome, retrotranspose into a genomic environment which permits retrotranspositional activity as an intact AviRTE ‘master gene’ and drift to fixation in the host population. It is therefore likely that the nine events of successful HT and germline infiltration, reconstructed here from dozens of sampled animal genomes, is but a small fraction of the actual number of prehistoric AviRTE exchanges between birds and filarial nematodes.
Discussion
Our study supports the notion that host–endoparasite interactions are prone to episodic gene exchange, including ‘selfish genes’ such as AviRTE and other TEs, a process that passes on genetic material as ‘public goods’36 among unrelated organisms. We demonstrate that HTs bear witness of long-extinct organismal interactions between birds and nematodes, although it remains undetermined whether the interactions leading to bird–nematode HTs were direct or indirect. The causative agents of lymphatic filariasis and loiasis infect humans as their adult host14,16 and many other filarial nematodes are known to only infect mammals, possibly the result of an ancestral mammalian host range27. The absence of AviRTE in mammalian genomes despite extensive bird–nematode HT seems to challenge this view. The two waves of HT spanned the avian Tree of Life and involved rapid movement among all five avifaunal regions of the tropics, which is puzzling given that tropical landbirds have limited propensities for inter-continental dispersal35. We thus hypothesize that the Brugia/Wuchereria and Loa lineages were Oligocene/Miocene parasites of tropical birds and dispersed pantropically through their dipteran vectors. This may explain the aforementioned complex paleobiogeography of HTs. Such a scenario requires that the nematode lineages underwent two subsequent host switches to humans or their hominid ancestors, likely after AviRTE ceased retrotranspositional activity in filarial nematodes and thus lost its potential for HT into hominid genomes. Our indirect evidence for ancient interactions between birds and the ancestors of the causative agents of lymphatic filariasis and loiasis raises the possibility that these widespread human pathogens may have independently evolved from prehistorically ubiquitous bird endoparasites. We anticipate that exploring the neglected biodiversity of extant bird-infecting nematodes will add further support to this hypothesis.
Methods
In silico screening
We initially detected a putatively RTE-mobilized SINE (later termed ‘ManaSINE1’) in RAD sequences of a Neotropical Zimmerius flycatcher37. The SINE sequence was BLASTN38 searched against a budgerigar39 repeat library that we had generated de novo using Repeatmodeler version 1.0.5 (http://www.repeatmasker.org/RepeatModeler.html). This led to the discovery of a nearly full-length consensus sequence of an autonomous RTE which we termed ‘AviRTE’. We then used the budgerigar AviRTE fragment as query for all subsequent BLASTN screens (cutoff e-value 1e–10) of animal genome assemblies and nucleotide sequences available in GenBank40, including the genomes of 48 birds13 and 2 additional parrots41,42. Furthermore, our screenings comprised all insect and tick genomes in VectorBase19, and all filarial nematode genomes in WormBase15,17. We made sure that, for each of the bird and nematode clades exhibiting AviRTE, we also sampled the closest relatives as outgroups (Supplementary Data 1).
In vitro screening
Our taxon sampling was complemented by species where genome or survey sequences were unavailable (Supplementary Data 1). We sampled these using a short PCR that amplifies a 126-bp region from the conserved 5′ UTR of AviRTE. PCR parameters were 40 cycles of 94 °C for 20 s, 53 °C for 45 s and 68 °C for 60 s, followed by final elongation for 120 s at 68 °C. We used the primers AviRTEint-F/R (5′- CCTGAGGACTTCACTGTCACC -3′+5′- CTTCAAGCCTGTGCAGTGG -3′) and interpreted the absence of an amplicon as an indication of genomic absence of AviRTE (Supplementary Fig. 3). In the case of Pitta moluccensis, we additionally confirmed the AviRTE presence by direct Sanger sequencing of the PCR amplicon (Supplementary Data 4). Finally, we were able to amplify the full length of AviRTE in Gymnopithys rufigula with four overlapping PCR amplicons and subsequent Sanger sequencing of four clones per amplicon, permitting the generation of a consensus sequence. PCR parameters were an initial denaturation for 120 s at 94 °C, 35 cycles of 94 °C for 30 s, 50/54 °C for 30 s and 72 °C for 80 s, followed by final elongation for 300 s at 72 °C. Purified PCR products were cloned into Escherichia coli JM09 cells using the pGEM-T Vector, followed by PCR amplification via standard M13 primers. The four primer pairs were AviRTEfull-1F/R (5′- TCGTGGGGAAAGAGCTTG -3′+5′- AATACAATCGGAATGACCTGTC -3′), AviRTEfull-2F/R (5′- AGGCATCTCTCAGGAGTTGG -3′+5′- CATAGAATCCTCTGTGGTCACC -3′), AviRTEfull-3F/R (5′- CAAGTGGTGGATCAACCTAGC -3′+5′- TGATTTAGGGTCTTGGTGTGG -3′), and AviRTEfull-4F/R (5′- CCTATTCAATCTAAGGCGACTG -3′+5′- ATCATCATGGCTTGGCTTC -3′).
Whole-genome survey sequencing
We obtained whole-genome survey sequences via paired-end sequencing on the Illumina HiSeq 2000 platform (100-bp reads). The insert size was 300 bp and the final coverage ∼6 × for the two suboscines. For the three hummingbirds, the insert sizes ranged from 275 to 450 bp and the final coverage was ∼0.1 × . Genomic presence of AviRTE was ascertained by BLASTN screens (cutoff e-value 1e–10). In the case of Oreotrochilus melanogaster and Zimmerius chrysops, we were able to infer complete and near-complete AviRTE consensus sequences, respectively.
Consensus sequences
Majority-rule consensus sequences were generated manually from each AviRTE-bearing host genome assembly. We used standard procedures43,44 to reconstruct full-length AviRTEs via BLASTN and extension by re-BLASTN searches to overcome incomplete 5′ and 3′ ends. For each of these multiple rounds of BLASTN searches, multiple sequence alignments of BLASTN hits were constructed using MAFFT45,46 version 7 (E-INS-i, http://mafft.cbrc.jp/alignment/server/index.html). The resultant consensus sequences span the full length of AviRTE or co-mobilized SINEs, in rare cases with ‘N’ nucleotides in ambiguous regions that resisted reconstruction. Full-length consensus sequences of AviRTEs and co-mobilized SINEs were submitted to Repbase (http://www.girinst.org/repbase/index.html) (see also Supplementary Data 5).
Phylogenetic analyses
We automatically aligned consensus sequences using MAFFT and then manually realigned ambiguous regions. For the AviRTE phylogenies (Fig. 3c, Supplementary Fig. 4), this was sufficient for generating nucleotide sequence alignments across the full length of AviRTE (Supplementary Data 6). On the other hand, the nucleotide sequence alignment for the phylogeny of all RTE subfamilies present in RepBase, additional GenBank BLASTn hits and all AviRTE subfamilies (Fig. 1c, Supplementary Fig. 1) contained many ambiguities and poorly aligned regions that were removed using Gblocks47 version 0.91b. We chose standard parameters in the Gblocks webserver (http://molevol.cmima.csic.es/castresana/Gblocks_server.html) for less stringent selection of alignment positions (that is, smaller final blocks, gap positions within the final blocks, less strict flanking positions), yielding a 429-bp high-confidence alignment (Supplementary Data 7) from the original 23,637 bp. Note that the low number of retained alignment positions reflects the fact that very distant RTE subfamilies were included, such as those from angiosperm plants. We excluded sequences that aligned poorly at the nucleotide level or comprised <200 bp of the filtered alignment, reducing the total amount of sequences in the RTE superfamily alignment from 444 to 370. We then conducted all phylogenetic analyses under maximum likelihood in RAxML48 version 8.1.11 (GTRCAT model, 1,000 bootstrap inferences) on the CIPRES Science Gateway49 (https://www.phylo.org/portal2/login!input.action). All phylogenetic trees are available in Newick format (Supplementary Data 8).
Distance and dating analyses
Pairwise nucleotide distances between AviRTE consensus sequences (Table 1) were calculated in MEGA6 (ref. 50) under the Kimura 2-parameter model51 with uniform rates among sites and pairwise deletion of gaps/missing data.
We then annotated the genomic copies of AviRTE using RepeatMasker version 3.3.0 (http://www.repeatmasker.org/RMDownload.html) with a custom repeat library for each of the AviRTE-bearing genome assemblies. This library contained the conspecific AviRTE consensus (and, if present, co-mobilized SINE consensus sequences). In the cases where this sequence was incomplete and contained ‘N’ residues, we instead used the full-length consensus sequence from the most closely related host genome. We subsequently calculated per-copy distances to consensus in the calcDivergenceFromAlign.pl script included in the RepeatMasker program package (Kimura 2-parameter model, excluding CpG sites) and plotted these as AviRTE divergence landscapes (Fig. 2b,c). Such a divergence distribution reflects the retrotranspositional activity of AviRTE on a relative time scale per genome. However, we hypothesize that its high-divergence and low-divergence extrema, which usually comprise merely <100 bp per divergence bin (Supplementary Data 2), may in fact arise from genomic outliers in substitution rates, such as conserved or hypervariable regions. In addition, it is plausible that some of the very short, high-divergence AviRTE fragments result from spurious hits to random non-AviRTE sequence during the BLAST-based RepeatMasker annotation. We therefore considered the boundaries of 95% of the distribution as suitable conservative estimates for the onset (latest point of genome invasion) and end (earliest point of extinction) of AviRTE retrotransposition, and the 99% interval as the maximum duration (Supplementary Data 2). Absolute dates were inferred by dividing these divergence values by two times the substitution rate (see Supplementary Data 2 for more details). For the seven birds, we used the fourfold degenerate site substitution rate derived from the respective bird genome13,52. Given the lack of substitution rates from filarial nematodes, we considered the neutral substitution rates of different nematodes, P. pacificus23 and Caenorhabditis species26, under the assumption of a generation time of 90 days for filarial nematodes24. Finally, we plotted the inferred AviRTE retrotranspositional activities on the dated genome-scale tree from the study by Jarvis et al.22,53 (Fig. 2a).
Biogeographic analyses
We used the S-DIVA method54 for biogeographic reconstruction of the site of HT occurrence under the assumption that extant bird species usually occur in less than two avifaunal regions35. Thus, analyses were done in RASP55 using standard parameters and allowing a maximum of two areas per node. Given the current cosmopolitan distribution of nematodes and their great age compared with birds56, we assumed a pantropical distribution of the Loa and Brugia/Wuchereria lineages at the time of HT. For birds, we considered existing evidence for the paleobiogeography of the respective avian clades to infer the areas each lineage inhabited at the time of the HT of AviRTE. Because there is no certainty about the paleobiogeography of each avian lineage, we carried out three different analyses at various levels of conservatism to infer the main location of HT.
First, we considered the widest possible distribution of each bird lineage at the time of HT (Supplementary Fig. 5a). In the absence of a detailed avian fossil record57, this meant that the area of occurrence of most bird lineages would be equated to their present-day distribution, such as hummingbirds (Trochilidae) in the Neotropics; psittacid parrots (Psittacidae) in Australasia, Africa, Indomalaya and the Neotropics; tinamous (Tinamidae) in the Neotropics; mesites (Mesitornithidae) in Madagascar; and trogons (Trogonidae) in Africa, Indomalaya and the Neotropics. We made an exception for the paleotropical hornbills (Bucerotidae) and the pantropical suboscine passerines (Suboscines): both lineages occur in Australasia, but do so only marginally with one and five species, respectively, all of which are known to be of fairly recent Indomalayan descent58,59. Therefore, hornbills were only coded for Africa and Indomalaya, whereas suboscines were only coded for Africa, Indomalaya and the Neotropics in this analysis.
Second, we repeated the first analysis but additionally included areas in which lineages may no longer be present now but are thought to have occurred around the time of HT based on the fossil record (Supplementary Fig. 5b). Given the poor avian fossil record57, this only changed the area designation for a single lineage, the hummingbirds, which are presently distributed only in the New World60 but are known from early-Oligocene Old World fossils61. Hence, in this analysis we coded hummingbirds for the Neotropics, Africa and Indomalaya.
Finally, in our main analysis (Fig. 3b) we only included regions thought to be the areas of occurrence of each respective bird clade roughly at the time of HT based on the current literature. For lineages in which the area of occurrence at the time of HT could not be further narrowed down in comparison to present-day distribution (that is trogons, mesites, tinamous, hornbills and hummingbirds), the same areas as in the first analysis (Supplementary Fig. 5a) were used. However, for psittacid parrots (Psittacidae), we used Australasia as the area of occurrence based on two considerations: (i) there is ample phylogenetic evidence on the Australasian origin of most deep parrot lineages33,35; and (ii) the internal topology of psittacid AviRTE subfamilies strongly suggests that the HT occurred into the last common ancestor of Psittacidae before the divergence of this family into Neotropical, African and Australasian clades. In the same way, we used the Neotropics as the area of occurrence for suboscines based on two similar considerations: (i) the Neotropics are the most likely ancestral area of suboscines35,58; and (ii) the internal topology of suboscine AviRTE subfamilies, as well as their presence in both major suboscine clades strongly suggest that the HT occurred into their last common ancestor before the break-up into Old World and New World suboscines.
Additional information
Accession codes: Raw survey sequence data have been deposited in the Sequence Read Archive (SRA) with accession codes SRS1259545 to SRS1259546, and SRS1303252 to SRS1303254.
How to cite this article: Suh, A. et al. Ancient horizontal transfers of retrotransposons between birds and ancestors of human pathogenic nematodes. Nat. Commun. 7:11396 doi: 10.1038/ncomms11396 (2016).
Accession codes
References
Keeling, P. J. & Palmer, J. D. Horizontal gene transfer in eukaryotic evolution. Nat. Rev. Genet. 9, 605–618 (2008) .
Hotopp, J. C. D. et al. Widespread lateral gene transfer from intracellular bacteria to multicellular eukaryotes. Science 317, 1753–1756 (2007) .
Mower, J. P., Stefanovic, S., Young, G. J. & Palmer, J. D. Plant genetics: Gene transfer from parasitic to host plants. Nature 432, 165–166 (2004) .
Pace, J. K., Gilbert, C., Clark, M. S. & Feschotte, C. Repeated horizontal transfer of a DNA transposon in mammals and other tetrapods. Proc. Natl Acad. Sci. USA 105, 17023–17028 (2008) .
Gilbert, C., Schaack, S., Pace, J. K., Brindley, P. J. & Feschotte, C. A role for host-parasite interactions in the horizontal transfer of transposons across phyla. Nature 464, 1347–1350 (2010) .
Houck, M., Clark, J., Peterson, K. & Kidwell, M. Possible horizontal transfer of Drosophila genes by the mite Proctolaelaps regalis. Science 253, 1125–1128 (1991) .
Ivancevic, A. M., Walsh, A. M., Kortschak, R. D. & Adelson, D. L. Jumping the fine LINE between species: Horizontal transfer of transposable elements in animals catalyses genome evolution. Bioessays 35, 1071–1082 (2013) .
Schaack, S., Gilbert, C. & Feschotte, C. Promiscuous DNA: horizontal transfer of transposable elements and why it matters for eukaryotic evolution. Trends Ecol. Evol. 25, 537–546 (2010) .
Silva, J., Loreto, E. & Clark, J. Factors that affect the horizontal transfer of transposable elements. Curr. Issues Mol. Biol. 6, 57–71 (2004) .
Kordiš, D. & Gubenšek, F. Unusual horizontal transfer of a long interspersed nuclear element between distant vertebrate classes. Proc. Natl Acad. Sci. USA 95, 10704–10709 (1998) .
Walsh, A. M., Kortschak, R. D., Gardner, M. G., Bertozzi, T. & Adelson, D. L. Widespread horizontal transfer of retrotransposons. Proc. Natl Acad. Sci. USA 110, 1012–1016 (2013) .
Sormacheva, I. et al. Vertical evolution and horizontal transfer of CR1 non-LTR retrotransposons and Tc1/mariner DNA transposons in Lepidoptera species. Mol. Biol. Evol. 29, 3685–3702 (2012) .
Zhang, G. et al. Comparative genomics reveals insights into avian genome evolution and adaptation. Science 346, 1311–1320 (2014) .
Ghedin, E. et al. Draft genome of the filarial nematode parasite Brugia malayi. Science 317, 1756–1760 (2007) .
Harris, T. W. et al. WormBase: a comprehensive resource for nematode research. Nucleic Acids Res. 38, D463–D467 (2010) .
Desjardins, C. A. et al. Genomics of Loa loa, a Wolbachia-free filarial parasite of humans. Nat. Genet. 45, 495–500 (2013) .
Howe, K. L. et al. WormBase 2016: expanding to enable helminth genomic research. Nucleic Acids Res. 44, D774–D780 (2016) .
Green, R. E. et al. Three crocodilian genomes reveal ancestral patterns of evolution among archosaurs. Science 346, 1335 (2014) .
Lawson, D. et al. VectorBase: a data resource for invertebrate vector genomics. Nucleic Acids Res. 37, D583–D587 (2009) .
Wallau, G. L., Ortiz, M. F. & Loreto, E. L. S. Horizontal transposon transfer in Eukarya: detection, bias, and perspectives. Genome Biol. Evol. 4, 801–811 (2012) .
Gogolevsky, K. P., Vassetzky, N. S. & Kramerov, D. A. Bov-B-mobilized SINEs in vertebrate genomes. Gene 407, 75–85 (2008) .
Jarvis, E. D. et al. Whole genome analyses resolve the early branches in the tree of life of modern birds. Science 346, 1320–1331 (2014) .
Weller, A. M., Rödelsperger, C., Eberhardt, G., Molnar, R. I. & Sommer, R. J. Opposing forces of A/T-biased mutations and G/C-biased gene conversions shape the genome of the nematode Pristionchus pacificus. Genetics 196, 1145–1152 (2014) .
Thomas, J. A., Welch, J. J., Lanfear, R. & Bromham, L. A generation time effect on the rate of molecular evolution in invertebrates. Mol. Biol. Evol. 27, 1173–1180 (2010) .
Blaxter, M. Nematodes (Nematoda). in The Timetree of Life eds Hedges S. B., Kumar S. 247–250Oxford Univ. Press (2009) .
Denver, D. R. et al. Variation in base-substitution mutation in experimental and natural lineages of Caenorhabditis nematodes. Genome Biol. Evol. 4, 513–522 (2012) .
Small, S. T., Tisch, D. J. & Zimmerman, P. A. Molecular epidemiology, phylogeny and evolution of the filarial nematode Wuchereria bancrofti. Infect. Genet. Evol. 28, 33–43 (2014) .
Blaxter, M. & Koutsovoulos, G. The evolution of parasitism in Nematoda. Parasitology 142, S26–S39 (2015) .
Newton, I. The Speciation and Biogeography of Birds Academic Press (2003) .
Claramunt, S. & Cracraft, J. A new time tree reveals Earth history’s imprint on the evolution of modern birds. Sci. Adv. 1, e1501005 (2015) .
Prum, R. O. et al. A comprehensive phylogeny of birds (Aves) using targeted next-generation DNA sequencing. Nature 526, 569–573 (2015) .
Ericson, P. G. et al. A Gondwanan origin of passerine birds supported by DNA sequences of the endemic New Zealand wrens. Proc. R. Soc. B 269, 235–241 (2002) .
Schweizer, M., Seehausen, O. & Hertwig, S. T. Macroevolutionary patterns in the diversification of parrots: effects of climate change, geological events and key innovations. J. Biogeogr. 38, 2176–2194 (2011) .
Suh, A. et al. Mesozoic retroposons reveal parrots as the closest living relatives of passerine birds. Nat. Commun. 2, 443 (2011) .
Ericson, P. G. P. Evolution of terrestrial birds in three continents: biogeography and parallel radiations. J. Biogeogr. 39, 813–824 (2012) .
McInerney, J., Pisani, D., Bapteste, E. & O'Connell, M. The public goods hypothesis for the evolution of life on Earth. Biol. Direct 6, 41 (2011) .
Rheindt, F. E., Fujita, M. K., Wilton, P. R. & Edwards, S. V. Introgression and phenotypic assimilation in Zimmerius flycatchers (Tyrannidae): population genetic and phylogenetic inferences from genome-wide SNPs. Syst. Biol. 63, 134–152 (2014) .
Altschul, S. F., Gish, W., Miller, W., Myers, E. W. & Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 215, 403–410 (1990) .
Ganapathy, G. et al. High-coverage sequencing and annotated assemblies of the budgerigar genome. GigaSci 3, 1–9 (2014) .
Benson, D. A. et al. GenBank. Nucleic Acids Res. 41, D36–D42 (2013) .
Oleksyk, T. K. et al. A locally funded Puerto Rican parrot (Amazona vittata) genome sequencing project increases avian data and advances young researcher education. Gigascience 1, 14–14 (2012) .
Seabury, C. M. et al. A multi-platform draft de novo genome assembly and comparative analysis for the scarlet macaw (Ara macao). PLoS ONE 8, e62415 (2013) .
Dasmahapatra, K. K. et al. Butterfly genome reveals promiscuous exchange of mimicry adaptations among species. Nature 487, 94–98 (2012) .
Suh, A. et al. Multiple lineages of ancient CR1 retroposons shaped the early genome evolution of amniotes. Genome Biol. Evol. 7, 205–217 (2015) .
Katoh, K. & Toh, H. Recent developments in the MAFFT multiple sequence alignment program. Brief Bioinform. 9, 286–298 (2008) .
Katoh, K. & Standley, D. M. MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol. Biol. Evol. 30, 772–780 (2013) .
Castresana, J. Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol. Biol. Evol. 17, 540–552 (2000) .
Stamatakis, A., Hoover, P. & Rougemont, J. A rapid bootstrap algorithm for the RAxML web servers. Syst. Biol. 75, 758–771 (2008) .
Miller, M. A., Pfeiffer, W. & Schwartz, T. in Proceedings of the Gateway Computing Environments Workshop (GCE), 1–8 (New Orleans, LA, USA, 2010) .
Tamura, K., Stecher, G., Peterson, D., Filipski, A. & Kumar, S. MEGA6: molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 30, 2725–2729 (2013) .
Kimura, M. A simple method for estimating evolutionary rates of base substitutions through comparative studies of nucleotide sequences. J. Mol. Evol. 16, 111–120 (1980) .
Zhang, G. et al. Comparative genomic data of the Avian Phylogenomics Project. Gigascience 3, 26 (2014) .
Jarvis, E. D. et al. Phylogenomic analyses data of the Avian Phylogenomics Project. Gigascience 4, 4 (2015) .
Yu, Y., Harris, A. J. & He, X. S-DIVA (Statistical Dispersal-Vicariance Analysis): a tool for inferring biogeographic histories. Mol. Phylogenet. Evol. 56, 848–850 (2010) .
Yu, Y., Harris, A. J., Blair, C. & He, X. RASP (Reconstruct Ancestral State in Phylogenies): a tool for historical biogeography. Mol. Phylogenet. Evol. 87, 46–49 (2015) .
Aguinaldo, A. M. A. et al. Evidence for a clade of nematodes, arthropods and other moulting animals. Nature 387, 489–493 (1997) .
Olson, S. L. The fossil record of birds. in Avian Biology eds Famer D. S., King J. R. 79–238Academic Press (1985) .
Irestedt, M., Ohlson, J. I., Zuccon, D., Källersjö, M. & Ericson, P. G. P. Nuclear DNA from old collections of avian study skins reveals the evolutionary history of the Old World suboscines (Aves, Passeriformes). Zool. Scripta 35, 567–580 (2006) .
Viseshakul, N. et al. A phylogeny of frugivorous hornbills linked to the evolution of Indian plants within Asian rainforests. J. Evol. Biol. 24, 1533–1545 (2011) .
McGuire, J. A. et al. Molecular phylogenetics and the diversification of hummingbirds. Curr. Biol. 24, 910–916 (2014) .
Mayr, G. Old World fossil record of modern-type hummingbirds. Science 304, 861–864 (2004) .
Yang, Y. et al. Regulation of a disintegrin and metalloprotease-33 expression by transforming growth factor-β. Am. J. Respir. Cell Mol. Biol. 46, 633–640 (2012) .
Longo, M. S., Brown, J. D., Zhang, C., O'Neill, M. J. & O'Neill, R. J. Identification of a recently active mammalian SINE derived from ribosomal RNA. Genome Biol. Evol. 7, 775–788 (2015) .
Acknowledgements
We thank the Genetic Resources Collection (Instituto Nacional de Pesquisas da Amazônia) for providing suboscine samples for survey sequencing, Christoph Bleidorn (Museo Nacional de Ciencias Naturales, Madrid), Cédric Feschotte (the University of Utah, Salt Lake City), Aurélie Kapusta (University of Utah, Salt Lake City), Ruxandra I. Molnar (Uppsala University), and Claudia C. Weber (Temple University, Philadelphia) for helpful discussions, Anja Bodenheim (University of Bonn) for lab assistance and Cai Li (China National Genebank, BGI-Shenzhen) for providing substitution rates. We are grateful to Jon Fjeldså (University of Copenhagen) for generating the bird paintings, Luk Cox & Idoya Lahortiga (somersault18:24) for providing the nematode image, and to Ronald Blakey (Colorado Plateau Geosystems, Inc.) for providing the paleogeographic map. A.S. acknowledges computational resources provided by the Swedish National Infrastructure for Computing (SNIC) through Uppsala Multidisciplinary Center for Advanced Computational Science (UPPMAX), access to which was kindly made possible by Hans Ellegren (Uppsala University). J.Me. was supported by a stipend from the Brazilian Science Funding Agency CAPES. F.E.R. was funded by the National University of Singapore (NUS) through Faculty of Science and Department of Biological Sciences grants (WBS R-154-000-570-133 and R-154-000-583-651). C.C.W. was funded by grants from the United States National Science Foundation (DEB-0543556 and DEB-1146491). S.V.E was funded by grants from the United States National Science Foundation.
Author information
Authors and Affiliations
Contributions
A.S. conceived the study. A.S., C.C.W. and F.E.R. designed in vitro experiments. J.Me., K.R.S, L.P., M.G. and A.W. conducted in vitro experiments. A.S. designed and conducted in silico experiments, and analysed the data. C.C.W., L.P., J.A.M., J.Mu., S.V.E. and F.E.R. provided materials/reagents/analysis tools. A.S. wrote the paper with input from C.C.W., S.V.E. and F.E.R.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures 1-5 and Supplementary References. (PDF 385 kb)
Supplementary Data 1
Taxon sampling of birds and nematodes used for the detection of AviRTE. (XLS 42 kb)
Supplementary Data 2
Activity periods of AviRTE derived from per-copy distances in bird and nematode genomes. (XLS 50 kb)
Supplementary Data 3
Amino acid alignment of AviRTE ORF from subfamily consensus sequences and single full-length copies. In addition to premature stop codons, all of the single full-length copies contain frameshifts, which was compensated by the insertion of alignment gaps prior to ORF translation. (TXT 31 kb)
Supplementary Data 4
Alignment of the AviRTE 5' UTR region amplified by short PCR. Primer sequences and the Old World suboscine Pitta moluccensis are included. (TXT 3 kb)
Supplementary Data 5
Fasta-formatted consensus sequences of AviRTE subfamilies and co-mobilized SINEs. (TXT 58 kb)
Supplementary Data 6
Nucleotide sequence alignment of AviRTE subfamilies, co-mobilized SINEs, and selected RTE outgroups. This alignment was used for phylogenetic analyses shown in Supplementary Fig. 4 and, after exclusion of SINEs, in Fig. 3c. (TXT 104 kb)
Supplementary Data 7
Nucleotide sequence alignment of AviRTE subfamilies, all RTE families present in RepBase, and additional GenBank BLASTn hits. This alignment was used for the phylogenetic analysis shown in Fig. 1c and Supplementary Fig. 1. (TXT 167 kb)
Supplementary Data 8
Newick-formatted phylogenetic trees from (a) Fig. 1c and Supplementary Fig. 1, (b) Fig. 3c, and (c) Supplementary Fig. 4. (TXT 32 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Suh, A., Witt, C., Menger, J. et al. Ancient horizontal transfers of retrotransposons between birds and ancestors of human pathogenic nematodes. Nat Commun 7, 11396 (2016). https://doi.org/10.1038/ncomms11396
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms11396
This article is cited by
-
Evidence of venom factor-like in crocodilians
Organisms Diversity & Evolution (2023)
-
Inner ear biomechanics reveals a Late Triassic origin for mammalian endothermy
Nature (2022)
-
The genome of the thin-necked bladder worm Taenia hydatigena reveals evolutionary strategies for helminth survival
Communications Biology (2021)
-
Hagfish genome reveals parallel evolution of 7SL RNA-derived SINEs
Mobile DNA (2020)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.