Abstract
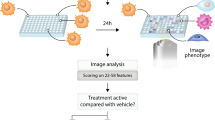
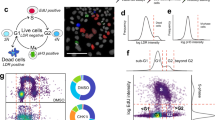
We present a strategy for identifying off-target effects and hidden phenotypes of drugs by directly probing biochemical pathways that underlie therapeutic or toxic mechanisms in intact, living cells. High-content protein-fragment complementation assays (PCAs) were constructed with synthetic fragments of a mutant fluorescent protein ('Venus', EYFP or both), allowing us to measure spatial and temporal changes in protein complexes in response to drugs that activate or inhibit particular pathways. One hundred and seven different drugs from six therapeutic areas were screened against 49 different PCA reporters for ten cellular processes. This strategy reproduced known structure-function relationships and also predicted 'hidden,' potent antiproliferative activities for four drugs with novel mechanisms of action, including disruption of mitochondrial membrane potential. A simple algorithm identified a 25-assay panel that was highly predictive of antiproliferative activity, and the predictive power of this approach was confirmed with cross-validation tests. This study suggests a strategy for therapeutic discovery that identifies novel, unpredicted mechanisms of drug action and thereby enhances the productivity of drug-discovery research.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Stoughton, R.B. & Friend, S.H. How molecular profiling could revolutionize drug discovery. Nat. Rev. Drug Discov. 4, 345–350 (2005).
Tian, Q. et al. Integrated genomic and proteomic analyses of gene expression in Mammalian cells. Mol. Cell. Proteomics 3, 960–969 (2004).
Miklos, G.L. & Maleszka, R. Microarray reality checks in the context of a complex disease. Nat. Biotechnol. 22, 615–621 (2004).
Ghosh, I., Hamilton, A.D. & Regan, L. Antiparallel leucine zipper-directed protein reassembly: application to the green fluorescent protein. J. Am. Chem. Soc. 122, 5658–5659 (2000).
Michnick, S.W., Remy, I., Campbell-Valois, F.X., Vallee-Belisle, A. & Pelletier, J.N. Detection of protein-protein interactions by protein fragment complementation strategies. Methods Enzymol. 328, 208–230 (2000).
Remy, I. & Michnick, S.W. Visualization of biochemical networks in living cells. Proc. Natl. Acad. Sci. USA 98, 7678–7683 (2001).
Nagai, T. et al. A variant of yellow fluorescent protein with fast and efficient maturation for cell-biological applications. Nat. Biotechnol. 20, 87–90 (2002).
Yu, H. et al. Measuring drug action in the cellular context using protein-fragment complementation assays. Assay Drug Dev. Technol. 1, 811–822 (2003).
Remy, I. & Michnick, S.W. Mapping biochemical networks with protein-fragment complementation assays. Methods Mol. Biol. 261, 411–426 (2004).
Remy, I. & Michnick, S.W. A cDNA library functional screening strategy based on fluorescent protein complementation assays to identify novel components of signaling pathways. Methods 32, 381–388 (2004).
Remy, I. & Michnick, S.W. Regulation of apoptosis by the Ft1 protein, a new modulator of protein kinase B/Akt. Mol. Cell. Biol. 24, 1493–1504 (2004).
Remy, I., Montmarquette, A. & Michnick, S.W. PKB/Akt modulates TGF-beta signalling through a direct interaction with Smad3. Nat. Cell Biol. 6, 358–365 (2004).
Nyfeler, B., Michnick, S.W. & Hauri, H.P. Capturing protein interactions in the secretory pathway of living cells. Proc. Natl. Acad. Sci. USA 102, 6350–6355 (2005).
Hu, C.D., Chinenov, Y. & Kerppola, T.K. Visualization of interactions among bZIP and Rel family proteins in living cells using bimolecular fluorescence complementation. Mol. Cell 9, 789–798 (2002).
Fang, D. & Kerppola, T.K. Ubiquitin-mediated fluorescence complementation reveals that Jun ubiquitinated by Itch/AIP4 is localized to lysosomes. Proc. Natl. Acad. Sci. USA 101, 14782–14787 (2004).
Zhang, J., Ferguson, S.S., Barak, L.S., Menard, L. & Caron, M.G. Dynamin and beta-arrestin reveal distinct mechanisms for G protein-coupled receptor internalization. J. Biol. Chem. 271, 18302–18305 (1996).
Lin, F.T. et al. Phosphorylation of beta-arrestin2 regulates its function in internalization of beta(2)-adrenergic receptors. Biochemistry 41, 10692–10699 (2002).
Nolte, R.T. et al. Ligand binding and co-activator assembly of the peroxisome proliferator-activated receptor-gamma. Nature 395, 137–143 (1998).
Winter-Vann, A.M. & Casey, P.J. Post-prenylation-processing enzymes as new targets in oncogenesis. Nat. Rev. Cancer 5, 405–412 (2005).
Maki, C.G., Huibregtse, J.M. & Howley, P.M. In vivo ubiquitination and proteasome-mediated degradation of p53(1). Cancer Res. 56, 2649–2654 (1996).
Treier, M., Staszewski, L.M. & Bohmann, D. Ubiquitin-dependent c-Jun degradation in vivo is mediated by the delta domain. Cell 78, 787–798 (1994).
Nefsky, B. & Beach, D. Pub1 acts as an E6-AP-like protein ubiquitin ligase in the degradation of cdc25. EMBO J. 15, 1301–1312 (1996).
Murray, A. Cyclin ubiquitination: the destructive end of mitosis. Cell 81, 149–152 (1995).
Bain, J., McLauchlan, H., Elliott, M. & Cohen, P. The specificities of protein kinase inhibitors: an update. Biochem. J. 371, 199–204 (2003).
Fotsis, T. et al. Flavonoids, dietary-derived inhibitors of cell proliferation and in vitro angiogenesis. Cancer Res. 57, 2916–2921 (1997).
Hoessel, R. et al. Indirubin, the active constituent of a Chinese antileukaemia medicine, inhibits cyclin-dependent kinases. Nat. Cell Biol. 1, 60–67 (1999).
Benzaquen, L.R., Brugnara, C., Byers, H.R., Gatton-Celli, S. & Halperin, J.A. Clotrimazole inhibits cell proliferation in vitro and in vivo. Nat. Med. 1, 534–540 (1995).
Neckers, L., Schulte, T.W. & Mimnaugh, E. Geldanamycin as a potential anti-cancer agent: its molecular target and biochemical activity. Invest. New Drugs 17, 361–373 (1999).
Tuynder, M. et al. Translationally controlled tumor protein is a target of tumor reversion. Proc. Natl. Acad. Sci. USA 101, 15364–15369 (2004).
Keblys, M. et al. The effects of the Penicillium mycotoxins citrinin, cyclopiazonic acid, ochratoxin A, patulin, penicillic acid, and roquefortine C on in vitro proliferation of porcine lymphocytes. Mycopathologia 158, 317–324 (2004).
Seigle-Murandi, F., Steiman, R., Krivobok, S., Beriel, H. & Benoit-Guyod, J.L. Antitumor activity of patulin and structural analogs. Pharmazie 47, 288–291 (1992).
Weinbach, E.C. & Garbus, J. Mechanism of action of reagents that uncouple oxidative phosphorylation. Nature 221, 1016–1018 (1969).
Burghardt, R.C. et al. Patulin-induced cellular toxicity: a vital fluorescence study. Toxicol. Appl. Pharmacol. 112, 235–244 (1992).
Minnich, A., Tian, N., Byan, L. & Bilder, G. A potent PPARα agonist stimulates mitochondrial fatty acid beta-oxidation in liver and skeletal muscle. Am. J. Physiol. Endocrinol. Metab. 280, E270–E279 (2001).
Froyland, L. et al. Mitochondrion is the principal target for nutritional and pharmacological control of triglyceride metabolism. J. Lipid Res. 38, 1851–1858 (1997).
Zhou, S. & Wallace, K.B. The effect of peroxisome proliferators on mitochondrial bioenergetics. Toxicol. Sci. 48, 82–89 (1999).
Nam, S. et al. Indirubin derivatives inhibit Stat3 signaling and induce apoptosis in human cancer cells. Proc. Natl. Acad. Sci. USA 102, 5998–6003 (2005).
Wald, N. & Cuckle, H. Reporting the assessment of screening and diagnostic tests. Br. J. Obstet. Gynaecol. 96, 389–396 (1989).
Parsonnet, J. & Axon, A.T. Principles of screening and surveillance. Am. J. Gastroenterol. 91, 847–849 (1996).
Hanahan, D. & Weinberg, R.A. The hallmarks of cancer. Cell 100, 57–70 (2000).
Galarneau, A., Primeau, M., Trudeau, L.E. & Michnick, S.W. β-Lactamase protein fragment complementation assays as in vivo and in vitro sensors of protein protein interactions. Nat. Biotechnol. 20, 619–622 (2002).
Leveson-Gower, D.B., Michnick, S.W. & Ling, V. Detection of TAP family dimerizations by an in vivo assay in mammalian cells. Biochemistry 43, 14257–14264 (2004).
Pelletier, J.N., Campbell-Valois, F. & Michnick, S.W. Oligomerization domain-directed reassembly of active dihydrofolate reductase from rationally designed fragments. Proc. Natl. Acad. Sci. USA 95, 12141–12146 (1998).
Subramaniam, R., Desveaux, D., Spickler, C., Michnick, S.W. & Brisson, N. Direct visualization of protein interactions in plant cells. Nat. Biotechnol. 19, 769–772 (2001).
Benton, R., Sachse, S., Michnick, S.W. & Vosshall, L.B. Atypical membrane topology and heteromeric function of Drosophila odorant receptors in vivo. PLoS Biol. 4, e20 (2006).
Ward, J. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 58, 236–244 (1963).
Reers, M., Smith, T.W. & Chen, L.B. J-aggregate formation of a carbocyanine as a quantitative fluorescent indicator of membrane potential. Biochemistry 30, 4480–4486 (1991).
Acknowledgements
The authors thank E. Smith, M. West and the molecular biology staff at Odyssey Thera for excellent technical support. S.W.M. is the Canada Research Chair in Integrative Genomics. This manuscript is dedicated to the memory of Anthony V. Carrano.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
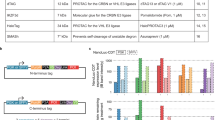
Supplementary Fig. 1
Enhanced sensitivity of IFP PCA versus EYFP (enhanced YFP) (PDF 61 kb)
Supplementary Fig. 2
Details of Figure 2 matrix showing compound activity versus PCA at different time points and additional treatments (PDF 166 kb)
Supplementary Fig. 3
Assay response matrix from Figure 5c (PDF 51 kb)
Supplementary Table 1
Assay components and assay conditions (PDF 88 kb)
Supplementary Table 2
Drugs and their screening doses (PDF 54 kb)
Supplementary Table 3
Effects of compounds on proliferation of PC3 Cells (PDF 57 kb)
Supplementary Table 4
Assay statistics and positive predictive values (PPVs) for the top 25 assays, and results of leave-n-out analyses (PDF 78 kb)
Rights and permissions
About this article
Cite this article
MacDonald, M., Lamerdin, J., Owens, S. et al. Identifying off-target effects and hidden phenotypes of drugs in human cells. Nat Chem Biol 2, 329–337 (2006). https://doi.org/10.1038/nchembio790
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio790
This article is cited by
-
Androgen receptor-dependent regulation of metabolism in high grade bladder cancer cells
Scientific Reports (2023)
-
HSP90β promotes osteoclastogenesis by dual-activation of cholesterol synthesis and NF-κB signaling
Cell Death & Differentiation (2023)
-
Chemoresistance Mechanisms in Non-Small Cell Lung Cancer—Opportunities for Drug Repurposing
Applied Biochemistry and Biotechnology (2023)
-
Collateral responses to classical cytotoxic chemotherapies are heterogeneous and sensitivities are sparse
Scientific Reports (2022)
-
Predictive modeling of estrogen receptor agonism, antagonism, and binding activities using machine- and deep-learning approaches
Laboratory Investigation (2021)