Abstract
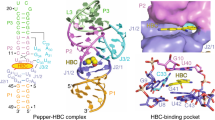
Genetically encoded fluorescent protein tags have revolutionized proteome studies, whereas the lack of intrinsically fluorescent RNAs has hindered transcriptome exploration. Among several RNA–fluorophore complexes that potentially address this problem, RNA Mango has an exceptionally high affinity for its thiazole orange (TO)-derived fluorophore, TO1–Biotin (Kd ∼3 nM), and, in complex with related ligands, it is one of the most redshifted fluorescent macromolecular tags known. To elucidate how this small aptamer exhibits such properties, which make it well suited for studying low-copy cellular RNAs, we determined its 1.7-Å-resolution co-crystal structure. Unexpectedly, the entire ligand, including TO, biotin and the linker connecting them, abuts one of the near-planar faces of the three-tiered G-quadruplex. The two heterocycles of TO are held in place by two loop adenines and form a 45° angle with respect to each other. Minimizing this angle would increase quantum yield and further improve this tool for in vivo RNA visualization.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Dolgosheina, E.V. et al. RNA mango aptamer-fluorophore: a bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 9, 2412–2420 (2014).
Ellington, A.D. & Szostak, J.W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Grate, D. & Wilson, C. Laser-mediated, site-specific inactivation of RNA transcripts. Proc. Natl. Acad. Sci. USA 96, 6131–6136 (1999).
Babendure, J.R., Adams, S.R. & Tsien, R.Y. Aptamers switch on fluorescence of triphenylmethane dyes. J. Am. Chem. Soc. 125, 14716–14717 (2003).
Paige, J.S., Wu, K.Y. & Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Jeng, S.C.Y., Chan, H.H.Y., Booy, E.P., McKenna, S.A. & Unrau, P.J. Fluorophore ligand binding and complex stabilization of the RNA Mango and RNA Spinach aptamers. RNA 22, 1884–1892 (2016).
Day, R.N. & Davidson, M.W. The Fluorescent Protein Revolution (CRC Press, 2014).
Baugh, C., Grate, D. & Wilson, C. 2.8 A crystal structure of the malachite green aptamer. J. Mol. Biol. 301, 117–128 (2000).
Strack, R.L., Disney, M.D. & Jaffrey, S.R. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat-containing RNA. Nat. Methods 10, 1219–1224 (2013).
Wang, P.C. et al. Photochemical properties of Spinach and its use in selective imaging. Chem. Sci. 4, 2865–2873 (2013).
Han, K.Y., Leslie, B.J., Fei, J., Zhang, J. & Ha, T. Understanding the photophysics of the spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. J. Am. Chem. Soc. 135, 19033–19038 (2013).
Escobedo, J.O., Rusin, O., Lim, S. & Strongin, R.M. NIR dyes for bioimaging applications. Curr. Opin. Chem. Biol. 14, 64–70 (2010).
Gellert, M., Lipsett, M.N. & Davies, D.R. Helix formation by guanylic acid. Proc. Natl. Acad. Sci. USA 48, 2013–2018 (1962).
Burge, S., Parkinson, G.N., Hazel, P., Todd, A.K. & Neidle, S. Quadruplex DNA: sequence, topology and structure. Nucleic Acids Res. 34, 5402–5415 (2006).
Warner, K.D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Huang, H. et al. A G-quadruplex-containing RNA activates fluorescence in a GFP-like fluorophore. Nat. Chem. Biol. 10, 686–691 (2014).
Ramesh, A., Wakeman, C.A. & Winkler, W.C. Insights into metalloregulation by M-box riboswitch RNAs via structural analysis of manganese-bound complexes. J. Mol. Biol. 407, 556–570 (2011).
Fiore, J.L. & Nesbitt, D.J. An RNA folding motif: GNRA tetraloop-receptor interactions. Q. Rev. Biophys. 46, 223–264 (2013).
Coonrod, L.A., Lohman, J.R. & Berglund, J.A. Utilizing the GAAA tetraloop/receptor to facilitate crystal packing and determination of the structure of a CUG RNA helix. Biochemistry 51, 8330–8337 (2012).
Michel, F. & Westhof, E. Modelling of the three-dimensional architecture of group I catalytic introns based on comparative sequence analysis. J. Mol. Biol. 216, 585–610 (1990).
Jaeger, L., Michel, F. & Westhof, E. Involvement of a GNRA tetraloop in long-range RNA tertiary interactions. J. Mol. Biol. 236, 1271–1276 (1994).
Costa, M. & Michel, F. Frequent use of the same tertiary motif by self-folding RNAs. EMBO J. 14, 1276–1285 (1995).
Nissen, P., Ippolito, J.A., Ban, N., Moore, P.B. & Steitz, T.A. RNA tertiary interactions in the large ribosomal subunit: the A-minor motif. Proc. Natl. Acad. Sci. USA 98, 4899–4903 (2001).
Howard, F.B. & Miles, H.T. Poly(inosinic acid) helices: essential chelation of alkali metal ions in the axial channel. Biochemistry 21, 6736–6745 (1982).
Williamson, J.R., Raghuraman, M.K. & Cech, T.R. Monovalent cation-induced structure of telomeric DNA: the G-quartet model. Cell 59, 871–880 (1989).
Sen, D. & Gilbert, W. A sodium-potassium switch in the formation of four-stranded G4-DNA. Nature 344, 410–414 (1990).
Mendez, M.A. & Szalai, V.A. Fluorescence of unmodified oligonucleotides: a tool to probe G-quadruplex DNA structure. Biopolymers 91, 841–850 (2009).
Dao, N.T., Haselsberger, R., Michel-Beyerle, M.-E. & Phan, A.T. Following G-quadruplex formation by its intrinsic fluorescence. FEBS Lett. 585, 3969–3977 (2011).
Kwok, C.K., Sherlock, M.E. & Bevilacqua, P.C. Effect of loop sequence and loop length on the intrinsic fluorescence of G-quadruplexes. Biochemistry 52, 3019–3021 (2013).
Molinaro, M. & Tinoco, I. Jr. Use of ultra stable UNCG tetraloop hairpins to fold RNA structures: thermodynamic and spectroscopic applications. Nucleic Acids Res. 23, 3056–3063 (1995).
Sherlock, M.E. et al. Steady-state and time-resolved studies into the origin of the intrinsic fluorescence of G-quadruplexes. J. Phys. Chem. B 120, 5146–5158 (2016).
Phan, A.T. et al. Structure-function studies of FMRP RGG peptide recognition of an RNA duplex-quadruplex junction. Nat. Struct. Mol. Biol. 18, 796–804 (2011).
Vasilyev, N. et al. Crystal structure reveals specific recognition of a G-quadruplex RNA by a β-turn in the RGG motif of FMRP. Proc. Natl. Acad. Sci. USA 112, E5391–E5400 (2015).
Nix, J., Sussman, D. & Wilson, C. The 1.3 Å crystal structure of a biotin-binding pseudoknot and the basis for RNA molecular recognition. J. Mol. Biol. 296, 1235–1244 (2000).
Hendrickson, W.A. et al. Crystal structure of core streptavidin determined from multiwavelength anomalous diffraction of synchrotron radiation. Proc. Natl. Acad. Sci. USA 86, 2190–2194 (1989).
Weber, P.C., Ohlendorf, D.H., Wendoloski, J.J. & Salemme, F.R. Structural origins of high-affinity biotin binding to streptavidin. Science 243, 85–88 (1989).
Nygren, J., Svanvik, N. & Kubista, M. The interactions between the fluorescent dye thiazole orange and DNA. Biopolymers 46, 39–51 (1998).
Carreon, J.R., Mahon, K.P. Jr. & Kelley, S.O. Thiazole orange-peptide conjugates: sensitivity of DNA binding to chemical structure. Org. Lett. 6, 517–519 (2004).
Leontis, N.B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
Xiao, H., Edwards, T.E. & Ferré-D'Amaré, A.R. Structural basis for specific, high-affinity tetracycline binding by an in vitro evolved aptamer and artificial riboswitch. Chem. Biol. 15, 1125–1137 (2008).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
Sheldrick, G.M. A short history of SHELX. Acta Crystallogr. A 64, 112–122 (2008).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Abrahams, J.P. & Leslie, A.G.W. Methods used in the structure determination of bovine mitochondrial F1 ATPase. Acta Crystallogr. D Biol. Crystallogr. 52, 30–42 (1996).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
DeLano, W.L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002).
Puglisi, J.D. & Tinoco, I. Jr. Absorbance melting curves of RNA. Methods Enzymol. 180, 304–325 (1989).
Acknowledgements
We thank the staff at beamlines 5.0.1 and 5.0.2 of the Advanced Light Source (ALS), Lawrence Berkeley National Laboratory, and beamline 24-ID-C of the Advanced Photon Source (APS), Argonne National Laboratory, for crystallographic data collection; G. Piszczek (Biophysics Core, US National Heart, Lung and Blood Institute, NHLBI, National Institutes of Health (NIH)) for analytical ultracentrifugation and dynamic light scattering; D. Lee and R. Levine (NHLBI) for mass spectrometry; and S. Bachas, M. Chen, C. Fagan, C. Jones, T. Numata, D. Sen, L. Sjekloca, L. Truong, K. Warner and J. Zhang for discussions. This work was partly conducted at the ALS, on the on the Berkeley Center for Structural Biology beamlines, and at the APS, on the NE-CAT beamlines, which are supported by the NIH. Use of the ALS and APS was supported by the US Department of Energy. P.J.U. was supported by an NSERC (Canada) operating grant. This work was supported in part by the intramural program of the NHLBI, NIH.
Author information
Authors and Affiliations
Contributions
P.J.U. and A.R.F.-D. conceived the project; M.W.L.L. performed initial crystallization screens; S.C.Y.J. synthesized ligands; R.J.T., N.A.D. and M.W.L.L. carried out preparative biochemistry; R.J.T. performed crystallization, diffraction data collection, structure determination and refinement; R.J.T., N.A.D., S.S.S.P. and S.C.Y.J. performed structure-guided analyses; R.J.T. and A.R.F.-D. prepared the manuscript with input from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results, Supplementary Tables 1–3 and Supplementary Figures 1–9. (PDF 3521 kb)
Rights and permissions
About this article
Cite this article
Trachman, R., Demeshkina, N., Lau, M. et al. Structural basis for high-affinity fluorophore binding and activation by RNA Mango. Nat Chem Biol 13, 807–813 (2017). https://doi.org/10.1038/nchembio.2392
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.2392
This article is cited by
-
ALS-linked TDP-43 mutations interfere with the recruitment of RNA recognition motifs to G-quadruplex RNA
Scientific Reports (2023)
-
Structure of the transcription open complex of distinct σI factors
Nature Communications (2023)
-
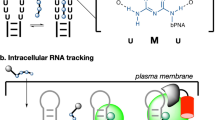
Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA
Nature Communications (2023)
-
Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine
Nature Communications (2021)
-
Structure-based investigation of fluorogenic Pepper aptamer
Nature Chemical Biology (2021)