Abstract
Squash is an RNA aptamer that strongly activates the fluorescence of small-molecule analogs of the fluorophore of green fluorescent protein (GFP). Unlike other fluorogenic aptamers, isolated de novo from random-sequence RNA, Squash was evolved from the bacterial adenine riboswitch to leverage its optimized in vivo folding and stability. We now report the 2.7-Å resolution cocrystal structure of fluorophore-bound Squash, revealing that while the overall fold of the riboswitch is preserved, the architecture of the ligand-binding core is dramatically transformed. Unlike previously characterized aptamers that activate GFP-derived fluorophores, Squash does not harbor a G-quadruplex, sandwiching its fluorophore between a base triple and a noncanonical base quadruple in a largely apolar pocket. The expanded structural core of Squash allows it to recognize unnatural fluorophores that are larger than the simple purine ligand of the parental adenine riboswitch, and suggests that stable RNA scaffolds can tolerate larger variation than has hitherto been appreciated.

This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Trachman, R. J. 3rd & Ferré-D’Amaré, A. R. Tracking RNA with light: selection, structure and design of fluorescence turn-on RNA aptamers. Q. Rev. Biophys. 52, e8 (2019).
Braselmann, E., Rathbun, C., Richards, E. M. & Palmer, A. E. Illuminating RNA biology: tools for imaging RNA in live mammalian cells. Cell Chem. Biol. 27, 891–903 (2020).
Neubacher, S. & Hennig, S. RNA structure and cellular applications of fluorescent light-up aptamers. Angew. Chem. Int. Ed. Engl. 58, 1266–1279 (2019).
Lee, E., Lee, J. M., Kim, H. & Y., K. Fluorogenic aptasensors with small molecules. Chemosensors 9, 54 (2021).
Ellington, A. D. & Szostak, J. W. In vitro selection of RNA molecules that bind specific ligands. Nature 346, 818–822 (1990).
Robertson, D. L. & Joyce, G. F. Selection in vitro of an RNA enzyme that specifically cleaves single-stranded DNA. Nature 344, 467–468 (1990).
Tuerk, C. & Gold, L. Systematic evolution of ligands by exponential enrichment: RNA ligands to bacteriophage T4 DNA polymerase. Science 249, 505–510 (1990).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Autour, A., Westhof, E. & Ryckelynck, M. iSpinach: a fluorogenic RNA aptamer optimized for in vitro applications. Nucleic Acids Res. 44, 2491–2500 (2016).
Trachman, R. J.3rd et al. Structure and functional reselection of the Mango-III fluorogenic RNA aptamer. Nat. Chem. Biol. 15, 472–479 (2019).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Han, K. Y., Leslie, B. J., Fei, J. Y., Zhang, J. C. & Ha, T. Understanding the photophysics of the Spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. J. Am. Chem. Soc. 135, 19033–19038 (2013).
Huang, H. et al. A G-quadruplex-containing RNA activates fluorescence in a GFP-like fluorophore. Nat. Chem. Biol. 10, 686–691 (2014).
Warner, K. D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Warner, K. D. et al. A homodimer interface without base pairs in an RNA mimic of red fluorescent protein. Nat. Chem. Biol. 13, 1195–1201 (2017).
Song, W. et al. Imaging RNA polymerase III transcription using a photostable RNA–fluorophore complex. Nat. Chem. Biol. 13, 1187–1194 (2017).
Mieczkowski, M. et al. Large Stokes shift fluorescence activation in an RNA aptamer by intermolecular proton transfer to guanine. Nat. Commun. 12, 3549 (2021).
Baugh, C., Grate, D. & Wilson, C. 2.8 Å crystal structure of the malachite green aptamer. J. Mol. Biol. 301, 117–128 (2000).
Shelke, S. A. et al. Structural basis for activation of fluorogenic dyes by an RNA aptamer lacking a G-quadruplex motif. Nat. Commun. 9, 4542 (2018).
Banco, M. T. & Ferré-D’Amaré, A. R. The emerging structural complexity of G-quadruplex RNAs. RNA 27, 390–402 (2021).
Dey, S. K. et al. Repurposing an adenine riboswitch into a fluorogenic imaging and sensing tag Nat. Chem. Biol. (in the press). https://doi.org/10.1038/s41589-021-00925-0 (2021).
Serganov, A. et al. Structural basis for discriminative regulation of gene expression by adenine- and guanine-sensing mRNAs. Chem. Biol. 11, 1729–1741 (2004).
Nissen, P., Ippolito, J. A., Ban, N., Moore, P. B. & Steitz, T. A. RNA tertiary interactions in the large ribosomal subunit: the A-minor motif. Proc. Natl Acad. Sci. USA 98, 4899–4903 (2001).
Sjekloća, L. & Ferré-D’Amaré, A. R. Binding between G-quadruplexes at the homodimer interface of the Corn RNA aptamer strongly activates thioflavin T fluorescence. Cell Chem. Biol. 26, 1159–1168 (2019).
Burley, S. K. & Petsko, G. A. Weakly polar interactions in proteins. Adv. Protein Chem. 39, 125–192 (1988).
Giese, M., Albrecht, M. & Rissanen, K. Anion-p interactions with fluoroarenes. Chem. Rev. 115, 8867–8895 (2015).
Dalvit, C. & Vulpetti, A. Weak intermolecular hydrogen bonds with fluorine: detection and implications for enzymatic/chemical reactions, chemical properties, and ligand/protein fluorine NMR screening. Chem. Eur. J. 22, 7592–7601 (2016).
Sykes, B. D., Weingarten, H. I. & Schlessinger, M. J. Fluorotyrosine alkaline phosphatase from Escherichia coli: preparation, properties and fluorine-19 nuclear magnetic resonance spectrum. Proc. Natl Acad. Sci. USA 71, 469–473 (1974).
Porter, E. B., Marcano-Velázquez, J. G. & Batey, R. T. The purine riboswitch as a model system for exploring RNA biology and chemistry. Biochim. Biophys. Acta 1839, 919–930 (2014).
Jeng, S. et al. Fluorogenic aptamers resolve the flexibility of RNA junctions using orientation-dependent FRET. RNA 27, 433–444 (2021).
Fernandez-Millan, P., Autour, A., Ennifar, E., Westhof, E. & Ryckelynck, M. Crystal structure and fluorescence properties of the iSpinach aptamer in complex with DFHBI. RNA 23, 1788–1795 (2017).
Truong, L. & Ferré-D’Amaré, A. R. From fluorescent proteins to fluorogenic RNAs: tools for imaging cellular macromolecules. Protein Sci. 28, 1374–1386 (2019).
Lau, M. W. L. & Ferré-D’Amaré, A. R. In vitro evolution of coenzyme-independent variants from the glmS ribozyme structural scaffold. Methods 106, 76–81 (2016).
Lau, M. W. L., Trachman, R. J. 3rd & Ferré-D’Amaré, A. R. A divalent cation-dependent variant of the glmS ribozyme with stringent Ca2+ selectivity co-opts a preexisting nonspecific metal ion-binding site. RNA 23, 355–364 (2017).
Porter, E. B., Polaski, J. T., Morck, M. M. & Batey, R. T. Recurrent RNA motifs as scaffolds for genetically encodable small-molecule biosensors. Nat. Chem. Biol. 13, 295–301 (2017).
de la Peña, M., Dufour, D. & Gallego, J. Three-way RNA junctions with remote tertiary contacts: a recurrent and highly versatile fold. RNA 15, 1949–1964 (2009).
Jones, C. P. & Ferré-D’Amaré, A. R. Long-range interactions in riboswitch control of gene expression. Annu. Rev. Biophys. 46, 455–481 (2017).
Serganov, A., Polonskaia, A., Phan, A. T., Breaker, R. R. & Patel, D. J. Structural basis for gene regulation by a thiamine pyrophosphate-sensing riboswitch. Nature 441, 1167–1171 (2006).
Thore, S., Leibundgut, M. & Ban, N. Structure of the eukaryotic thiamine pyrophosphate riboswitch with its regulatory ligand. Science 312, 1208–1211 (2006).
Edwards, T. E. & Ferré-D’Amaré, A. R. Crystal structures of the thi-box riboswitch bound to thiamine pyrophosphate analogs reveal adaptive RNA-small molecule recognition. Structure 14, 1459–1468 (2006).
Smith, K. D. et al. Structural basis of ligand binding by a c-di-GMP riboswitch. Nat. Struct. Mol. Biol. 16, 1218–1223 (2009).
Kulshina, N., Baird, N. J. & Ferré-D’Amaré, A. R. Recognition of the bacterial second messenger cyclic diguanylate by its cognate riboswitch. Nat. Struct. Mol. Biol. 16, 1212–1217 (2009).
Xiao, H., Edwards, T. E. & Ferré-D’Amaré, A. R. Structural basis for specific, high-affinity tetracycline binding by an in vitro evolved aptamer and artificial riboswitch. Chem. Biol. 15, 1125–1137 (2008).
Stagno, J. R. et al. Structures of riboswitch RNA reaction states by mix-and-inject XFEL serial crystallography. Nature 541, 242–246 (2017).
Leontis, N. B. & Westhof, E. Geometric nomenclature and classification of RNA base pairs. RNA 7, 499–512 (2001).
DeLano, W. L. The PyMOL Molecular Graphics System (DeLano Scientific, 2002).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Kabsch, W. XDS. Acta Crystallogr. D 66, 125–132 (2010).
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
McCoy, A. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Franke, D. et al. ATSAS 2.8: a comprehensive data analysis suite for small-angle scattering from macromolecular solutions. J. Appl. Crystallogr. 50, 1212–1225 (2017).
Keller, S. et al. High-precision isothermal titration calorimetry with automated peak-shape analysis. Anal. Chem. 84, 5066–5073 (2012).
Zhao, H., Piszczek, G. & Schuck, P. SEDPHAT–a platform for global ITC analysis and global multi-method analysis of molecular interactions. Methods 76, 137–148 (2015).
Acknowledgements
We thank the staff of beamlines 5.0.1 of the Advanced Light Source (ALS), Lawrence Berkeley National Laboratory, and 24-ID-C and 24-ID-E of the Advanced Photon Source (APS), Argonne National Laboratory for crystallographic data collection; the staff of beamline 12-ID-B of APS for SAXS; G. Piszczeck and of the Biophysics Core of the National Heart, Lung and Blood Institute (NHLBI) for fluorescence and ITC; and M. Banco, N. Demeshkina, C. Jones, T. Numata and R. Trachman for discussions. This research used resources of the APS, a US Department of Energy (DOE) Office of Science User Facility operated for the DOE Office of Science by Argonne National Laboratory under contract no. DE-AC02-06CH11357. This work is based upon research conducted at the Northeastern Collaborative Access Team beamlines, which are funded by the National Institute of General Medical Sciences from the National Institutes of Health (NIH P30 GM124165). The Pilatus 6M detector on 24-ID-C beam line is funded by an NIH-ORIP HEI grant (S10 RR029205). This work was supported in part by NIH award R35NS111631 (to S.R.J.) and by the intramural program of the NHLBI, NIH.
Author information
Authors and Affiliations
Contributions
S.R.J. and A.R.F.D’A. initiated the project. L.T. performed fluorescence, crystallographic, SAXS and ITC experiments. S.K.D. and X.L. performed initial biochemistry and synthesized fluorophores. H.K. and N.T. performed NMR experiments. A.R.F.D’A. and L.T. wrote the manuscript with contributions from all authors.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Peer review information Nature Chemical Biology thanks Sven Hennig and other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
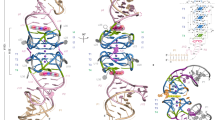
Extended Data Fig. 1 Electron density maps for structures determined in this study.
a, Density-modified SAD electron density map for the Squash-DFHBI-1T complex contoured at 1.5 σ (blue mesh), superimposed on the final refined model. b, Composite simulated annealing-omit 2|Fo | -|Fc | electron density map for the Squash-DFHO complex contoured at 1.5 σ (blue mesh), superimposed on the final refined model.
Extended Data Fig. 2 Small-angle X-ray scattering analysis of Squash.
a, Experimental scattering profile of Squash-DFHBI-1T and scattering profile back-calculated2 using CRYSOL, from the co-crystal structure (Methods). I and q are the intensity and the scattering vector, respectively. b, Scattering profile of Squash in the presence and absence of excess DFHBI-1T. c, Scattering profile of Squash mutant with disrupted L2-L3 kissing loop interactions (MM) in the presence and absence of excess DFHBI-1T.
Extended Data Fig. 3 Eight tiers comprise the Squash aptamer core.
Ball-and-stick representation of each tier (numbered 1-8 in bold numerals) in the Squash core. Panel with DFHBI-1T and G14 (tier 2) shows A69 and A70 from adjacent tiers for reference. Orange dashed lines denote hydrogen bonds. Secondary structure representation corresponds to that in Fig. 1e.
Extended Data Fig. 4 ThT recognition by Squash.
a, Fluorescence of Squash RNA (0.5 μM) and thioflavin T (ThT; 1.0 μM) measured in the absence or the presence of increasing concentrations of DFHO (Methods). b, DFHO fluorescence measured from the same mixtures.
Extended Data Fig. 5 Cartoon representation of orthogonal views of DFHO-bound Squash.
RNA color scheme as in Fig. 1. Metal ions and water molecules omitted for clarity.
Extended Data Fig. 6 Isothermal titration calorimetry.
Triplicate, baseline-corrected thermograms and fits for Squash titrated with a, DFHBI-1T (means ± s.d., n = 3). ΔH and ΔG are 41.0 ± 20.8 nM and -21.3 ± 2.3 kcal/Mol, respectively. b, Triplicate, baseline-corrected thermograms and fits for Squash titrated with a, DFHBI-1E (means ± s.d., n = 3). ΔH and ΔG are 30.3 ± 16.6 and -19.2 ± 0.8 kcal/Mol, respectively.
Extended Data Fig. 7 Fluorescence of DFHBI-1T and DFHBI-1E bound Squash.
Absorbance spectra (dashed lines) were recorded at 503 nm and 492 nm, and emission spectra (solid lines) at 451 nm and 431 nm, respectively. Green lines, Squash with DFHBI-1T; blue lines, Squash with DFHBI-1E.
Extended Data Fig. 8 The adenine riboswitch core is smaller than that of Squash.
a, Ball-and-stick representations of each of the five tiers (numbered 1-5 in bold numerals) that comprise the adenine riboswitch core (PDB ID:1Y26; ref. 3). Orange dashed lines denote hydrogen bonds. Base pairs in the secondary structure are in Leontis-Westhof symbols4. Compare with Supplementary Fig. 3. b, Structure-based sequence alignment of Squash (sq) and the V. vulnificus add adenine riboswitch aptamer domain (ar). Residue numbers correspond to Squash. Color scheme is that of Fig. 1. Asterisks and periods denote sequence identity and gaps needed to account for Squash being longer, respectively. Junction regions J1/2, J2/3 and J3/1 have no clear structural correspondence and the residues in these regions are not aligned.
Extended Data Fig. 9 Orthogonal ligand selectivity of the adenine riboswitch and Squash.
a, Fluorescence activity of the adenine riboswitch in the presence of excess DFHBI-1T, normalized to Squash fluorescence (means ± s.d., n = 3; Methods). Ordinate is discontinuous to accommodate different scale of the two samples. b, Normalized fluorescence of Squash-DFHBI-1T in the presence of varying excess of adenine.
Extended Data Fig. 10 Comparison of Squash and iSpinach recognition of DFHBI-1T.
a, Cartoon representation of the fluorophore binding site of iSpinach (PDB:7L0Z; ref. 5). Orange dashed lines denote hydrogen bonds. Residue numbers correspond to those of Spinach (ref. 6). b, Fluorescence lifetimes (Methods) of DFHBI-1T bound to Squash and iSpinach (green squares and cyan triangles, respectively; means ± s.d., n = 3). Black circles, instrument response function (IRF).
Supplementary information
Supplementary Information
Supplementary Tables 1–3, Extended Data Figs. 1–10, Note and refs.
Rights and permissions
About this article
Cite this article
Truong, L., Kooshapur, H., Dey, S.K. et al. The fluorescent aptamer Squash extensively repurposes the adenine riboswitch fold. Nat Chem Biol 18, 191–198 (2022). https://doi.org/10.1038/s41589-021-00931-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41589-021-00931-2
This article is cited by
-
Fluorogenic CRISPR for genomic DNA imaging
Nature Communications (2024)
-
Co-crystal structures of the fluorogenic aptamer Beetroot show that close homology may not predict similar RNA architecture
Nature Communications (2023)
-
Growing a garden of fluorescent RNAs
Nature Chemical Biology (2022)