Abstract
The most widely used method for intracellular RNA fluorescence labeling is MS2 labeling, which generally relies on the use of multiple protein labels targeted to multiple RNA (MS2) hairpin structures installed on the RNA of interest (ROI). While effective and conveniently applied in cell biology labs, the protein labels add significant mass to the bound RNA, which potentially impacts steric accessibility and native RNA biology. We have previously demonstrated that internal, genetically encoded, uridine-rich internal loops (URILs) comprised of four contiguous UU pairs (8 nt) in RNA may be targeted with minimal structural perturbation by triplex hybridization with 1 kD bifacial peptide nucleic acids (bPNAs). A URIL-targeting strategy for RNA and DNA tracking would avoid the use of cumbersome protein fusion labels and minimize structural alterations to the RNA of interest. Here we show that URIL-targeting fluorogenic bPNA probes in cell media can penetrate cell membranes and effectively label RNAs and RNPs in fixed and live cells. This method, which we call fluorogenic U-rich internal loop (FLURIL) tagging, was internally validated through the use of RNAs bearing both URIL and MS2 labeling sites. Notably, a direct comparison of CRISPR-dCas labeled genomic loci in live U2OS cells revealed that FLURIL-tagged gRNA yielded loci with signal to background up to 7X greater than loci targeted by guide RNA modified with an array of eight MS2 hairpins. Together, these data show that FLURIL tagging provides a versatile scope of intracellular RNA and DNA tracking while maintaining a light molecular footprint and compatibility with existing methods.
Similar content being viewed by others
Introduction
There remains a need for improved and orthogonal methods for localizing and tracking intracellular RNA and DNA molecules in real time1. Despite the existence of many useful RNA imaging strategies, there are limitations on the placement of probe binding sites and concerns regarding the overall structural impact of labeling on the RNA of interest. Modification of internal sites in RNA via a chemical reaction or duplex hybridization can be technically challenging or sequence-limited2,3. While fluorescence in situ (duplex) hybridization4,5 (FISH) with DNA or PNA probes is broadly used for labeling RNAs, FISH is limited to fixed cell experiments. Molecular beacons may be used with unmodified cells and transcripts in live cell labeling but require electroporation or microinjection strategies and have not been as widely adopted as other methods1,6. In addition, duplex hybridization to internal sites destroys existing secondary structures via strand invasion or transforms a single-stranded loop into a new duplex stem where none existed previously4, introducing potentially disruptive steric interactions. Beyond chemical modification and FISH, approaches to intracellular RNA tracking may be roughly divided into small molecule dye binding and protein labels. Dye-binding SELEX-derived aptamers (e.g., Spinach7, Broccoli8, Corn9, Mango10, Pepper11) have been inserted into RNAs of interest for tracking and sensing applications12,13. A variation on this approach exploits RNA binding to dequench dye-quencher conjugates (e.g., SRB214, Gemini-56115, Riboglow16) and can utilize native aptamers to universal quenchers such as cobalamin16; this method may be applied to a wider range of fluorophores without the need for additional aptamer selection. However, all aptamer methods require the insertion of a distinct folded structure within the ROI that has the potential to alter RNA sterics or lifetime. Further, early fluorescent aptamer designs suffered from insufficient brightness, thought to be due to misfolded RNA loops17. In particular, the G-quadruplex motif common to Mango and Spinach aptamers18,19,20 is especially labile to intracellular degradation21 and requires non-native levels of potassium and magnesium8, which may result in deficient intracellular aptamer performance1. Bacteriophage-derived RNA hairpins MS2 and PP7 are native sequences that bind to phage coat proteins MCP (MS2 coat protein) and PCP (PP7 coat protein), respectively. MS2-labeling requires insertion of multiple MS2 hairpins into ROIs and with subsequent RNP imaging via binding of MCP fusion proteins that facilitate fluorescent readout (eg.- fluorescent proteins22,23 (MCP-FPs), MCP-HaloTag24 and MCP-SNAP tag11,25 fusions). Though effective, the use of constitutively fluorescent dyes and FPs generates a significant background signal, requiring multiple (often >20) MS2 hairpins to generate a sufficient signal to noise. Further, these protein labeling systems carry significant steric bulk (>40 kD for a single MCP-FP) with the risk of altered transcript decay22,26,27,28 and inhibited native contacts. Despite these drawbacks, MS2/PP7 labeling remains the most widely used method for RNA tracking. Similarly, CRISPR-Cas/guide RNA complexes29 can be used to track of DNA and RNA targets via fluorescent readout of protein fusions to Cas variants, but also via formation of sterically encumbering protein binding to native RNA30 targets or genomic loci31.
Here, in a method we call fluorogenic U-rich internal loop (FLURIL) tagging, we show that intracellular RNAs bearing an 8-nt (U4xU4) U-rich internal loop (URIL) can be selectively labeled with a fluorogenic bPNA probe via triplex hybridization. The FLURIL system minimizes the introduction of new RNA secondary structures by use of duplex-for-triplex stem replacement, is considerably smaller than typical RNA aptamer or protein labeling systems, and exhibits improved signal-to-background and stability over comparable methods (Supplementary Fig. 3). Prior studies on triazine assembly32,33,34,35 and targeting36,37 led to our development of bifacial peptide nucleic acid (bPNA)34,38,39, which presents synthetic melamine bases40,41 on an α-peptide backbone42,43,44 (Fig. 1). The bPNA family of compounds selectively hybridize with UnxUn internal bulges38,39,45,46 via formation of uracil-melamine-uracil (UMU) base triples, generating triplex stems that can functionally replace native RNA stems45,46,47, tertiary contacts46,47, block protein readthrough48, direct chemistry49, and modulate lncRNA lifetime50. Optimized51 cationic 4 M bPNAs (with four melamine bases, Fig. 2) are cell-permeable49,50 and can bind to structured U4xU4 loops (URILs) while retaining nanomolar affinity45, thus enabling specific intracellular bPNA targeting of URIL-tagged ROIs. via formation of triplex hybrid stems that are similar in mass to a 4 bp RNA duplex. Via URIL-tagging, bPNA hybridization can place prosthetic groups at internal RNA sites without aptamer selection; this manuscript describes fluorogen-modified bPNA binding to the URIL (FLURIL) tag. We demonstrate FLURIL tagging of intracellular RNAs and RNPs in both fixed and live cell contexts.
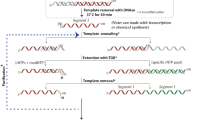
a Triplex hybridization of a U-rich internal loop (URIL) with bPNA (blue) via base triple formation between the melamine base (M) and two uracil bases (inset). b General schematic of labeling strategy described herein. An RNA of interest is engineered to contain a URIL and expressed within the cell, with a fluorogenic bPNA probe introduced via cell culture media. Successful URIL targeting is reported by an increase in emission (green) and confirmed by colocalization with an RBP fusion with a (red) fluorescent protein.
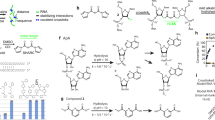
Structures of fluorogenic bPNA probes studied, with thiazole orange (TO) dye and K2M sidechain structure shown inset. Lower case k2M residue denotes D-configuration, and KTO indicates sidechain is acylated with R1 (TO). For TO-K2MAK2M and TO-K2MIK2M, R1 includes a β-alanine spacer between TO and bPNA.
Results
Design and synthesis of fluorogenic bPNA probes for URIL RNAs
We previously reported that cyanine dye (Cy3, Cy5) modified bPNAs exhibited fluorescence enhancement upon triplex hybridization to URIL RNA and subsequent hybrid RNA-protein binding45. This was expected52 based on prior observations with DNA binding53. Thiazole orange dye is a more typical choice to signal intercalative binding due to its high emission in the bound state and low emission in the unbound state: Seitz and others54 have demonstrated sequence-selective “forced intercalation” (FIT) emission with PNA-thiazole orange (TO) conjugates55 that place TO at the hybridization interface. Similarly, Dervan has attached TO to pyrrole-imidazole polyamide (Py-Im) minor groove intercalators that fluoresce upon sequence recognition56. While these probes have utility, they are limited with regard to transport and RNA binding57. To study fluorogenic RNA binding in the context of URIL triplex hybridization, we synthesized bPNAs N-terminated with Cy5, difluorohydroxybenzylidene (DFHBI)7 and TO derivatives (Fig. 2 and Supplementary Fig. 18). Prior work has established that compact dipeptide or tripeptide bPNA scaffolds containing just four melamine base-tripling units (4 M bPNAs) can exhibit high affinity binding to a structured U4xU4 internal loop (URIL). Structure-function studies51,58,59 indicated that presentation of two melamine bases on the ε-amine of lysine (K2M) results in a solubilizing cationic sidechain that enables strong URIL binding even with short, low molecular weight di and tri-peptides. Even following N-terminal modification with fluorogenic modules, these efficient bPNA probes are in a low molecular weight regime (~1 kD). Moreover, Fmoc-K2M can be obtained on a multigram scale by double reductive alkylation of Fmoc-lysine with melamine acetaldehyde without column purification60, making these bPNA probes are highly accessible and synthetically scalable. We prepared a small family of dye-modified (Cy3, Cy5, DFHBI, TO) 4 M bPNAs (Fig. 2), focusing primarily on the tripeptide (K2MX-K2M) scaffold where X is an α-amino acid and dipeptide isomers. Fluorogenic turn-on upon RNA complexation was modest for the cyanine and DFHBI dyes (Supplementary Figs. 7 and 14); we thus focused on TO derivatives. In K2MX-K2M, thiazole orange carboxylic acid was coupled via a β-alanine linker to the N-terminus of (X=Ala, Ile) or directly to the ε-amine of lysine (X = KTO). As dipeptide and tripeptide bPNAs of alternating amino acid configuration had previously shown an advantage in triplex hybridization over the homochiral dipeptides due to syndiotactic base presentation51,61,62, we set out to test the impact of stereochemistry on the fluorogenic binding when modified with TO on the N-terminus (TO-k2MK2M, TO-K2Mk2M) and ε-amine of the central amino acid (K2MkTO-K2M). Additionally, we synthesized and evaluated the isodipeptide (TO-(αK2M)2) that features a sidechain linkage, with base presentation on the α-nitrogens and TO modification on the ε-amine.
In vitro evaluation of fluorogenic URIL-RNA binding by bPNA variants
With URIL-binding bPNAs in hand, we tested fluorogenic binding in vitro to hairpin and duplex URIL-RNAs (Fig. 3A). The absorbance and emission properties of the free TO-bPNAs were similar to the parent thiazole orange dye, which exhibits weak emission in solution (quantum yield 10−4)63. However, when TO-K2MAla-K2M binds to URIL RNAs (Fig. 3A), emission increases by up to 600-fold (Fig. 4, Supplementary Figs. 12 and 13). Indeed, the TO-bPNA:RNA hybrid exhibited a robust relative quantum yield of 43% (Fig. 4B). This TO-bPNA:URIL hybrid quantum yield exceeds that observed with the RNA aptamer Mango bound to TO-biotin (quantum yield = 14%)10. An RNA hairpin construct with a fully base-paired duplex stem (RNAII) does not improve the brightness of the TO-bPNA derivatives (Fig. 4C). While thiazole orange fluorescence turn-on upon RNA binding is expected for non-specific intercalation, this only occurs at higher concentrations64; the null response with non-URIL RNA (RNAII) indicates the critical role of bPNA triplex hybridization in fluorogenic binding. All bPNAs gave a strong fluorogenic response to URIL RNAs, but there appeared to be significant differences among bPNA variants51,58. Further, despite the biophysical advantage previously observed for triplex hybridization with L,D and D,L dipeptide bPNAs51, these fluorogenic variants were not as bright as the tripeptides (K2MAlaK2M, K2MIleK2M). Based on these results, we chose the TO-K2MAlaK2M probe (1) for subsequent intracellular studies.
A RNAs for in vitro evaluation were prepared via run-off transcription or purchased directly. The 12-U6-12 and 12-U4-12 constructs feature 12 bp duplexes (indicated by dashed lines) separated by U6xU6 and U4xU4 internal bulges, respectively. B RNA constructs for fixed cell labeling (HEK-293T) of RNPs are shown; hairpin RNAs were used to replace the anticodon stem of tRNALys (dashed lines) or appended to a TDP-43 binding (GU)8 sequence. Constructs for live cell (U2OS) RNA tracking with C FLURIL tagging and D Sirius-IDR2-(MS2)8 with MS2 (MBSV5)84 hairpins are shown in red. IDR3-U4 carries a single PP7 hairpin as an internal control. RNAs for cell experiments were delivered by plasmid transfection. See “Methods” and Supplementary Fig. 2 for sequences and protocols.
A Normalized absorbance and emission of a representative TO-bPNA 1 hybrid with RNA. B Relative fluorescence units (RFU) of fluorescein standard and the hybrid in (A) indicating relative brightness (Φ = 43%, ε507 = 44,883 M−1cm−1). In vitro fluorescence of C bPNAs alone or with RNAII (lacking a URIL) and D upon treatment with RNA constructs that have a URIL binding site (Fig. 2). The mean value of three separate measurements with standard deviation error for all RNA and bPNA combinations are shown under identical conditions (50 mM HEPES, pH 7.5, 100 mM NaCl, 1 µM bPNA and RNA).
Intracellular labeling of URIL RNAs and bacteriophage RNPs with fluorogenic bPNA
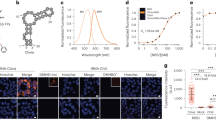
Using similar design principles as the in vitro studies, plasmid vectors were constructed to encode tRNALys with URIL RNA hairpins (Fig. 3B, Supplementary Fig. 1) installed in place of the anticodon loop; this platform affords stable intracellular RNA expression7 upon transfection into HEK-293T cells. A construct encoding a tRNA scaffold with a fully base-paired stem in place of the URIL (NEG tRNA) was designed to serve as an intracellular negative control that lacks bPNA binding. The U4-tRNA construct was used to determine intracellular fluorogenic triplex hybridization with TO-bPNA. Indeed, treatment of HEK-293T cells with probe 1 in cell culture media following transfection with either U4 or NEG tRNA resulted in a 5- to 10-fold brighter fluorescence intensity in the U4-tRNA transfected cells (Fig. 5, Supplementary Figs. 9 and 10). These data were supportive of intracellular targeting of URIL RNA, albeit with diminished enhancement relative to in vitro conditions.
A Integrated fluorescence of U4 tRNA, NEG tRNA and untreated cells normalized to untreated cells. The mean value of 10 independent cell measurements of normalized fluorescence (scatter plot) and standard deviation error is shown. All cell samples were treated with TO-bPNA 1. B Representative confocal fluorescence microscopy images from triplicate studies of HEK-293T cells treated with 1 in media and transfection with plasmid encoding (Top row) U4 tRNA and (Bottom row) NEG tRNA. 170/177 (97%) of cells were labeled with TO-bPNA. Scale bar = 50 µm.
To benchmark bPNA labeling of RNA against known RNA tracking strategies, we juxtaposed the U4 URIL with the MS2 hairpin sequence in the tRNALys scaffold to yield a construct encoding U4-MS2 tRNA (Fig. 3B, Supplementary Fig. 1). The U4-MS2 tRNA plasmid was co-transfected into HEK-293T cells with a plasmid encoding an MCP-RFP fusion, which also bears a nuclear localization signal (NLS)65. Cells imaged 2 h after transfection and treatment in media with 1 revealed clear co-localization of red and green fluorescence, supportive of labeling of the U4-MS2 tRNA with TO-bPNA and subsequent binding of the MCP-RFP protein fusion to the triplex hybrid in the cytoplasm (Fig. 6, Supplementary Figs. 5, 6 and 11). This red-green co-localization was accompanied by a condensation of signal intensity, possibly due to RFP aggregation66. Notably, imaging 8 h after transfection indicated that both green and red fluorescence had co-localized to the nucleus, consistent with time-dependent transport of both MCP-RFP (which bears the NLS) along with the TO-bPNA hybrid with U4-MS2 tRNA. Thus, treatment of cell culture with TO-bPNA in media can effectively FLURIL-tag intracellular RNA and RNP targets and track RNP transport from cytoplasm to nucleus. Importantly, without MCP-RFP, green fluorescence from TO-bPNA remains cytoplasmic; without TO-bPNA, MCP-RFP remains localized to the nucleus as expected.
Representative confocal fluorescence microscopy images of HEK-293T cells treated as indicated at the left of each row and imaged under the dye channels indicated at the top of each column. Experiments were performed in triplicate. Scale bar = 10 µm. The MCP-RFP fusion contains a nuclear localization tag. The first row is imaged 2 h after transfection and treatment with TO-K2MAla-K2M (1 µM) in media, while subsequent rows are imaged 8 h after transfection.
Intracellular labeling of mammalian RNPs with fluorogenic bPNA
To complement the bacteriophage RBP system, we conducted an identical experiment with a native mammalian RNA and protein pair to verify intracellular targeting by TO-bPNA. There have been extensive studies on TAR DNA/RNA binding protein (TDP-43) due to its central role in the progression of neurodegenerative diseases such as ALS67,68,69, where it is mislocalized from the nucleus to the cytoplasm70. As TDP-43 is known to bind to repeat sequences of UG/TG71,72, we co-transfected HEK-293T cells with plasmids encoding URIL-UG repeat RNA (U4-(GU)8, Fig. 3B, Supplementary Fig. 1) and TDP-43-tdTomato73. Upon Treatment with TO-bPNA in cell media resulted in co-localized red (tdTomato) and green fluorescence in the nucleus (Fig. 7, Supplementary Fig. 8). This is supportive of successful FLURIL tagging of the TDP-43/(GU)8 mammalian RNP, in the correct subcellular compartment. With the MS2/MCP and (GU)8/TDP-43 systems verified, we transfected with swapped RNA-protein pairings that should not result in complex formation: U4-MS2 was co-transfected with TDP-43-tdTomato, and U4-(GU)8 with MCP-RFP (Fig. 7). In these control experiments, both red fluorescent proteins were retained in the nucleus, but the TO-bPNA tagged RNAs remained cytoplasmic, indicating that RNA-protein binding is driving co-localization of red and green fluorescence signals.
Representative confocal fluorescence microscopy images of HEK-293 cells treated as indicated at the left of each row and imaged under the dye channels indicated at the top of each column. Experiments were performed in triplicate. Scale bar = 10 µm. The top row indicates nuclear co-localization of TDP-43-tdTomato with TO-K2MAla-K2M fluorescence when co-expressed with U4-(GU)8. Lower two rows feature mismatches between RNA and RBP (U4-(GU)8 with MCP-RFP and U4-MS2 with TDP-43-tdTomato, respectively), resulting in partitioning of TO-K2MAla-K2M emission to the cytoplasm.
Live cell tracking of genomic loci with bPNA-labeled CRISPR-dCas/gRNA complexes
To test the scope of bPNA cell imaging, we incorporated TO-bPNA reporting into an established CRISPR imaging approach for live cell genomic loci tracking30. CRISPRainbow is a live cell imaging strategy that uses MS2 or PP7 modified guide RNA (gRNA) with deactivated Cas9 (dCas9) to precisely track endogenous DNA repeats marking specific genomic loci74. Incorporation of MS2/PP7 bacteriophage RNA hairpin sequences into the gRNA enables the dCas-gRNA complexes to be tracked in live cells by fluorescence imaging upon co-expression of phage coat protein (MCP/PCP) fusions with a “rainbow” of fluorescent proteins; sequence-optimization of the bacteriophage hairpin array yielded more stable and brighter imaging systems known as CRISPR-Sirius gRNAs75. We selected two CRISPRainbow/Sirius gRNAs that target proximal intergenic DNA regions (IDR2, IDR3) and modified one to utilize FLURIL tagging only, with the other serving as an internal benchmark of MS2 labeling. On the gRNA targeting IDR3, a single U4XU4 bulge (URIL) was installed in a stabilized hairpin (Fig. 3C, Supplementary Fig. 2), rendering the construct (IDR3-U4) trackable by FLURIL tagging. The IDR3-U4 gRNA carries a PP7 hairpin that serves as an internal control for labeling experiments that lack a bPNA probe and was indeed found to be unreactive to labeling in the absence of TO-bPNA. To target IDR2, a CRISPR-Sirius gRNA called Sirius-IDR2-(MS2)8 was used (Fig. 3D), which features an array of eight MS2 hairpins optimized for stability. A plasmid was constructed encoding both IDR3-U4 and Sirius-IDR2-(MS2)8, and this vector was transfected into U2OS cells stably expressing dCas9 and MCP-HaloTag. Staining of live cells in culture with TO-bPNA and HaloTag-JF549 dye thus enabled simultaneous and orthogonal imaging of IDR3 and IDR2, respectively (Fig. 8A).
A Illustration of dual-color genomic labeling of IDR2 and IDR3 by CRISPR-dCas9 targeting. IDR3 is tracked by FLURIL tagging of IDR3-targeting gRNA, while MS2 labeling of IDR2-targeting gRNA is used to track IDR2. FLURIL tags are stained with TO-bPNA, while MS2 labels are stained with MCP-HaloTag binding and HaloTag-JF549 reaction with the complex. MS2 labeling is accomplished using the CRISPR-Sirius gRNA design in Sirius-IDR2-(MS2)8 gRNA, which has an array of eight MS2 hairpins. B Plasmids used: (Top) dual gRNA plasmid driven by two promoters (hU6 for Sirius-IDR2-(MS2)8, CMV for IDR3-U4), (Middle) control dual gRNA plasmid with IDR3-U4 gRNA replaced by Sirius-IDR3-(PP7)8, (Bottom) plasmids carrying dCas9 under an inducible promoter, MCP-Halotag fusion under continuous expression. NLS nuclear localization signal, P2A cleavage peptide, HSA mouse heat-stable antigen.
The final vector design was driven by the low efficiency of initial efforts in the expression of the modified gRNAs. We speculated that the oligo-U domains triggered early termination by RNA Pol III, which is known to significantly reduce the gRNA targeting efficiency in the CRISPR system65,76. To bypass this issue, we expressed gRNA under a CMV Pol II promoter and ensured precise transcript generation by flanking the gRNA sequence with Hammerhead (HH) and Hepatitis Delta Virus (HDV) self-cleaving ribozymes77. We thus modified the gRNA plasmid to contain the Hammerhead (HH) and Hepatitis delta virus (HDV) ribozymes on either side of the gRNA under a CMV promoter and inserted it into a dual gRNA plasmid expressing CRISPR-Sirius gRNA (Sirius-IDR2-(MS)8) for dual-color genomic loci targeting (Fig. 8). Indeed, the resulting dual-color genomic loci labeling system for IDR2 (Sirius-IDR2-(MS2)8 + MCP-Halotag + Halotag-JF549) and IDR3 (IDR3-U4 + TO-bPNA) was readily established in U2OS cells by lipofectamine plasmid transfection and staining with dyes delivered in media (HaloTag-JF549, TO-bPNA). Live cell imaging revealed partially overlapped red (IDR2, HaloTag-JF549) and green (IDR3, TO-bPNA) foci (Fig. 9, top row, Supplementary Fig. 4), which was expected given the close proximity (~4.6 kB) of the two loci on chromosome 19. The exclusion of TO-bPNA from media resulted in the loss of IDR3 foci, leaving only red IDR2 foci visible (Fig. 9, middle row). Additionally, identical plasmid transfection protocols were carried out in which the FLURIL gRNA was replaced with a CRISPRainbow gRNA bearing PP7 hairpins (Sirius-IDR3-8xPP7). When stained with HaloTag-JF549 and TO-bPNA, only red foci were visible (Fig. 8, bottom row), consistent with URIL-selective fluorogenic binding of the TO-bPNA probe previously observed in fixed cells (Fig. 5). FLURIL-tagged IDR3 loci was easily tracked over 13 s without significant photobleaching or dye blinking, performing comparably to IDR2 loci tracking with the established CRISPR-Sirius (8X-MS2) system75. Time-lapse imaging was carried out for 96 frames at a capture rate of 136 ms per frame (Supplementary Movie S1), yielding IDR2 and IDR3 trajectories on identical and homologous chromosomes of similar range (~100–200 nm) but in different patterns of loci territories, consistent with the labeling of proximal, but distinct loci (Fig. 10). FLURIL tagging of gRNA thus gave a highly usable fluorescence signal-to-noise ratio with single locus labeling, likely due to minimal background fluorescence from unbound TO-bPNA, unlike constitutively fluorescent FP and HaloTag systems. Though single molecule fluorescence tracking has not been demonstrated, we note that the sole FLURIL tag was sufficient for labeling the low-copy number IDR3 locus (45 copies over 1.5 kB). Under standard operating conditions, the low background of a single FLURIL gRNA tag affords 7X greater brightness (Supplementary Fig. 3) than the CRISPR-Sirius gRNA tags that have a substantially larger molecular footprint of eight bacteriophage RNA hairpins bound by bacteriophage coat protein fusions.
(Top row) Representative loci imaging from triplicate independent measurements, following transfection with a dual plasmid encoding Sirius-IDR2-(MS2)8 and IDR3-U4, followed by staining with Halo-JF549 (red) and TO-bPNA (green), respectively. Genomic loci highlighted in the white square are shown enlarged, inset bottom right-hand corner. (Middle row) Same plasmid transfection as top row without TO-bPNA treatment. (Bottom row) Loci imaging following transfection with a dual plasmid encoding Sirius-IDR2-(MS2)8 and Sirius-IDR3-(PP7)8, followed by staining with Halo-JF549 (red) and TO-bPNA (green), respectively. Bar in the inset, 1 μm.
U2OS cells were imaged 48 h post-transfection with a dual plasmid encoding Sirius-IDR2-(MS2)8 and IDR3-U4 and stained with HaloTag-JF549 (red) and TO-bPNA (green). Representative loci trajectories of 96 frames from homologous chromosomes are shown in the top and bottom from triplicate independent measurements. The imaging rate is 136 ms per frame. Bar in the inset, 1 μm. For comparison, trajectories were aligned to start from the origin (0,0).
Discussion
Taken together, these data establish FLURIL tagging as an efficient and convenient strategy for intracellular RNA, RNP, and DNA tracking that leverages selective URIL targeting with fluorogenic binding to establish a robust and bright RNA tracking signal. We find that TO-bPNA exhibits significant (~600X) enhancement of emission intensity upon bPNA triplex hybridization with structured RNAs URILs in vitro and a substantive increase in quantum yield from ~0.01% in the unbound state to 43% in the RNA complex. This significant fluorogenic response to URIL-RNAs translated to 5–10X intracellular signal-to-noise and a highly usable, low-background cellular imaging of URILs. While an engineered biomolecule (URIL-RNA) is needed, this is a universal requirement for the most broadly used, genetically encoded live cell tracking strategies such as MS2-labeling.
Intracellular FLURIL tagging of RNAs in both fixed and live cells was directly verified using established protein labels from endogenous mammalian RNPs as well as bacteriophage MS2 labeling, which remains the gold standard for RNA tracking. In the present study, we have inserted stabilized hairpin structures to host the URIL site and included a PP7 hairpin as a control element in IDR3-U4; however, a URIL could be inserted into a native fold by replacement of an existing native 4 bp stem buttressed by other structures in the ROI, further streamlining the FLURIL-tag footprint. Importantly, RBP fusions with fluorescent proteins (FP) confirmed that FLURIL tags were labeling the correct RNA species, while cells lacking the correct RNA-protein pairing resulted in the segregation of FLURIL tag and RBP-FP fusion signals. Under live cell tracking conditions, FLURIL tags also performed comparably to established MS2-based methods (CRISPR-Sirius) while occupying a considerably more compact molecular footprint. It is particularly noteworthy that the FLURIL tag only requires the replacement of a 4 bp stem with an 8-nt URIL to enable RNP live cell tracking with a cell-permeable, ~1kD bPNA probe.
Though there remain many appealing aspects to MS2 labeling, including multiplex and multicolor approaches, the use of protein fusions and multiple RNA hairpin sites encumbers the RNA of interest with substantial steric bulk, adding 41–47 kD of mass per MCP-FP and MCP-HaloTag fusion, respectively. The steric bulk and RNA secondary structures required for protein labeling have raised concerns that this could affect native transcript processing and tertiary contacts1; however, inhibition of transcript degradation by insertion of bacteriophage (MS2/PP7) hairpins is less of an issue in mammalian cells22,27,28. In contrast, FLURIL tags add negligible additional mass to the RNA target and a structural perturbation that may be as minor as the replacement of a duplex stem with a triplex stem. Prior work has demonstrated that this replacement does not disrupt proximal RNA domains, retaining tertiary interactions as well as catalytic function. In addition to minimizing the addition of obtrusive non-native structures, FLURIL tagging also avoids the use of labile21 G-quadruplex domains found in SELEX-derived dye-binding aptamers such as Spinach and Mango. Furthermore, as fluorogenic dye binding is driven by bPNA triplex hybridization to the URIL rather than direct RNA recognition of the dye, prosthetic groups50 may be used to decorate RNPs without the technical demand of a SELEX campaign. In particular, other fluorogens in the cyanine family could potentially be used in FLURIL tagging, thus expanding the color range available without laborious aptamer selection procedures. Analogously, the aptamer Pepper11 binds a family of fluorogenic dyes with significant structural variation, suggesting that minimal specific contacts are needed to access fluorogenic binding.
The unique advantages of FLURIL tagging are counterbalanced by some limitations. As mentioned above, a URIL motif must be installed in the RNA of interest, requiring non-native RNA expression by transfection. The efficiency of tracking, as described herein, thus depends on the efficiency of transfection and vector design, which can lead to variability in labeling outcomes; additionally, transfection yields elevated, non-native transcript levels. These issues may be alleviated with genome editing, though this comes with a significant cost and effort per experiment. Currently, FLURIL tagging has only been demonstrated with a single color, though efforts to expand both color selection and sequence scope41 are underway. Despite these caveats, the present work demonstrates the utility of FLURIL tagging, as well as its appealing compatibility with popular methods such as MS2/Halotag labeling. We anticipate that FLURIL tagging will be a useful tool that may also be used in conjunction with other methods (eg-dye aptamers, Riboglow, dCas13, FISH) to facilitate RNA tracking while minimizing the steric burdens of RNA labeling.
Methods
Materials and handling
All chemicals were used without further purification from commercial sources as indicated unless otherwise noted. DNAs and RNAs were purchased from Integrated DNA Technologies (IDT). Nucleic acid strands shorter than 20 nt were used without further purification. Otherwise, the DNAs/RNAs were purified by TBE-urea denaturing gel. DNA stock solutions were serially diluted in MilliQ water, and concentrations were determined by measuring solution absorbance at 260 nm on a Thermo Fisher Nanodrop 2000. Sample fluorescence was measured on a Thermo Fisher Nanodrop 3300. Cell lines were acquired from ATCC (HEK-293T, cat# CRL-2316; U2OS, cat# HTB-96) and cultured according to ATCC protocols.
Nucleic acid sequences
RNAs for in vitro fluorescence studies shown below were purchased. All RNAs were annealed prior to use. Duplexes were annealed together from A and B strands indicated.
12-U4-12 A: 5’-CGCAUAGCUCAGUUUUGACUCGAUACGC-3’
12-U4-12 B: 5’-GCGUAUCGAGUCUUUUCUGAGCUAUGCG-3’
12-U6-12 A: 5’-CGCAUAGCUCAGUUUUUUGACUCGAUACGC-3’
12-U6-12 B: 5’-GCGUAUCGAGUCUUUUUUCUGAGCUAUGCG-3’
RNAI U6: 5’-GGCAGCUUUUUUUUGGUAGUUUUUUGCUGCC-3’
RNAII WT: 5’-GCACCGCUACCAACGGUGC-3’
The following RNA sequences were delivered into cells by plasmid transfection for intracellular labeling.
U4 tRNA: 5’-GCCCGGAUAGCUCAGUCGGUAGAGCAGCGGCCGUUUUCGCUCCGGCGUUUUCGGCCGCGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGCCA-3’
U4-MS2 (MBSV5) tRNA: 5’-GCCCGGAUAGCUCAGUCGGUAGAGCAGCGGCCGUUUUCGCACAUGAGGAUCACCCAUGUGCGUUUUCGGCCGCGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGCCA-3’
NEG tRNA: 5’-GCCCGGAUAGCUCAGUCGGUAGAGCAGCGGCCGCGCGCGCUCCGGCGCGCGCGGCCGCGGGUCCAGGGUUCAAGUCCCUGUUCGGGCGCCA
(GU)8-U4 RNA: 5’-CGGCCGUUUUCGCUCCGGCGUUUUCGGCCGGUGUGUGUGUGUGUGU-3’
In vitro fluorescence measurement
RNAs were annealed with bPNA at a 1:1 ratio and incubated for 30 min following slow cooling (50 mM HEPES, pH 7.5, 100 mM NaCl, 2 μM RNA, 2 μM bPNA). Sample fluorescence was measured using a Thermo Fisher Nanodrop 3300 (relative fluorescence units (RFU), excitation = 470 nm, emission = 522 nm, 2 μl sample volume). All fluorescence values are the average of triplicate measurements, starting from fresh sample preparation. Error bars indicate standard deviation. Data were processed using OriginPro and Graphpad Prism.
Quantum yield measurement
Relative quantum yield was calculated against fluorescein (quantum yield = 92%)78. All dye samples (TO, TO-bPNA) were prepared at 1 μM with hybrid samples prepared at 1:1 TO:RNA ratio (50 mM HEPES, pH 7.5, 100 mM NaCl). Sample concentrations were determined by UV absorbance (Cary UV-Vis-NIR Spectrophotometer) and adjusted to obtain matched sample absorbances, not to exceed 0.04 at 470 nm (±0.0002). The fluorescence was measured by Quantamaster 8000 spectrofluorometer and corrected for anisotropy with a vertical polarizer in the excitation path (470 nm) and a polarizer in the emission path (507 nm) set at the magic angle (54.7°), with excitation and emission slit widths = 5 nm)79. The integrated emission of the bPNA hybrid was 49.7% that of fluorescein, indicating a relative quantum yield of 43%.
Plasmid construction
The DNA inserts corresponding to modified tRNA scaffolds (U4, U4-MS2, NEG tRNA) were annealed into duplexes from which 2 µg were digested with SalI and XbaI (Thermo Fisher, 2x Tango buffer, 12 h), following Thermo Fisher protocol. Vector pAV U6 + 27 (1 µg) was separately digested with SalI and XbaI (12 h), and all digested products were gel purified (band isolation from with Qiagen QIAquick Gel Extraction Kit). Purified products were ligated (DNA insert:linearized vector=5:1) with T4 DNA ligase (Thermo Fisher), following published protocol80. The ligation product was transformed into DH5α for amplification, and the resulting mixture was inoculated on an agar plate with ampicillin selection. After 16 h, several colonies were picked and amplified in LB media containing ampicillin. The harvested plasmid was isolated by miniprep (Qiagen) and verified by sequencing. Tomato-TDP43 and MCP-TagRFPt plasmids were obtained from Addgene (#28205 and #64541)73,81 and amplified in the corresponding E.coli cell lines.
Cell treatment (fixed cell analysis)
HEK-293T cells were cultured based on ATCC protocol. Cells were seeded to a 35-mm culture dish (Thermo Fisher) with a clean coverslip (Thermo Fisher) attached to the bottom at a concentration of 5 × 105/ml. After one day of incubation, the cells were transfected with corresponding plasmids (1000 ng each per dish) with Lipofectamine 3000 (Invitrogen) following the online protocol, and the cells were incubated for 24 h. Culture medium was removed and the fresh medium containing 1 µM bPNA-dye was added to the attached cells. The cells were incubated at 37 °C for 2–8 h, medium discarded, and Hoechst 33258 (Invitrogen) in fresh culture medium was added at the final concentration of 200 ng/ml to the cells and incubated for 15 min at 37 °C. Cells were washed with PBS, fixed with 4% formaldehyde in PBS for 15 min at room temperature and rinsed with PBS. The coverslip was transferred to glass slides for fluorescence microscopy and imaged under Olympus FV3000 systems (Objective lens: ×40, Zoom: ×1. Blue channel: (excitation) 405 nm, (emission) 422 nm, Voltage: 355 V, detection range: 420–470 nm; Green: (excitation) 488 nm, (emission) 520 nm, Voltage: 580 V, detection range: 499–599 nm). Three or more replicate data sets were obtained that showed consistent findings, starting from cell seeding.
Fluorescence microscopy (live cell imaging)
Cell imaging (U2OS) was carried out on an Olympus IX83 microscope equipped with three EMCCD cameras (Andor iXon 897) mounted on a 4-camera splitter, four lasers (405 nm, 488 nm, 561 nm and 647 nm), mounted with a ×1.6 magnification adapter and ×60 apochromatic oil objective lens (NA 1.5), resulting in a total of ×96 magnification. The microscope stage incubation chamber was maintained at 37 °C with CO2 and humidity supplement. A laser quad-band filter set for TIRF (emission filters at 445/58, 525/50, 595/44, 706/95) was used to collect fluorescence signals simultaneously. Data acquisition was carried out with CellSens 4.1.1 software. Localization precision was ~5 nm in 4 s, ~6 nm in 16 s, and ~10 nm in 80 s82. The video was recorded at 136 ms per frame with a total of 96 frames and 100 ms exposure time. Image size was adjusted to show individual nuclei, and intensity thresholds were set on the basis of the ratios between nuclear foci signals to background nucleoplasmic fluorescence.
Image processing
The images were registered and analyzed by Fiji83 and Mathematica (Wolfram) software. To achieve subpixel registration accuracy, parameters for shifting, scaling, and rotating camera images were determined by the least-squares fitting of fluorescent bead images (100 nm TetraSpeck fluorescent microspheres, Invitrogen). The experimental data from each channel were processed through an affine transformation and overlapped in false-color channels for visualization. The locus trajectory was obtained by tracking the locus position over time, and graphs were generated by OriginPro (OriginLab version 2019b).
Reporting summary
Further information on research design is available in the Nature Portfolio Reporting Summary linked to this article.
Data availability
All data that support this manuscript, including source data files, are available in the Supplementary Information. Source data are provided with this paper.
References
Braselmann, E., Rathbun, C., Richards, E. M. & Palmer, A. E. Illuminating RNA biology: tools for imaging RNA in live mammalian cells. Cell Chem. Biol. 27, 891–903 (2020).
Alexander, S. C., Busby, K. N., Cole, C. M., Zhou, C. Y. & Devaraj, N. K. Site-specific covalent labeling of RNA by enzymatic transglycosylation. J. Am. Chem. Soc. 137, 12756–12759 (2015).
Jahn, K. et al. Site-specific chemical labeling of long RNA molecules. Bioconjug. Chem. 22, 95–100 (2011).
Smith, G. J., Sosnick, T. R., Scherer, N. F. & Pan, T. Efficient fluorescence labeling of a large RNA through oligonucleotide hybridization. RNA 11, 234–239 (2005).
Schmitz, A. G., Zelger-Paulus, S., Gasser, G. & Sigel, R. K. O. Strategy for internal labeling of large RNAs with minimal perturbation by using fluorescent PNA. Chembiochem 16, 1302–1306 (2015).
Mao, S., Ying, Y., Wu, R. & Chen, A. K. Recent advances in the molecular beacon technology for live-cell single-molecule imaging. iScience 23, 101801 (2020).
Paige, J. S., Wu, K. Y. & Jaffrey, S. R. RNA mimics of green fluorescent protein. Science 333, 642–646 (2011).
Filonov, G. S., Moon, J. D., Svensen, N. & Jaffrey, S. R. Broccoli: rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 136, 16299–16308 (2014).
Warner, K. D. et al. A homodimer interface without base pairs in an RNA mimic of red fluorescent protein. Nat. Chem. Biol. 13, 1195–1201 (2017).
Dolgosheina, E. V. et al. RNA Mango aptamer-fluorophore: a bright, high-affinity complex for RNA labeling and tracking. ACS Chem. Biol. 9, 2412–2420 (2014).
Chen, X. et al. Visualizing RNA dynamics in live cells with bright and stable fluorescent RNAs. Nat. Biotechnol. 37, 1287–1293 (2019).
George, L., Indig, F. E., Abdelmohsen, K. & Gorospe, M. Intracellular RNA-tracking methods. Open Biol. 8, 180104 (2018).
Schmidt, A., Gao, G., Little, S. R., Jalihal, A. P. & Walter, N. G. Following the messenger: recent innovations in live cell single molecule fluorescence imaging. Wiley Interdiscip. Rev. RNA 11, e1587 (2020).
Sunbul, M. & Jäschke, A. SRB-2: a promiscuous rainbow aptamer for live-cell RNA imaging. Nucleic Acids Res. 46, e110 (2018).
Bouhedda, F. et al. A dimerization-based fluorogenic dye-aptamer module for RNA imaging in live cells. Nat. Chem. Biol. 16, 69–76 (2020).
Braselmann, E. et al. A multicolor riboswitch-based platform for imaging of RNA in live mammalian cells. Nat. Chem. Biol. 14, 964–971 (2018).
Trachman, R. J. & Ferré-D’Amaré, A. R. Tracking RNA with light: selection, structure, and design of fluorescence turn-on RNA aptamers. Q. Rev. Biophys. 52, e8 (2019).
Warner, K. D. et al. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 21, 658–663 (2014).
Huang, H. et al. A G-quadruplex–containing RNA activates fluorescence in a GFP-like fluorophore. Nat. Chem. Biol. 10, 686–691 (2014).
Trachman, R. J. III et al. Structural basis for high-affinity fluorophore binding and activation by RNA Mango. Nat. Chem. Biol. 13, 807–813 (2017).
Guo, J. U. & Bartel, D. P. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science 353, aaf5371 (2016).
Tutucci, E. et al. An improved MS2 system for accurate reporting of the mRNA life cycle. Nat. Methods 15, 81–89 (2018).
Bertrand, E. et al. Localization of ASH1 mRNA particles in living yeast. Mol. Cell 2, 437–445 (1998).
Grimm, J. B. et al. A general method to improve fluorophores for live-cell and single-molecule microscopy. Nat. Methods 12, 244–250 (2015).
Hoelzel, C. A. & Zhang, X. Visualizing and manipulating biological processes by using HaloTag and SNAP-tag technologies. Chembiochem 21, 1935–1946 (2020).
Garcia, J. F. & Parker, R. MS2 coat proteins bound to yeast mRNAs block 5’ to 3’ degradation and trap mRNA decay products: implications for the localization of mRNAs by MS2-MCP system. RNA 21, 1393–1395 (2015).
Garcia, J. F. & Parker, R. Ubiquitous accumulation of 3’ mRNA decay fragments in Saccharomyces cerevisiae mRNAs with chromosomally integrated MS2 arrays. RNA 22, 657–659 (2016).
Heinrich, S., Sidler, C. L., Azzalin, C. M. & Weis, K. Stem-loop RNA labeling can affect nuclear and cytoplasmic mRNA processing. RNA 23, 134–141 (2017).
Pickar-Oliver, A. & Gersbach, C. A. The next generation of CRISPR–Cas technologies and applications. Nat. Rev. Mol. Cell Biol. 20, 490–507 (2019).
Nelles, D. A. et al. Programmable RNA tracking in live cells with CRISPR/Cas9. Cell 165, 488–496 (2016).
Chen, B. et al. Dynamic imaging of genomic loci in living human cells by an optimized CRISPR/Cas system. Cell 155, 1479–1491 (2013).
Lange, R. F. M. et al. Crystal engineering of melamine–imide complexes; tuning the stoichiometry by steric hindrance of the imide carbonyl groups. Angew. Chem. Int. Ed. Engl. 36, 969–971 (1997).
Whitesides, G. M. et al. Noncovalent synthesis: using physical-organic chemistry to make aggregates. Acc. Chem. Res. 28, 37–44 (1995).
Ma, M. & Bong, D. Determinants of cyanuric acid and melamine assembly in water. Langmuir 27, 8841–8853 (2011).
Ma, M. & Bong, D. Controlled fusion of synthetic lipid membrane vesicles. Acc. Chem. Res. 46, 2988–2997 (2013).
Arambula, J. F., Ramisetty, S. R., Baranger, A. M. & Zimmerman, S. C. A simple ligand that selectively targets CUG trinucleotide repeats and inhibits MBNL protein binding. Proc. Natl Acad. Sci. USA 106, 16068–16073 (2009).
Sanjayan, G. J., Pedireddi, V. R. & Ganesh, K. N. Cyanuryl-PNA monomer: synthesis and crystal structure. Org. Lett. 2, 2825–2828 (2000).
Piao, X., Xia, X. & Bong, D. Bifacial peptide nucleic acid directs cooperative folding and assembly of binary, ternary, and quaternary DNA complexes. Biochemistry 52, 6313–6323 (2013).
Zeng, Y., Pratumyot, Y., Piao, X. & Bong, D. Discrete assembly of synthetic peptide-DNA triplex structures from polyvalent melamine-thymine bifacial recognition. J. Am. Chem. Soc. 134, 832–835 (2012).
Miao, S. et al. Unnatural bases for recognition of noncoding nucleic acid interfaces. Biopolymers 112, e23399 (2021).
Liang, Y. et al. Screening of minimalist noncanonical sites in duplex DNA and RNA reveals context and motif-selective binding by fluorogenic base probes. Chemistry 28, e202103616 (2022).
Mittapalli, G. K. et al. Mapping the landscape of potentially primordial informational oligomers: oligodipeptides and oligodipeptoids tagged with triazines as recognition elements. Angew. Chem. Int. Ed. Engl. 46, 2470–2477 (2007).
Mittapalli, G. K. et al. Mapping the landscape of potentially primordial informational oligomers: oligodipeptides tagged with 2,4-disubstituted 5-aminopyrimidines as recognition elements. Angew. Chem. Int. Ed. Engl. 46, 2478–2484 (2007).
Ura, Y., Beierle, J. M., Leman, L. J., Orgel, L. E. & Ghadiri, M. R. Self-assembling sequence-adaptive peptide nucleic acids. Science 325, 73–77 (2009).
Miao, S. et al. Duplex stem replacement with bPNA+ triplex hybrid stems enables reporting on tertiary interactions of internal RNA domains. J. Am. Chem. Soc. 141, 9365–9372 (2019).
Mao, J., DeSantis, C. & Bong, D. Small molecule recognition triggers secondary and tertiary interactions in DNA folding and hammerhead ribozyme catalysis. J. Am. Chem. Soc. 139, 9815–9818 (2017).
Xia, X., Piao, X. & Bong, D. Bifacial peptide nucleic acid as an allosteric switch for aptamer and ribozyme function. J. Am. Chem. Soc. 136, 7265–7268 (2014).
Xia, X., Piao, X., Fredrick, K. & Bong, D. Bifacial PNA complexation inhibits enzymatic access to DNA and RNA. Chembiochem 15, 31–36 (2013).
Liang, Y., Miao, S., Mao, J., DeSantis, C. & Bong, D. Context-sensitive cleavage of folded DNAs by loop-targeting bPNAs. Biochemistry 59, 2410–2418 (2020).
Miao, S. et al. Bifacial PNAs destabilize MALAT1 by 3’ A-tail displacement from the U-rich internal loop. ACS Chem. Biol. 16, 1600–1609 (2021).
Rundell, S., Munyaradzi, O. & Bong, D. Enhanced triplex hybridization of DNA and RNA via syndiotactic side chain presentation in minimal bPNAs. Biochemistry 61, 85–91 (2022).
Klymchenko, A. S. Solvatochromic and fluorogenic dyes as environment-sensitive probes: design and biological applications. Acc. Chem. Res. 50, 366–375 (2017).
Mohammed, H. S., Delos Santos, J. O. & Armitage, B. A. Noncovalent binding and fluorogenic response of cyanine dyes to DNA homoquadruplex and PNA-DNA heteroquadruplex structures. Artif. DNA PNA XNA 2, 43–49 (2011).
Vilaivan, T. Fluorogenic PNA probes. Beilstein J. Org. Chem. 14, 253–281 (2018).
Köhler, O., Jarikote, D. V. & Seitz, O. Forced intercalation probes (FIT Probes): thiazole orange as a fluorescent base in peptide nucleic acids for homogeneous single-nucleotide-polymorphism detection. Chembiochem 6, 69–77 (2005).
Fechter, E. J., Olenyuk, B. & Dervan, P. B. Sequence-specific fluorescence detection of DNA by polyamide-thiazole orange conjugates. J. Am. Chem. Soc. 127, 16685–16691 (2005).
Chenoweth, D. M., Meier, J. L. & Dervan, P. B. Pyrrole-imidazole polyamides distinguish between double-helical DNA and RNA. Angew. Chem. Int. Ed. Engl. 52, 415–418 (2013).
Munyaradzi, O., Rundell, S. & Bong, D. Impact of bPNA backbone structural constraints and composition on triplex hybridization with DNA. Chembiochem 23, e202100707 (2022).
Devari, S., Bhunia, D. & Bong, D. Synthesis of bifacial peptide nucleic acids with diketopiperazine backbones. Synlett 33, 965–968 (2022).
Liang, Y., Mao, J. & Bong, D. In Methods in Enzymology, Vol. 623 (ed. Hargrove, A. E.) Ch. 8, 151–175 (Academic Press, 2019).
Bong, D. T., Clark, T. D., Granja, J. R. & Ghadiri, M. R. Self-assembling organic nanotubes. Angew. Chem. Int. Ed. Engl. 40, 988–1011 (2001).
Ghadiri, M. R., Granja, J. R., Milligan, R. A., McRee, D. E. & Khazanovich, N. Self-assembling organic nanotubes based on a cyclic peptide architecture. Nature 366, 324–327 (1993).
Nygren, J., Svanvik, N. & Kubista, M. The interactions between the fluorescent dye thiazole orange and DNA. Biopolymers 46, 39–51 (1998).
Suss, O., Motiei, L. & Margulies, D. Broad applications of thiazole orange in fluorescent sensing of biomolecules and ions. Molecules 26, 2828 (2021).
Ma, H. et al. Multiplexed labeling of genomic loci with dCas9 and engineered sgRNAs using CRISPRainbow. Nat. Biotechnol. 34, 528–530 (2016).
Piatkevich, K. D. & Verkhusha, V. V. Guide to red fluorescent proteins and biosensors for flow cytometry. Methods Cell Biol. 102, 431–461 (2011).
Ma, X. R. et al. TDP-43 represses cryptic exon inclusion in the FTD-ALS gene UNC13A. Nature 603, 124–130 (2022).
Brown, A.-L. et al. TDP-43 loss and ALS-risk SNPs drive mis-splicing and depletion of UNC13A. Nature 603, 131–137 (2022).
Lagier-Tourenne, C. & Cleveland, D. W. Rethinking ALS: the FUS about TDP-43. Cell 136, 1001–1004 (2009).
Suk, T. R. & Rousseaux, M. W. C. The role of TDP-43 mislocalization in amyotrophic lateral sclerosis. Mol. Neurodegener. 15, 45 (2020).
Kuo, P.-H., Doudeva, L. G., Wang, Y.-T., Shen, C.-K. J. & Yuan, H. S. Structural insights into TDP-43 in nucleic-acid binding and domain interactions. Nucleic Acids Res. 37, 1799–1808 (2009).
Kuo, P.-H., Chiang, C.-H., Wang, Y.-T., Doudeva, L. G. & Yuan, H. S. The crystal structure of TDP-43 RRM1-DNA complex reveals the specific recognition for UG- and TG-rich nucleic acids. Nucleic Acids Res. 42, 4712–4722 (2014).
Yang, C. et al. The C-terminal TDP-43 fragments have a high aggregation propensity and harm neurons by a dominant-negative mechanism. PLoS ONE 5, e15878 (2010).
Ma, H. et al. CRISPR-Cas9 nuclear dynamics and target recognition in living cells. J. Cell Biol. 214, 529–537 (2016).
Ma, H. et al. CRISPR-Sirius: RNA scaffolds for signal amplification in genome imaging. Nat. Methods 15, 928–931 (2018).
Nielsen, S., Yuzenkova, Y. & Zenkin, N. Mechanism of eukaryotic RNA polymerase III transcription termination. Science 340, 1577–1580 (2013).
Xu, L., Zhao, L., Gao, Y., Xu, J. & Han, R. Empower multiplex cell and tissue-specific CRISPR-mediated gene manipulation with self-cleaving ribozymes and tRNA. Nucleic Acids Res. 45, e28 (2017).
Brouwer, A. M. Standards for photoluminescence quantum yield measurements in solution (IUPAC Technical Report). Pure Appl. Chem. 83, 2213–2228 (2011).
Levitus, M. Tutorial: measurement of fluorescence spectra and determination of relative fluorescence quantum yields of transparent samples. Methods Appl. Fluoresc. 8, 033001 (2020).
Song, W. et al. Imaging RNA polymerase III transcription using a photostable RNA-fluorophore complex. Nat. Chem. Biol. 13, 1187–1194 (2017).
Halstead, J. M. et al. An RNA biosensor for imaging the first round of translation from single cells to living animals. Science 347, 1367–1671 (2015).
Chung, Y. C., Bisht, M., Thuma, J. & Tu, L. C. Single Chromosome Dynamics Reveals Locus-Dependent Dynamics and Chromosome Territory Orientation. J. Cell Sci. 136, jcs260137 (2023).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Tutucci, E., Vera, M. & Singer, R. H. Single-mRNA detection in living S. cerevisiae using a re-engineered MS2 system. Nat. Protoc. 13, 2268–2296 (2018).
Acknowledgements
We thank Andy Sak and Bern Kohler for guidance on quantum yield determination. We thank the Department of Biological Chemistry and Pharmacology at OSU Medical Center for the use of the BD FACSAria Fusion sorter. This work was supported by the NIH 1R01GM143543-01 (D.B.), NIH 4R00GM126810-05 (L.T.), P30 CA016058 (Ohio State University Comprehensive Cancer Center), Ohio State University start up funds (L.T.), STEP fellowship (S.W.) and the Center for RNA Biology at the Ohio State University.
Author information
Authors and Affiliations
Contributions
Y.L., Y.M.L., and S.R. synthesized compounds; Y.L., S.W., Y.C.C., Y.M.L. and S.M. carried out experiments and performed analysis. L.C.T. and D.B. designed experiments and supervised the work.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Peer review
Peer review information
Nature Communications thanks Ling-Ling Chen and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Source data
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Liang, Y., Willey, S., Chung, YC. et al. Intracellular RNA and DNA tracking by uridine-rich internal loop tagging with fluorogenic bPNA. Nat Commun 14, 2987 (2023). https://doi.org/10.1038/s41467-023-38579-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41467-023-38579-2
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.