Abstract
Obesity is associated with chronic, low-grade inflammation, which can disrupt homeostasis within tissue microenvironments. Given the correlation between obesity and relative risk of death from cancer, we investigated whether obesity-associated inflammation promotes metastatic progression. We demonstrate that obesity causes lung neutrophilia in otherwise normal mice, which is further exacerbated by the presence of a primary tumour. The increase in lung neutrophils translates to increased breast cancer metastasis to this site, in a GM-CSF- and IL5-dependent manner. Importantly, weight loss is sufficient to reverse this effect, and reduce serum levels of GM-CSF and IL5 in both mouse models and humans. Our data indicate that special consideration of the obese patient population is critical for effective management of cancer progression.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Quail, D. F. & Joyce, J. A. Microenvironmental regulation of tumor progression and metastasis. Nat. Med. 19, 1423–1437 (2013).
Hanahan, D. & Weinberg, R. A. Hallmarks of cancer: the next generation. Cell 144, 646–674 (2011).
McAllister, S. S. & Weinberg, R. A. The tumour-induced systemic environment as a critical regulator of cancer progression and metastasis. Nat. Cell Biol. 16, 717–727 (2014).
Peinado, H. et al. Pre-metastatic niches: organ-specific homes for metastases. Nat. Rev. Cancer 17, 302–317 (2017).
Grivennikov, S. I., Greten, F. R. & Karin, M. Immunity, inflammation, and cancer. Cell 140, 883–899 (2010).
Calle, E. E. & Kaaks, R. Overweight, obesity and cancer: epidemiological evidence and proposed mechanisms. Nat. Rev. Cancer 4, 579–591 (2004).
Ligibel, J. A. et al. American Society of Clinical Oncology position statement on obesity and cancer. J. Clin. Oncol. 32, 3568–3574 (2014).
Fantuzzi, G. Adipose tissue, adipokines, and inflammation. J. Allergy Clin. Immunol. 115, 911–919 (2005).
Xia, S. et al. Gr-1+ CD11b+ myeloid-derived suppressor cells suppress inflammation and promote insulin sensitivity in obesity. J. Biol. Chem. 286, 23591–23599 (2011).
Kanneganti, T. D. & Dixit, V. D. Immunological complications of obesity. Nat. Immunol. 13, 707–712 (2012).
Iyengar, N. M., Hudis, C. A. & Dannenberg, A. J. Obesity and inflammation: new insights into breast cancer development and progression. Am. Soc. Clin. Oncol. Educ. Book 33, 46–51 (2013).
Iyengar, N. M. et al. Systemic correlates of white adipose tissue inflammation in early-stage breast cancer. Clin. Cancer Res. 22, 2283–2289 (2015).
Coussens, L. M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Ewertz, M. et al. Effect of obesity on prognosis after early-stage breast cancer. J. Clin. Oncol. 29, 25–31 (2011).
Osman, M. A. & Hennessy, B. T. Obesity correlation with metastases development and response to first-line metastatic chemotherapy in breast cancer. Clin. Med. Insights Oncol. 9, 105–112 (2015).
Khandekar, M. J., Cohen, P. & Spiegelman, B. M. Molecular mechanisms of cancer development in obesity. Nat. Rev. Cancer 11, 886–895 (2011).
Weigelt, B., Peterse, J. L. & van ’t Veer, L. J. Breast cancer metastasis: markers and models. Nat. Rev. Cancer 5, 591–602 (2005).
Mancuso, P. Obesity and lung inflammation. J. Appl. Physiol. 108, 722–728 (2010).
Julia, V., Macia, L. & Dombrowicz, D. The impact of diet on asthma and allergic diseases. Nat. Rev. Immunol. 15, 308–322 (2015).
Lackey, D. E. & Olefsky, J. M. Regulation of metabolism by the innate immune system. Nat. Rev. Endocrinol. 12, 15–28 (2016).
Catena, R. et al. Bone marrow-derived Gr1+ cells can generate a metastasis-resistant microenvironment via induced secretion of thrombospondin-1. Cancer Discov. 3, 578–589 (2013).
Liu, Y. et al. Tumor exosomal RNAs promote lung pre-metastatic niche formation by activating alveolar epithelial TLR3 to recruit neutrophils. Cancer Cell 30, 243–256 (2016).
Gao, D. et al. Myeloid progenitor cells in the premetastatic lung promote metastases by inducing mesenchymal to epithelial transition. Cancer Res. 72, 1384–1394 (2012).
Wculek, S. K. & Malanchi, I. Neutrophils support lung colonization of metastasis-initiating breast cancer cells. Nature 528, 413–417 (2015).
Casbon, A. J. et al. Invasive breast cancer reprograms early myeloid differentiation in the bone marrow to generate immunosuppressive neutrophils. Proc. Natl Acad. Sci. USA 112, E566–E575 (2015).
Sharma, S. K. et al. Pulmonary alveolar macrophages contribute to the premetastatic niche by suppressing antitumor T cell responses in the lungs. J. Immunol. 194, 5529–5538 (2015).
Coffelt, S. B. et al. IL-17-producing γδ T cells and neutrophils conspire to promote breast cancer metastasis. Nature 522, 345–348 (2015).
Qian, B. Z. et al. CCL2 recruits inflammatory monocytes to facilitate breast-tumour metastasis. Nature 475, 222–225 (2011).
Kowanetz, M. et al. Granulocyte-colony stimulating factor promotes lung metastasis through mobilization of Ly6G+ Ly6C+ granulocytes. Proc. Natl Acad. Sci. USA 107, 21248–21255 (2010).
Zhang, Y. et al. Positional cloning of the mouse obese gene and its human homologue. Nature 372, 425–432 (1994).
Montgomery, M. K. et al. Mouse strain-dependent variation in obesity and glucose homeostasis in response to high-fat feeding. Diabetologia 56, 1129–1139 (2013).
Gabrilovich, D. I., Ostrand-Rosenberg, S. & Bronte, V. Coordinated regulation of myeloid cells by tumours. Nat. Rev. Immunol. 12, 253–268 (2012).
Delano, M. J. et al. Neutrophil mobilization from the bone marrow during polymicrobial sepsis is dependent on CXCL12 signaling. J. Immunol. 187, 911–918 (2011).
Eash, K. J., Greenbaum, A. M., Gopalan, P. K. & Link, D. C. CXCR2 and CXCR4 antagonistically regulate neutrophil trafficking from murine bone marrow. J. Clin. Invest. 120, 2423–2431 (2010).
Fan, J. & Malik, A. B. Toll-like receptor-4 (TLR4) signaling augments chemokine-induced neutrophil migration by modulating cell surface expression of chemokine receptors. Nat. Med. 9, 315–321 (2003).
Fridlender, Z. G. et al. Polarization of tumor-associated neutrophil phenotype by TGF-β: “N1” versus “N2” TAN. Cancer Cell 16, 183–194 (2009).
Granot, Z. et al. Tumor entrained neutrophils inhibit seeding in the premetastatic lung. Cancer Cell 20, 300–314 (2011).
Bronte, V. et al. IL-4-induced arginase 1 suppresses alloreactive T cells in tumor-bearing mice. J. Immunol. 170, 270–278 (2003).
Kusmartsev, S., Nefedova, Y., Yoder, D. & Gabrilovich, D. I. Antigen-specific inhibition of CD8 + T cell response by immature myeloid cells in cancer is mediated by reactive oxygen species. J. Immunol. 172, 989–999 (2004).
Sinha, P., Clements, V. K., Fulton, A. M. & Ostrand-Rosenberg, S. Prostaglandin E2 promotes tumor progression by inducing myeloid-derived suppressor cells. Cancer Res 67, 4507–4513 (2007).
Nagareddy, P. R. et al. Adipose tissue macrophages promote myelopoiesis and monocytosis in obesity. Cell Metab. 19, 821–835 (2014).
Sinha, P. et al. Proinflammatory S100 proteins regulate the accumulation of myeloid-derived suppressor cells. J. Immunol. 181, 4666–4675 (2008).
Spiegel, A. et al. Neutrophils suppress intraluminal NK cell-mediated tumor cell clearance and enhance extravasation of disseminated carcinoma cells. Cancer Discov. 6, 630–649 (2016).
Yan, H. H. et al. Gr-1+ CD11b+ myeloid cells tip the balance of immune protection to tumor promotion in the premetastatic lung. Cancer Res. 70, 6139–6149 (2010).
Sevenich, L. et al. Analysis of tumour- and stroma-supplied proteolytic networks reveals a brain-metastasis-promoting role for cathepsin S. Nat. Cell Biol. 16, 876–888 (2014).
Herishanu, Y., Rogowski, O., Polliack, A. & Marilus, R. Leukocytosis in obese individuals: possible link in patients with unexplained persistent neutrophilia. Eur. J. Haematol. 76, 516–520 (2006).
Dranoff, G. et al. Involvement of granulocyte-macrophage colony-stimulating factor in pulmonary homeostasis. Science 264, 713–716 (1994).
Stanley, E. et al. Granulocyte/macrophage colony-stimulating factor-deficient mice show no major perturbation of hematopoiesis but develop a characteristic pulmonary pathology. Proc. Natl Acad. Sci. USA 91, 5592–5596 (1994).
Uchida, K. et al. GM-CSF autoantibodies and neutrophil dysfunction in pulmonary alveolar proteinosis. N. Engl. J. Med. 356, 567–579 (2007).
Hoshi, H. et al. IL-5, IL-8 and GM-CSF immunostaining of sputum cells in bronchial asthma and chronic bronchitis. Clin. Exp. Allergy 25, 720–728 (1995).
Lambrecht, B. N. & Hammad, H. The immunology of asthma. Nat. Immunol. 16, 45–56 (2015).
Asquith, K. L. et al. The IL-3/IL-5/GM-CSF common receptor plays a pivotal role in the regulation of Th2 immunity and allergic airway inflammation. J. Immunol. 180, 1199–1206 (2008).
Rosenberg, H. F., Dyer, K. D. & Foster, P. S. Eosinophils: changing perspectives in health and disease. Nat. Rev. Immunol. 13, 9–22 (2013).
Linch, S. N. et al. Interleukin 5 is protective during sepsis in an eosinophil-independent manner. Am. J. Respir. Crit. Care Med. 186, 246–254 (2012).
Hakonarson, H., Maskeri, N., Carter, C., Chuang, S. & Grunstein, M. M. Autocrine interaction between IL-5 and IL-1β mediates altered responsiveness of atopic asthmatic sensitized airway smooth muscle. J. Clin. Invest. 104, 657–667 (1999).
Tournoy, K. G., Kips, J. C. & Pauwels, R. A. The allergen-induced airway hyperresponsiveness in a human-mouse chimera model of asthma is T cell and IL-4 and IL-5 dependent. J. Immunol. 166, 6982–6991 (2001).
Wahl, S. et al. Epigenome-wide association study of body mass index, and the adverse outcomes of adiposity. Nature 541, 81–86 (2017).
Alemán, J. O. et al. Effects of rapid weight loss on systemic and adipose tissue inflammation and metabolism in obese postmenopausal women. J. Endocr. Soc. 1, 625–637 (2017).
Chen, S. B. et al. Serum C-reactive protein and white blood cell count in morbidly obese surgical patients. Obes. Surg. 19, 461–466 (2009).
Cottam, D. et al. Tumor growth factor expression in obesity and changes in expression with weight loss: another cause of increased virulence and incidence of cancer in obesity. Surg. Obes. Relat. Dis. 6, 538–541 (2010).
Molofsky, A. B. et al. Innate lymphoid type 2 cells sustain visceral adipose tissue eosinophils and alternatively activated macrophages. J. Exp. Med. 210, 535–549 (2013).
Brestoff, J. R. et al. Group 2 innate lymphoid cells promote beiging of white adipose tissue and limit obesity. Nature 519, 242–246 (2015).
Fainardi, V., Pisi, G. & Chetta, A. Mepolizumab in the treatment of severe eosinophilic asthma. Immunotherapy 8, 27–34 (2016).
Bhardwaj, P. et al. Caloric restriction reverses obesity-induced mammary gland inflammation in mice. Cancer Prev. Res. 6, 282–289 (2013).
Pietrocola, F. et al. Caloric restriction mimetics enhance anticancer immunosurveillance. Cancer Cell 30, 147–160 (2016).
Botelho, F. M. et al. A mouse GM-CSF receptor antibody attenuates neutrophilia in mice exposed to cigarette smoke. Eur. Respir. J. 38, 285–294 (2011).
Zaynagetdinov, R. et al. Interleukin-5 facilitates lung metastasis by modulating the immune microenvironment. Cancer Res. 75, 1624–1634 (2015).
Binder, C. J. et al. IL-5 links adaptive and natural immunity specific for epitopes of oxidized LDL and protects from atherosclerosis. J. Clin. Invest. 114, 427–437 (2004).
Esper, R. M. et al. Leptin and adiponectin modulate the self-renewal of normal human breast epithelial stem cells. Cancer Prev. Res. 8, 1174–1183 (2015).
Swamydas, M. & Lionakis, M. S. Isolation, purification and labeling of mouse bone marrow neutrophils for functional studies and adoptive transfer experiments. J. Vis. Exp. 77, e50586 (2013).
Acknowledgements
We thank members of the Joyce and Dannenberg laboratories and V. Mittal for insightful comments and discussion. We acknowledge J. O. Alemán for assistance in providing human sera from weight loss trials. We thank H.-W. Wang for originally isolating the PyMT-BL6 cell lines used herein, and F. Klemm and J. Kowal for critically reading the manuscript. This research was supported by the Breast Cancer Research Foundation (J.A.J., A.J.D.), the Ludwig Institute for Cancer Research (J.A.J.), NIH/NCI U54 CA210184-01 (A.J.D.), the Botwinick-Wolfensohn Foundation (in memory of Mr and Mrs Benjamin Botwinick) (A.J.D.), the Sackler Center for Biomedicine and Nutrition Research at The Rockefeller University (P.R.H.), a National Cancer Institute Cancer Center Support Grant awarded to MSKCC (P30 CA008748), and fellowships from the Canadian Institutes of Health Research (D.F.Q., L.A.W.), National Cancer Institute F31CA171384 (O.C.O.), and the American Brain Tumor Association in honour of Joel A. Gingras (L.A.).
Author information
Authors and Affiliations
Contributions
D.F.Q., O.C.O. and J.A.J. conceived the study, designed and interpreted experiments, and wrote the manuscript. D.F.Q., O.C.O., P.B., L.A.W., L.A., M.L.Q., I.-C.C., N.W. and N.B.-C. performed experiments and analysed results. J.W., P.R.H. and A.J.D. provided human sera and blood, and A.J.D. helped design and interpret experiments. J.A.J. supervised the study. All authors commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Adiposity supports lung neutrophilia during obesity.
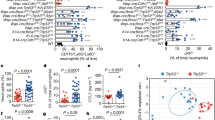
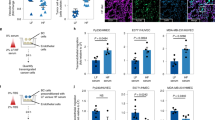
(a) Gating strategy for flow cytometry of CD11b+Gr1+ cells throughout this study, demonstrating that the Gr1+ cells are Ly6CloLy6G+ neutrophils. CD11b+Gr1+ populations are shown as a red overlay upon total CD11b+ cells, graphed on Ly6C (x-axis) by Ly6G (y-axis) staining intensity dot plots. (b) Giemsa stain for FACS-purified CD11b+Gr1+ cells, representing banded and hyper-segmented neutrophils (left), with a smaller subset showing a myelocyte precursor morphology (right). Scale bar = 10 μm. (c) Flow cytometry analysis of total CD45+CD3+ T cells, CD45+CD3+CD4+ helper T cells, and CD45+CD3+CD8+ cytotoxic T cells in the lungs of mice from the diet-induced obesity (DIO) model. n = 4 mice; mean ± s.e.m., 2-tailed unpaired Student’s t-test. (d) Flow cytometry analysis comparing myeloid cell populations in lung, liver and brain in the context of the DIO model. n = 4 mice, matched for each organ presented; mean ± s.e.m., 2-tailed unpaired Student’s t-test. (e) Flow cytometry analysis comparing myeloid cell populations in liver in wild-type (WT) and ob/ob mice. n = 10 mice; mean ± s.e.m., 2-tailed unpaired Student’s t-test. (f) Schematic representation of diet-switch trial. 4 week-old female WT BL6 mice were enrolled on high fat (HF) diet for 15 w, followed by a switch to low fat (LF) diet for an additional 7w.
Supplementary Figure 2 Development and characterization of an immune competent model of breast cancer in the BL6 genetic background.
(a) Cell lines were isolated from MMTV-PyMT mice backcrossed into a BL6 background and tested for their capacity to grow in vivo in WT BL6 animals. Graph shows primary tumor volume over time, measured by calipers biweekly (beginning at 19 d-post injection, when tumors were first palpable), after injecting a panel of seven PyMT-BL6 cell lines into the mammary fat pad of WT BL6 mice (700,000 cells/mouse). n = 5 mice per group; mean ± s.e.m. Three cell lines were selected from this panel for further use in the study: 99LN, 86R2, and 91R2. (b) qRT–PCR analysis of hormone receptor status, and epithelial versus mesenchymal cell marker expression in 99LN, 86R2 and 91R2 cell lines. n = 4 passages per line; mean ± s.e.m. (c) In vitro migration (left) and invasion (right) assays through a Transwell chamber, comparing baseline capacity for metastatic phenotypes in 99LN, 86R2 and 91R2 cell lines. FOV: field of view. n = 20 chambers/group; mean ± s.e.m. Kruskal-Wallis test and Dunn’s multiple comparisons test. (d) H&E image of lung showing spontaneous metastases arising from 99LN primary tumors after ∼2 months. Scale bar = 1,000 μm. (e) Primary tumor volume measured every 10 d over 60 d for 86R2 cells (LF, n = 5 mice; HF, n = 4 mice; mean ± s.e.m., 2-tailed unpaired Student’s t-test) or 99LN cells (LF, n = 10 mice; HF, n = 9 mice; mean ± s.e.m., 2-tailed unpaired Student’s t-test) injected into the mammary fat pad, in the context of the diet-induced obesity (DIO) model (1.5 × 106 cells/mouse). LF: low fat, HF: high fat. (f) Representative H&E image of lungs with spontaneous micro-metastases (black arrows) arising from 99LN mammary fat pad tumors implanted in LF or HF mice after ∼2 months. n = 5 mice/group corresponding to Fig. 2e. Scale bar = 500 μm.
Supplementary Figure 3 The lung neutrophilia and metastasis phenotype from the DIO model is recapitulated in ob/ob mice.
(a) Bioluminescent imaging (BLI) 48 h post-injection of 99LN breast cancer cells into the tail vein of WT or ob/ob animals. Quantification (left) and representative images (right) are presented, showing elevated metastasis in the ob/ob setting. (b) Flow cytometric analysis of lung neutrophils (CD45+CD11b+Ly6CloLy6G+) from trial presented in a. (c) qRT–PCR analysis of Cxcr2, Cxcr4, S100a8, and S100a9 gene expression in FACS-purified neutrophils from lungs of WT or ob/ob mice. For (a–c), n = 5 mice/group; mean ± s.e.m., 2-tailed unpaired Student’s t-test. (d) CFSE assay to assess suppressive activity of LF or HF neutrophils on CD8+ cytotoxic T cells. T cell proliferation was assessed by dilution of CFSE dye (that is, each division results in reduced brightness). Cells were isolated by FACS from n = 6 mice, one representative histogram is shown; all replicates showed no changes between LF and HF groups. (e) Flow cytometry gating strategy for NK cell cytotoxicity assay corresponding to (f). (f) Flow cytometry analysis of the degranulation marker CD107a on NK1.1+ cells after in vitro co-culture with neutrophils isolated from LF or HF peripheral blood. Cells were isolated by FACS from n = 6 mice/group; mean ± s.e.m., one-way ANOVA and Bonferroni’s multiple comparisons test. (g) Flow cytometry analysis of peripheral NK cell cytotoxicity in tumor-bearing mice from the DIO model, as quantified by percent of CD107a+ cells of total NK1.1+ cells in blood. Data are shown as Tukey’s box and whisker plots. All data points were included in statistical analyses, Mann-Whitney test (non-significant), n = 6 mice/group. Box plots represent median and interquartile range while whiskers represent maximum and minimum values excluding outliers. Each symbol represents one mouse.
Supplementary Figure 4 Optimization of Gr1 neutralization in vivo.
(a) Schematic representation of Gr1 neutralization trial in the context of the DIO model and experimental metastasis, as shown in Fig. 3h–i. (b) Validation in non-tumor bearing animals that Gr1 antibody-mediated neutralization is effective at depleting target cells over three days in blood, spleen, and lung. (c) Flow cytometry quantification of CD11b+Ly6G+ cells (left) and CD11b+Ly6C+ cells (right) in blood from the anti-Gr1 neutralization trial. LF, n = 5 mice; HF + IgG, n = 9 mice; HF + αGr1, n = 8 mice; mix-max boxplot, all data points shown, one-way ANOVA and Dunnett’s multiple comparisons test. Box plots represent median and interquartile range while whiskers represent maximum and minimum values excluding outliers. Each symbol represents one mouse.
Supplementary Figure 5 Effect of obesity on circulating factors in mice and humans.
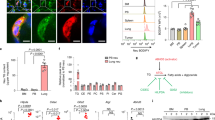
(a) Cross-species cytokine array heatmap showing log2(fold) changes of species-concordant factors that were significantly different between obese mouse (Mu) and human (Hu) serum, versus lean Mu and Hu serum. Red indicates factors that were upregulated in obese serum, and blue indicates factors that were downregulated in obese serum. For a full list of factors and data quantification, see Supplementary Table 2. (b) Quantification of log2(fold) and P-values for cytokine array results as shown in a. (c) ELISA analysis of serum GM-CSF or (d) IL5 in lean and obese human donors. n = 10 donors/group; mean ± s.d., Mann-Whitney test. (e–k) qRT–PCR analysis of mRNA levels for 8 candidate factors identified from cytokine array across different bulk tissues from n = 4 HF-fed mice; mean ± s.e.m.
Supplementary Figure 6 Effect of recombinant GM-CSF treatment on myeloid cell subsets in spleen and bone marrow.
(a) Flow cytometry analysis of CD11b+Gr1+ cells in spleen or (b) bone marrow in WT BL6 animals treated for 5 consecutive days with rGM-CSF versus a PBS control, as shown in Fig. 5a. n = 5 mice/group; mean ± s.e.m., 2-tailed unpaired Student’s t-test. Box plots represent median and interquartile range while whiskers represent maximum and minimum values excluding outliers. Each symbol represents one mouse.
Supplementary Figure 7 IL5 contributes to lung neutrophilia in obesity.
(a) Representative flow cytometry plots (left) and quantification (right) of lung CD3+ T cells in WT BL6 animals after treatment with rIL5, in the trial as shown in Fig. 6b. n = 5 mice/group; mean ± s.e.m., 2-tailed unpaired Student’s t-test. (b) Trial design for rIL5 +/- anti-GM-CSF experiment in WT BL6 mice. Briefly, WT mice were treated for 5 consecutive days with rIL5 prior to tail vein injection with 99LN breast cancer cells at t = 0 d (48 h metastasis assay). At −3 d and −1 d, mice were also treated with a neutralizing antibody against GM-CSF. Mice were treated continually with rmIL5 for the duration of the 48 h metastasis trial. (c) Bioluminescent imaging (BLI) after tail vein injection with breast cancer cells for the trial shown in b. Quantification of photons/second (left) and representative images (right) are presented. n = 10 mice/group; Tukey boxplot, one-way ANOVA and Bonferroni’s multiple comparisons test. (d) Flow cytometry analysis of lung neutrophils from the trial shown in b. n = 5 mice/group from the full trial in b were used for flow analysis; Tukey boxplot, one-way ANOVA and Bonferroni’s multiple comparisons test. (e) Flow cytometry analysis of CD45+CD11b+Ly6CloLy6G+ lung neutrophils in athymic/nude or NOD-scid Il2rgnull (NSG) mice after treatment with either PBS or rIL5 for 5 consecutive days. n = 5 mice/group; mean ± s.e.m., 2-tailed unpaired Student’s t test. (f) Flow cytometry comparison of lung IL5ra + neutrophils, eosinophils, or monocytes in BL6 (n = 5 mice/group), NSG (n = 10 mice/group), or nude mice (n = 10 mice/group). Data are presented as mean ± s.e.m. Box plots represent median and interquartile range while whiskers represent maximum and minimum values excluding outliers. Each symbol represents one mouse.
Supplementary Figure 8 Neutrophil turnover occurs at 8 h post-adoptive cell transfer in WT and ob/ob mice.
(a) Flow cytometry analysis of neutrophil numbers (events per million, y-axis × 103) at 4 h and 8 h post-adoptive transfer, showing that neutrophils turnover by 8 h across all groups. n = 5 mice/recipient group; mean ± s.e.m displayed. (b) Flow cytometry analysis of fluorescently labeled circulating neutrophils 8 h post-adoptive transfer, demonstrating relatively equal representation of both red (ob/ob donor) and green (WT donor) cells. n = 5 mice/recipient group; mean ± s.e.m displayed.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1791 kb)
Supplementary Information
Supplementary Information (PDF 83 kb)
Supplementary Table 1
Supplementary Information (XLS 67 kb)
Supplementary Table 2
Supplementary Information (XLS 117 kb)
Supplementary Table 3
Supplementary Information (XLSX 37 kb)
Supplementary Table 4
Supplementary Information (XLSX 2236 kb)
Rights and permissions
About this article
Cite this article
Quail, D., Olson, O., Bhardwaj, P. et al. Obesity alters the lung myeloid cell landscape to enhance breast cancer metastasis through IL5 and GM-CSF. Nat Cell Biol 19, 974–987 (2017). https://doi.org/10.1038/ncb3578
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3578
This article is cited by
-
A high-cholesterol diet promotes the intravasation of breast tumor cells through an LDL–LDLR axis
Scientific Reports (2024)
-
Made to order: emergency myelopoiesis and demand-adapted innate immune cell production
Nature Reviews Immunology (2024)
-
Locally sourced: site-specific immune barriers to metastasis
Nature Reviews Immunology (2023)
-
Chronic pulmonary bacterial infection facilitates breast cancer lung metastasis by recruiting tumor-promoting MHCIIhi neutrophils
Signal Transduction and Targeted Therapy (2023)
-
SHP2 regulates adipose maintenance and adipocyte-pancreatic cancer cell crosstalk via PDHA1
Journal of Cell Communication and Signaling (2023)