Abstract
The development and maintenance of tissues requires collective cell movement, during which neighbouring cells coordinate the polarity of their migration machineries. Here, we ask how polarity signals are transmitted from one cell to another across symmetrical cadherin junctions, during collective migration. We demonstrate that collectively migrating endothelial cells have polarized VE-cadherin-rich membrane protrusions, ‘cadherin fingers’, which leading cells extend from their rear and follower cells engulf at their front, thereby generating opposite membrane curvatures and asymmetric recruitment of curvature-sensing proteins. In follower cells, engulfment of cadherin fingers occurs along with the formation of a lamellipodia-like zone with low actomyosin contractility, and requires VE-cadherin/catenin complexes and Arp2/3-driven actin polymerization. Lateral accumulation of cadherin fingers in follower cells precedes turning, and increased actomyosin contractility can initiate cadherin finger extension as well as engulfment by a neighbouring cell, to promote follower behaviour. We propose that cadherin fingers serve as guidance cues that direct collective cell migration.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Friedl, P. & Gilmour, D. Collective cell migration in morphogenesis, regeneration and cancer. Nat. Rev. Mol. Cell Biol. 10, 445–457 (2009).
Mayor, R. & Etienne-Manneville, S. The front and rear of collective cell migration. Nat. Rev. Mol. Cell Biol. 17, 97–109 (2016).
Cai, D. et al. Mechanical feedback through E-cadherin promotes direction sensing during collective cell migration. Cell 157, 1146–1159 (2014).
Reffay, M. et al. Interplay of RhoA and mechanical forces in collective cell migration driven by leader cells. Nat. Cell Biol. 16, 217–223 (2014).
Ng, M. R., Besser, A., Danuser, G. & Brugge, J. S. Substrate stiffness regulates cadherin-dependent collective migration through myosin-II contractility. J. Cell Biol. 199, 545–563 (2012).
Khalil, A. A. & Friedl, P. Determinants of leader cells in collective cell migration. Integr. Biol. (Camb). 2, 568–574 (2010).
Das, T. et al. A molecular mechanotransduction pathway regulates collective migration of epithelial cells. Nat. Cell Biol. 17, 276–287 (2015).
Yonemura, S., Itoh, M., Nagafuchi, A. & Tsukita, S. Cell-to-cell adherens junction formation and actin filament organization: similarities and differences between non-polarized fibroblasts and polarized epithelial cells. J. Cell Sci. 108, 127–142 (1995).
Vasioukhin, V., Bauer, C., Yin, M. & Fuchs, E. Directed actin polymerization is the driving force for epithelial cell–cell adhesion. Cell 100, 209–219 (2000).
Millán, J. et al. Adherens junctions connect stress fibres between adjacent endothelial cells. BMC Biol. 8, 11 (2010).
Hoelzle, M. K. & Svitkina, T. The cytoskeletal mechanisms of cell–cell junction formation in endothelial cells. Mol. Biol. Cell 23, 310–323 (2012).
Taguchi, K., Ishiuchi, T. & Takeichi, M. Mechanosensitive EPLIN-dependent remodeling of adherens junctions regulates epithelial reshaping. J. Cell Biol. 194, 643–656 (2011).
Huveneers, S. et al. Vinculin associates with endothelial VE-cadherin junctions to control force-dependent remodeling. J. Cell Biol. 196, 641–652 (2012).
Ando, K. et al. Rap1 potentiates endothelial cell junctions by spatially controlling myosin I activity and actin organization. J. Cell Biol. 202, 901–916 (2013).
Peglion, F., Llense, F. & Etienne-Manneville, S. Adherens junction treadmilling during collective migration. Nat. Cell Biol. 16, 639–651 (2014).
Kametani, Y. & Takeichi, M. Basal-to-apical cadherin flow at cell junctions. Nat. Cell Biol. 9, 92–98 (2007).
Brevier, J., Montero, D., Svitkina, T. & Riveline, D. The asymmetric self-assembly mechanism of adherens junctions: a cellular push-pull unit. Phys. Biol. 5, 16005 (2008).
van Geemen, D. et al. F-actin-anchored focal adhesions distinguish endothelial phenotypes of human arteries and veins. Arterioscler. Thromb. Vasc. Biol. 34, 2059–2067 (2014).
Jakobsson, L. et al. Endothelial cells dynamically compete for the tip cell position during angiogenic sprouting. Nat. Cell Biol. 12, 943–953 (2010).
Bentley, K. et al. The role of differential VE-cadherin dynamics in cell rearrangement during angiogenesis. Nat. Cell Biol. 16, 309–321 (2014).
Angelini, T. E. et al. Glass-like dynamics of collective cell migration. Proc. Natl Acad. Sci. USA 108, 4714–4719 (2011).
Haga, H., Irahara, C., Kobayashi, R., Nakagaki, T. & Kawabata, K. Collective movement of epithelial cells on a collagen gel substrate. Biophys. J. 88, 2250–2256 (2005).
Vitorino, P. & Meyer, T. Modular control of endothelial sheet migration. Genes Dev. 1, 3268–3281 (2008).
Schell, M. J., Erneux, C. & Irvine, R. F. Inositol 1,4,5-trisphosphate 3-kinase A associates with F-actin and dendritic spines via its N terminus. J. Biol. Chem. 276, 37537–37546 (2001).
Peter, B. J. et al. BAR domains as sensors of membrane curvature: the amphiphysin BAR structure. Science 303, 495–499 (2004).
Galic, M. et al. External push and internal pull forces recruit curvature-sensing N-BAR domain proteins to the plasma membrane. Nat. Cell Biol. 14, 874–881 (2012).
Dorland, Y. L. et al. The F-BAR protein pacsin2 inhibits asymmetric VE-cadherin internalization from tensile adherens junctions. Nat. Commun. 7, 12210 (2016).
Haeger, A., Wolf, K., Zegers, M. M. & Friedl, P. Collective cell migration: guidance principles and hierarchies. Trends Cell Biol. 25, 556–566 (2015).
Collins, C. & Nelson, W. J. Running with neighbors: coordinating cell migration and cell–cell adhesion. Curr. Opin. Cell Biol. 36, 62–70 (2015).
Etienne-Manneville, S. Neighborly relations during collective migration. Curr. Opin. Cell Biol. 30C, 51–59 (2014).
Abu Taha, A., Taha, M., Seebach, J. & Schnittler, H.-J. ARP2/3-mediated junction-associated lamellipodia control VE-cadherin-based cell junction dynamics and maintain monolayer integrity. Mol. Biol. Cell 25, 245–256 (2014).
Lou, S. S., Diz-Munoz, A., Weiner, O. D., Fletcher, D. A. & Theriot, J. A. Myosin light chain kinase regulates cell polarization independently of membrane tension or Rho kinase. J. Cell Biol. 209, 275–288 (2015).
Farooqui, R. & Fenteany, G. Multiple rows of cells behind an epithelial wound edge extend cryptic lamellipodia to collectively drive cell-sheet movement. J. Cell Sci. 118, 51–63 (2005).
Asokan, S. B. et al. Mesenchymal chemotaxis requires selective inactivation of myosin II at the leading edge via a noncanonical PLCγ/PKCα pathway. Dev. Cell 31, 747–760 (2014).
Theveneau, E. et al. Chase-and-run between adjacent cell populations promotes directional collective migration. Nat. Cell Biol. 15, 1–12 (2013).
Inoue, T., Heo, W. D., Grimley, J. S., Wandless, T. J. & Meyer, T. An inducible translocation strategy to rapidly activate and inhibit small GTPase signaling pathways. Nat. Methods 2, 415–418 (2005).
Maddox, A. S. & Burridge, K. RhoA is required for cortical retraction and rigidity during mitotic cell rounding. J. Cell Biol. 160, 255–265 (2003).
Matthews, H. K. et al. Changes in Ect2 localization couple actomyosin-dependent cell shape changes to mitotic progression. Dev. Cell 23, 371–383 (2012).
Tatsumoto, T., Xie, X., Blumenthal, R., Okamoto, I. & Miki, T. Human ECT2 is an exchange factor for Rho GTPases, phosphorylated in G2/M phases, and involved in cytokinesis. Cell 147, 921–927 (1999).
Wagner, E. & Glotzer, M. Local RhoA activation induces cytokinetic furrows independent of spindle position and cell cycle stage. J. Cell Biol. 213, 641–649 (2016).
Phng, L.-K. et al. Formin-mediated actin polymerization at endothelial junctions is required for vessel lumen formation and stabilization. Dev. Cell 32, 123–132 (2015).
Wilson, C. W. et al. Rasip1 regulates vertebrate vascular endothelial junction stability through Epac1-Rap1 signaling. Blood 122, 3678–3690 (2013).
Arrieumerlou, C. & Meyer, T. A local coupling model and compass parameter for eukaryotic chemotaxis. Dev. Cell 8, 215–227 (2005).
Swaney, K. F., Huang, C.-H. & Devreotes, P. N. Eukaryotic chemotaxis: a network of signaling pathways controls motility, directional sensing, and polarity. Annu. Rev. Biophys. 39, 265–289 (2010).
Young, L., Sung, J., Stacey, G. & Masters, J. R. Detection of Mycoplasma in cell cultures. Nat. Protoc. 5, 929–934 (2010).
Bajar, B. T. et al. Improving brightness and photostability of green and red fluorescent proteins for live cell imaging and FRET reporting. Sci. Rep. 6, 20889 (2016).
Gibson, D. G. et al. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 6, 343–345 (2009).
Conway, D. E. et al. Fluid shear stress on endothelial cells modulates mechanical tension across VE-cadherin and PECAM-1. Curr. Biol. 23, 1024–1030 (2013).
Wollman, R. & Meyer, T. Coordinated oscillations in cortical actin and Ca2+ correlate with cycles of vesicle secretion. Nat. Cell Biol. 14, 1261–1269 (2012).
Szymczak-Workman, A. L., Vignali, K. M. & Vignali, D. A. A. Design and construction of 2A peptide-linked multicistronic vectors. Cold Spring Harb. Protoc. 2012, 199–204 (2012).
Ferreira, J. P., Overton, K. W. & Wang, C. L. Tuning gene expression with synthetic upstream open reading frames. Proc. Natl Acad. Sci. USA 110, 11284–11289 (2013).
Counter, C. M. et al. Dissociation among in vitro telomerase activity, telomere maintenance, and cellular immortalization. Proc. Natl Acad. Sci. USA 95, 14723–14728 (1998).
Yang, H. W. et al. Cooperative activation of PI3K by Ras and Rho family small GTPases. Mol. Cell 47, 281–290 (2012).
Hart, M. J. et al. Identification of a novel guanine nucleotide exchange factor for the Rho GTPase. J. Biol. Chem. 271, 25452–25458 (1996).
Campeau, E. et al. A versatile viral system for expression and depletion of proteins in mammalian cells. PLoS ONE 4, e6529 (2009).
Tsai, F.-C. et al. A polarized Ca2+, diacylglycerol and STIM1 signalling system regulates directed cell migration. Nat. Cell Biol. 16, 133–144 (2014).
Tsai, F.-C. & Meyer, T. Ca2+ pulses control local cycles of lamellipodia retraction and adhesion along the front of migrating cells. Curr. Biol. 22, 837–842 (2012).
Otsu, N. A threshold selection method from gray-level histograms. IEEE Trans. Syst. Man Cybern. SMC-9, 62–66 (1979).
Edelstein, A., Amodaj, N., Hoover, K., Vale, R. & Stuurman, N. Computer control of microscopes using μManager. Curr. Protoc. Mol. Biol. http://dx.doi.org/10.1002/0471142727.mb1420s92 (2010).
Fiolka, R., Shao, L., Rego, E. H., Davidson, M. W. & Gustafsson, M. G. L. Time-lapse two-color 3D imaging of live cells with doubled resolution using structured illumination. Proc. Natl Acad. Sci. USA 109, 5311–5315 (2012).
Gustafsson, M. G. L. et al. Three-dimensional resolution doubling in wide-field fluorescence microscopy by structured illumination. Biophys. J. 94, 4957–4970 (2008).
Preibisch, S., Saalfeld, S. & Tomancak, P. Globally optimal stitching of tiled 3D microscopic image acquisitions. Bioinformatics 25, 1463–1465 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Acknowledgements
We thank G. Crabtree, M. Lin and X. Liu for providing constructs, M. Teruel for reagents to generate Giardia-diced siRNA, and the Stanford Shared FACS Facility for support. E. Wagner and M. Glotzer generously provided constructs and advice for local Rho activation. We are grateful to S. Collins, D. Garbett, A. Suvrathan, G. Dey and M. Galic for helpful discussions and comments on the manuscript. A.H. was supported by postdoctoral fellowships from the Swiss National Science Foundation and from the Human Frontiers Science Program Organization. This work was supported by NIH grants GM063702 and MH095087.
Author information
Authors and Affiliations
Contributions
A.H. and T.M. conceived the study. A.H. performed all experiments and analysed the data, with the help of L.S. and E.B. for 3D-SIM experiments, with the help of H.W.Y. for synthetic Rac/Rho activation and cadherin finger tracking, and except for SEM analysis, which was done by L.-M.J. F.-C.T. and M.C. contributed MATLAB code for cell tracking, and M.C. analysed data for Fig. 5c–g. A.B. helped to develop the myosin II/F-actin reporter. A.H. and T.M. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
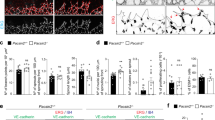
Supplementary Figure 2 Examples of junctional morphologies in HUVEC monolayers, plated at increasing cell density, and stained with Hoechst/phalloidin/anti-CDH5 (VE-cadherin).
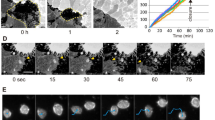
(a) Scale bars, 10 μm. Finger-like VE-cadherin-positive structures were observed across a range of cell densities, reticular junctions at very high densities. (b,c) Scratch wounds were made in HUVEC monolayers to induce collective migration of cells into the cell free area. (b) Cells were fixed 8 h later, stained with Hoechst/phalloidin/anti-CDH5, and imaged by confocal microscopy. Magnifications highlight cadherin fingers pointing away from the rear of leader cells towards their followers. Scale bar, 20 μm. (c) HUVEC stably expressing CDH5-mCitrine and migrating into the cell-free area were image by live-cell microscopy. Magnifications show the presence of cadherin fingers between cells near the wound edge as well as several rows of cells behind. Scale bar, 50 μm. (d) Half-life of cadherin fingers in migrating cells. In time-lapse sequences of HUVEC stably expressing CDH5-mCitrine, captured at 1 min intervals, individual cadherin fingers present in a given cell at t = 0 min were tracked for up to 1 h or until they disappeared. Mean ± S.D. from n = 14 cells (227 cadherin fingers total), from three independent experiments.
Supplementary Figure 3 Additional characterization of cadherin fingers.
(a) 3D-SIM of endothelial cell-cell junctions, additional examples. HUVEC stably expressing CDH5-mEGFP were fixed and stained with fluorescent phalloidin. 3D-SIM revealed polarized cadherin fingers and their underlying actin cytoskeleton. Scale bars, 2 μm. (b) HUVEC expressing CDH5-mCitrine were fixed and stained with the focal adhesion marker anti-paxillin and imaged by confocal microscopy. Line scans across cadherin fingers confirmed that they did not colocalize with paxillin antibody signal. Scale bar, 10 μm. (c) Endothelial intercellular bridges and cadherin fingers do not fuse the cytoplasms or the plasma membranes of neighboring cells. HUVEC stably expressing either Ftractin-mCherry (a soluble maker) or CDH5-mEGFP (a membrane marker) were co-plated and imaged by 3D-SIM. Arrowheads point towards engulfed cadherin fingers or intercellular bridges, where no exchange of soluble or membrane markers was detectable. Scale bars, 2 μm.
Supplementary Figure 4 Effect of Arp2/3 inhibition on single cell velocity.
(a) In cell monolayers either control treated or treated with CK666, cell nuclei were tracked and single cell velocities determined based on nuclear displacements between frames (10 min intervals), averaged over 10 frames. Arp2/3 inhibition caused a dose-dependent inhibition of single cell velocity. Bars are means ± S.D., n = 7 wells from two independent experiments. (b–d) Myosin II activity profiles along the axis of movement – single cell profiles and means. The front-back profiles of (b) mTurquoise-MLC, (c) Ftractin-mCherry and (d) the ratio mTurquoise-MLC/Ftractin-mCherry were measured as in Figure 4. Means and single-cell profiles, normalized to their maximum, of n = 181 cells, from two independent experiments are shown.
Supplementary Figure 5 Correlation between incoming cadherin finger bias and local edge velocity.
Analysis performed as in Figure 5c-g, data for four additional cells are shown.
Supplementary Figure 6 Turning analysis – single cell traces.
The spatiotemporal relationship between incoming/outgoing cadherin fingers and cell turning was analyzed as in Figure 6, but means and single cell traces (n = 33 cells) are shown. (a) Time-course of cell turning aligned to the time point of maximum turning. (b) Time-course of incoming cadherin finger angular bias aligned to the time point of maximum turning. (c) Cross-correlation analysis between incoming cadherin finger angular bias and cell turning shows that incoming cadherin finger angular bias peaks prior to cell turning. (d) Time-course of outgoing cadherin finger angular bias aligned to the time point of maximum turning. (e) Cross-correlation analysis between outgoing cadherin finger angular bias and cell turning shows that outgoing cadherin finger angular bias peaks after cell turning.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1291 kb)
Coordinated movement of HUVEC in monolayers depends on cell density.
HUVEC plated at low and high density were stained with Hoechst and imaged using widefield fluorescence microscopy (4×, 0.2 NA) at 10 min intervals for 4 h. Streams and swirls of coordinately moving HUVEC can be seen in high-density cultures (right, 792 cells/mm2, coordination score 0.24), but not in low-density cultures (274 cells/mm2, coordination score 0.07). Scale bar, 200 μm. Displayed at 15 fps. (AVI 6156 kb)
Polarized cadherin fingers at the interface between collectively migrating cells.
A monolayer of HUVEC stably expressing CDH5-mCitrine was wounded and collective migration was monitored using widefield fluorescence microscopy (20×, 0.75 NA) at 5 min intervals. Cadherin fingers extending from the rear of migrating cells are highlighted by circles. Scale bar, 20 μm. Displayed at 3 fps. (AVI 3199 kb)
Dynamics and half-life of cadherin fingers.
HUVEC stably expressing CDH5-mCitrine were imaged at 1 min intervals using widefield fluorescence microscopy (40×, 1.3 NA). The life-time of individual cadherin fingers ranges from minutes to hours. Scale bar, 10 μm. Displayed at 10 fps. (AVI 6032 kb)
Loss of polarized cadherin fingers in cells treated with the Arp2/3 inhibitor CK666, the ROCK inhibitor Y27632, or Thrombin.
HUVEC stably expressing CDH5-mCitrine were subjected to live-cell imaging (40× 1.3 NA, 1 min intervals) and were either control-treated or treated with CK666 (200 μM), Y27632 (20 μM), or Thrombin (1U/ml) after the first 5 frames. In CK666 and Y27632-treated cells the number of cadherin fingers decreases after drug addition. Thrombin treatment causes serrated, symmetric cell-cell junction. Scale bar, 10 μm. Displayed at 5 fps. (AVI 1870 kb)
The stoichiometric F-actin/myosin II activity reporter Ftractin-mCherry-P2A-mTurquoise-MLC.
In HUVEC stably expressing the reporter, and plated at low density, mTurquoise-MLC is depleted from protrusions, indicating depletion of myosin II activity from protrusive actin networks. Acquired at 5 s intervals, displayed at 8 fps. Scale bar, 10 μm. (AVI 39175 kb)
Depletion of myosin II activity in the front of follower cells.
HUVEC stably expressing Ftractin-mCherry-P2A-mTurquoise-MLC were imaged at 5 s intervals. At the interface between the leader (top) and the follower cell (bottom), mTurquoise-MLC is depleted in the front of the follower cells. Acquired at 15 s intervals, displayed at 8 fps. Scale bar, 10 μm. (AVI 19426 kb)
Depletion of myosin II activity in the front of collectively migrating cells.
HUVEC stably expressing CDH5-mCitrine were co-plated with HUVEC stably expressing Ftractin-mCherry-P2A-mTurquoise-MLC at a ratio of 10:1 and imaged at 30 s intervals. Separate channels, overlay and the ratio of mTurquoise-MLC/Ftractin are shown. A low mTurquoise-MLC/Ftractin-mCherry ratio was observed in the front near where cadherin fingers are present. Scale bar, 10 μm, displayed at 15 fps. (AVI 40225 kb)
Automated tracking of presence and orientation of cadherin fingers.
HUVEC stably expressing CDH5-mCitrine were imaged at 1 min intervals using widefield fluorescence microscopy (40×, 1.3 NA). Automatically detected incoming (green) and outgoing (red) cadherin fingers are shown. Scale bar, 10 μm. Displayed at 8 fps. (AVI 2429 kb)
Incoming cadherin fingers accumulate laterally prior to cell turning.
A monolayer of HUVEC stably expressing CDH5-mCitrine (white) was stained with Hoechst (blue) and imaged at 4 min intervals using widefield fluorescence microscopy (20×, 0.75 NA). The video focuses on a turning cell. The nuclear trajectory and incoming cadherin fingers (+) are marked. Incoming cadherin fingers accumulate laterally and predict the direction of cell turning. Scale bar, 20 μm, displayed at 8 fps. (AVI 23451 kb)
Outgoing cadherin fingers follow turning.
The same turning cell as in video 9 is shown. The nuclear trajectory and outgoing cadherin fingers (+) are marked. Outgoing cadherin fingers do not predict the direction of cell turning. Scale bar, 20 μm, displayed at 8 fps. (AVI 23451 kb)
Increased protrusive actin polymerization is insufficient to induce incoming cadherin fingers.
HUVEC stably expressing CDH5-mCitrine and transiently transfected with Lyn11-FRB and mCherry-FKBP-GEF(TIAM1) were imaged live using a 40× 1.3 NA objective and at 20 s intervals. Rapamycin was added after frame 7 to synthetically activate Rac and induce protrusive actin polymerization. Increased Rac activity was insufficient to increase the number of incoming or outgoing cadherin fingers. Scale bar, 10 μm, displayed at 5 fps. (AVI 22529 kb)
Increased contractility through synthetic Rho activation is sufficient to induce outgoing cadherin fingers.
HUVEC stably expressing CDH5-mCitrine and transiently transfected with Lyn11-FRB and mCherry-FKBP-GEF(TIAM1) were imaged live using a 40× 1.3 NA objective and at 1 min intervals. Rapamycin was added after frame 5 to synthetically activate Rho and induce actomyosin contractility. Increased Rho activity was sufficient to induce outgoing cadherin fingers, while incoming cadherin fingers were lost. Scale bar, 10 μm, displayed at 5 fps. (AVI 30345 kb)
Increased contractility at the onset of mitotic cell rounding is sufficient to induce outgoing cadherin fingers.
HUVEC stably expressing CDH5-mCitrine were subjected to live-cell imaging, using a 40× 1.3NA objective and at 1 min intervals. The increased contractility in the cell undergoing mitotic cell rounding causes a loss of incoming cadherin fingers, but initiates formation of outgoing cadherin fingers between itself and its neighbors. Scale bar, 10 μm, displayed at 5 fps. (AVI 1256 kb)
Local optogenetic Rho activation induces outgoing cadherin fingers.
HUVEC stably expressing CDH5-mCitrine were transiently transfected with Stargazin-GFP-LOVpep and (PDZ)2-mCherry-GEF(LARG). Live-cell imaging was done using a 60× 1.35 NA objective for 60 frames at 30 s intervals using a 514 nm laser for illumination. Transfected cells were identified by mCherry fluorescence (not shown) and Rho activity was locally activated in circular regions using 455 nm laser at frames 11-21. Locally increased contractility induced the formation of outgoing cadherin fingers. Scale bar, 10 μm, displayed at 7 fps. (AVI 33122 kb)
Rights and permissions
About this article
Cite this article
Hayer, A., Shao, L., Chung, M. et al. Engulfed cadherin fingers are polarized junctional structures between collectively migrating endothelial cells. Nat Cell Biol 18, 1311–1323 (2016). https://doi.org/10.1038/ncb3438
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3438
This article is cited by
-
Skin muscle is the initial site of viral replication for arboviral bunyavirus infection
Nature Communications (2024)
-
Tankyrase inhibition interferes with junction remodeling, induces leakiness, and disturbs YAP1/TAZ signaling in the endothelium
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)
-
A robust reprogramming strategy for generating hepatocyte-like cells usable in pharmaco-toxicological studies
Stem Cell Research & Therapy (2023)
-
Functional classification of DDOST variants of uncertain clinical significance in congenital disorders of glycosylation
Scientific Reports (2023)
-
Aberrant gene activation in synovial sarcoma relies on SSX specificity and increased PRC1.1 stability
Nature Structural & Molecular Biology (2023)