Abstract
Half of the human genome is made up of repetitive DNA1. However, mechanisms underlying replication of chromosome regions containing repetitive DNA are poorly understood. We reconstituted replication of defined human chromosome segments using bacterial artificial chromosomes in Xenopus laevis egg extract. Using this approach we characterized the chromatin assembly and replication dynamics of centromeric alpha-satellite DNA. Proteomic analysis of centromeric chromatin revealed replication-dependent enrichment of a network of DNA repair factors including the MSH2–6 complex, which was required for efficient centromeric DNA replication. However, contrary to expectations, the ATR-dependent checkpoint monitoring DNA replication fork arrest could not be activated on highly repetitive DNA due to the inability of the single-stranded DNA binding protein RPA to accumulate on chromatin. Electron microscopy of centromeric DNA and supercoil mapping revealed the presence of topoisomerase I-dependent DNA loops embedded in a protein matrix enriched for SMC2–4 proteins. This arrangement suppressed ATR signalling by preventing RPA hyper-loading, facilitating replication of centromeric DNA. These findings have important implications for our understanding of repetitive DNA metabolism and centromere organization under normal and stressful conditions.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Mirkin, S. M. Expandable DNA repeats and human disease. Nature 447, 932–940 (2007).
Zhao, J., Bacolla, A., Wang, G. & Vasquez, K. M. Non-B DNA structure-induced genetic instability and evolution. Cell. Mol. Life Sci. 67, 43–62 (2010).
Zeman, M. K. & Cimprich, K. A. Causes and consequences of replication stress. Nat. Cell Biol. 16, 2–9 (2014).
Murphy, T. D. & Karpen, G. H. Centromeres take flight: alpha satellite and the quest for the human centromere. Cell 93, 317–320 (1998).
Rudd, M. K., Wray, G. A. & Willard, H. F. The evolutionary dynamics of alpha-satellite. Genome Res. 16, 88–96 (2006).
Bloom, K. S. Centromeric heterochromatin: the primordial segregation machine. Annu. Rev. Genet. 48, 457–484 (2014).
McGarry, T. J. & Kirschner, M. W. Geminin, an inhibitor of DNA replication, is degraded during mitosis. Cell 93, 1043–1053 (1998).
Meijer, L. et al. Biochemical and cellular effects of roscovitine, a potent and selective inhibitor of the cyclin-dependent kinases cdc2, cdk2 and cdk5. Eur. J. Biochem. 243, 527–536 (1997).
Bernad, R. et al. Xenopus HJURP and condensin II are required for CENP-A assembly. J. Cell Biol. 192, 569–582 (2011).
Hirano, T. Condensins: universal organizers of chromosomes with diverse functions. Genes Dev. 26, 1659–1678 (2012).
Cimprich, K. A. & Cortez, D. ATR: an essential regulator of genome integrity. Nat. Rev. Mol. Cell Biol. 9, 616–627 (2008).
Errico, A. & Costanzo, V. Mechanisms of replication fork protection: a safeguard for genome stability. Crit. Rev. Biochem. Mol. Biol. 47, 222–235 (2012).
Hashimoto, Y., Tsujimura, T., Sugino, A. & Takisawa, H. The phosphorylated C-terminal domain of Xenopus Cut5 directly mediates ATR-dependent activation of CHK1. Genes Cells 11, 993–1007 (2006).
Olivera Harris, M. et al. Mismatch repair-dependent metabolism of O-methylguanine-containing DNA in Xenopus laevis egg extracts. DNA Repair (Amst) 28C, 1–7 (2015).
Hashimoto, Y., Ray Chaudhuri, A., Lopes, M. & Costanzo, V. Rad51 protects nascent DNA from Mre11-dependent degradation and promotes continuous DNA synthesis. Nat. Struct. Mol. Biol. 17, 1305–1311 (2010).
Furuyama, T. & Henikoff, S. Centromeric nucleosomes induce positive DNA supercoils. Cell 138, 104–113 (2009).
Bermudez, I., Garcia-Martinez, J., Perez-Ortin, J. E. & Roca, J. A method for genome-wide analysis of DNA helical tension by means of psoralen-DNA photobinding. Nucleic Acids Res. 38, e182 (2010).
Naughton, C. et al. Transcription forms and remodels supercoiling domains unfolding large-scale chromatin structures. Nat. Struct. Mol. Biol. 20, 387–395 (2013).
Ray Chaudhuri, A. et al. Topoisomerase I poisoning results in PARP-mediated replication fork reversal. Nat. Struct. Mol. Biol. 19, 417–423 (2012).
Vos, S. M., Tretter, E. M., Schmidt, B. H. & Berger, J. M. All tangled up: how cells direct, manage and exploit topoisomerase function. Nat. Rev. Mol. Cell Biol. 12, 827–841 (2011).
Costanzo, V., Paull, T., Gottesman, M. & Gautier, J. Mre11 assembles linear DNA fragments into DNA damage signaling complexes. PLoS Biol. 2, E110 (2004).
Hashimoto, Y., Puddu, F. & Costanzo, V. RAD51- and MRE11-dependent reassembly of uncoupled CMG helicase complex at collapsed replication forks. Nat. Struct. Mol. Biol. 19, 17–24 (2012).
Slean, M. M., Panigrahi, G. B., Ranum, L. P. & Pearson, C. E. Mutagenic roles of DNA ‘repair’ proteins in antibody diversity and disease-associated trinucleotide repeat instability. DNA Repair (Amst) 7, 1135–1154 (2008).
Burdova, K., Mihaljevic, B., Sturzenegger, A., Chappidi, N. & Janscak, P. The mismatch-binding factor MutSβ can mediate ATR activation in response to DNA double-strand breaks. Mol. Cell 59, 603–614 (2015).
Koch, J. Neocentromeres and alpha satellite: a proposed structural code for functional human centromere DNA. Hum. Mol. Genet. 9, 149–154 (2000).
Jonstrup, A. T. et al. Hairpin structures formed by alpha satellite DNA of human centromeres are cleaved by human topoisomerase IIα. Nucleic Acids Res. 36, 6165–6174 (2008).
Lupardus, P. J., Byun, T., Yee, M. C., Hekmat-Nejad, M. & Cimprich, K. A. A requirement for replication in activation of the ATR-dependent DNA damage checkpoint. Genes Dev. 16, 2327–2332 (2002).
Voineagu, I., Freudenreich, C. H. & Mirkin, S. M. Checkpoint responses to unusual structures formed by DNA repeats. Mol. Carcinog. 48, 309–318 (2009).
Nasmyth, K. Disseminating the genome: joining, resolving, and separating sister chromatids during mitosis and meiosis. Annu. Rev. Genet. 35, 673–745 (2001).
Sanborn, A. L. et al. Chromatin extrusion explains key features of loop and domain formation in wild-type and engineered genomes. Proc. Natl Acad. Sci. USA 112, E6456–E6465 (2015).
Dunleavy, E. M. et al. HJURP is a cell-cycle-dependent maintenance and deposition factor of CENP-A at centromeres. Cell 137, 485–497 (2009).
Bermejo, R. et al. The replication checkpoint protects fork stability by releasing transcribed genes from nuclear pores. Cell 146, 233–246 (2011).
Palm, W. & de Lange, T. How shelterin protects mammalian telomeres. Annu. Rev. Genet. 42, 301–334 (2008).
Amiard, S. et al. A topological mechanism for TRF2-enhanced strand invasion. Nat. Struct. Mol. Biol. 14, 147–154 (2007).
Balestrini, A., Cosentino, C., Errico, A., Garner, E. & Costanzo, V. GEMC1 is a TopBP1-interacting protein required for chromosomal DNA replication. Nat. Cell Biol. 12, 484–491 (2010).
Errico, A. et al. Tipin/Tim1/And1 protein complex promotes Pol alpha chromatin binding and sister chromatid cohesion. EMBO J. 28, 3681–3692 (2009).
Marheineke, K. & Hyrien, O. Control of replication origin density and firing time in Xenopus egg extracts: role of a caffeine-sensitive, ATR-dependent checkpoint. J. Biol. Chem. 279, 28071–28081 (2004).
Costanzo, V. et al. Mre11 protein complex prevents double-strand break accumulation during chromosomal DNA replication. Mol. Cell 8, 137–147 (2001).
Acknowledgements
We thank H. Mahbubani and F. Pezzimenti for technical support with Xenopus laevis, J. Gannon for antibody production, M. Lopes for helpful discussions, J. Jiricny for MMR antibodies and A. Losada for CENP-A antibodies. This work was funded by the Associazione Italiana per la Ricerca sul Cancro (AIRC), a European Research Council (ERC) consolidator grant (614541), a Giovanni Armenise-Harvard foundation award, the EPIGEN Progetto Bandiera (4.7), AICR-Worldwide Cancer Research (13-0026) and a Fondazione Telethon (GGP13-071) grant awarded to V.C. A.B. is funded by AIRC grant IG 14578.
Author information
Authors and Affiliations
Contributions
A.A. and V.S. prepared the materials and carried out the experiments. P.S. and A.B. carried out MS analysis. A.A. and V.C. designed, analysed and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 BACs replication in egg extract.
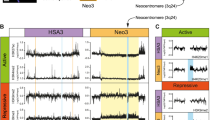
(a) L10 DNA BAC or Sperm chromatin incubated in Xenopus egg extract with [32P]dCTP. DNA synthesis was monitored by measuring the percentage of radiolabelled nucleotide incorporation relative to the input DNA. Error bars represent ±s.d. of the mean. n = 3 independent experiments; P < 0.05 when comparing L10 and B18 mean values; unpaired two-tailed t-test. (b) L10 BAC was replicated in egg extract in the presence of geminin or roscovitine and [32P]dCTP. Incorporation of radiolabelled nucleotides was monitored by autoradiography and relative intensities were plotted on the graph, considering maximum values as 1. Error bars represent ±s.d. of the mean. n = 3 independent experiments; P < 0.001 when comparing mean values for all the indicated treatments; one-way Anova. (c) Replicating L10 BAC was pulse labelled with [32P]dCTP for 20 min and samples were fixed and analysed at the indicated time by autoradiography. Quantification of one representative experiment is shown. (d) Chromatin was isolated at different times using L10 BAC DNA incubated in egg extracts treated with geminin or roscovitine and then analysed by WB as indicated. (e) Graph showing incorporation of [32P]dCTP in fractions derived from CsCl gradient centrifugation of sperm and L10 BAC DNA replicated in egg extract in the presence of [32P]dCTP and BrdUTP. Heavy-Light (HL) and Heavy-Heavy (HH) DNA fractions are indicated. Quantification performed as in (b) from one representative experiment is shown. (f) Molecular combing of RP11-1051L10 (L10) and RP11-5B18 (B18) BACs fully replicated in the presence of digoxygenin-dUTP (green). (g) Table representing the nomenclature, the size and the chromosome regions of the BACs used in the present study. (h) Chromosome mapping of the centromere BACs. (i) Graph showing the percentage of alpha-satellite sequences in each centromeric BAC used. (j) Graph showing the GC base content in the BAC DNA sequences. (k) Replication of B18 and L10 DNA BACs incubated for five hours in Xenopus egg extracts supplemented with 32PdCTP. DNA synthesis was monitored by measuring the percentage of radiolabelled nucleotides incorporation relative to the input DNA. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.001 when comparing L10 and B18 mean values; unpaired two-tailed t-test. (l) DNA replication of the different BACs DNA described in (g). Average percentage of 32PdCTP incorporation. L10 values were considered as 100%. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.001 when comparing DNA replication mean values for all the indicated BACs; Two-way Anova. (m,n) Control and centromeric chromatin was isolated at the indicated times and then analysed by WB using the indicated antibodies. (o) Analysis of the average inter-origin distance (IODs) relative to at least hundred fibres of DNA in replicated L10 and B18 BACs. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.001 when comparing L10 and B10 mean values; unpaired two tailed t-test.
Supplementary Figure 2 Proteomic analysis and validation of control and centromeric chromatin.
(a) Scheme of MS experiments (see text). (b) STRING analysis of DNA repair factors enriched on centromeric chromatin. (c) WB analysis of L10 and B18 chromatin isolated at 150 min from DNA addition to egg extract and probed with the indicated antibodies. MSH2 antibodies used here were different from the ones used in Fig. 3. Graphs showing TopBP1 (d) and MSH2 (e) relative abudance on control L10 and centromeric B18 chromatin compared to ATR. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.05 when comparing L10 and B10 mean values for TopBp1-ATR and MSH2-ATR ratios; One-way Anova.
Supplementary Figure 3 Visualization by EM of L10 and B18 DNA intermediates.
(a) EM of a segment of L10 DNA isolated after 150 minutes incubation in interphase egg extract. (b) EM of intact circular centromeric B18 DNA molecule not incubated in egg extract. (c) A typical replication bubble made of doubled stranded DNA (dsDNA). (d) Typical ssDNA bubble observed on centromeric b18 DNA. Differences in ssDNA and dsDNA fibre thickness can be appreciated. (e) Graph showing the frequency of ssDNA bubbles for each mega base of non-centromeric DNA (L10) or centromeric B18 DNA incubated in egg extracts that were untreated (B18) or treated with geminin (B18 + Geminin). Experiments were repeated three times and one mega base of total DNA was scored. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.001 when comparing L10, B10 and B10 + Geminin mean values; One-way Anova. (f) Typical replication fork intermediate present on B18 DNA.
Supplementary Figure 4 DNA isolation procedure for EM.
(a) Nuclei were isolated at different times from addition of L10 or B18 BAC DNA to egg extract and subjected to psoralen photo-crosslinking with 365 nm UV light. Chromatin was completely digested with 1 mg ml−1 Proteinase K for 2 h at 37 °C. DNA was isolated by phenol:chloroform extraction (25:25 v v−1) and spread in the presence of 1.5% formamide on carbon coated EM grids, which were subjected to rotary shadowing. (b) Scheme showing DNA positive supercoil (+) mediated inhibition of psoralen crosslinking, resulting in ssDNA bubbles after melting in mild denaturing conditions used for the EM preparation.
Supplementary Figure 5 Condensins accumulation and structural arrangement of centromeric chromatin.
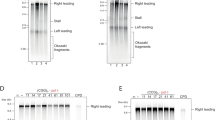
(a) WB analysis showing SMC2 loading on L10 and B18 chromatin isolated at 150 and 300 min from BAC DNA addition to egg extract. MCM7 was used as loading control (b) Graphs showing SMC2 relative abundance on control L10 and centromeric B18 chromatin compared to MCM7. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.01 when comparing L10 and B18 mean values for the indicated times; One-way Anova. (c) Tunel assay showing 32PdCTP incorporation catalysed by Terminal Transferase (TdT) on DNA derived from L10 and B18 nuclei isolated from egg extracts that were untreated, treated with topotecan (TPT) or HAEIII restriction enzyme. The latter treatment was used as positive control of double strand breaks formation. Error bars represent ±s.d. of the mean. n = 3 experiments; P < 0.05 when comparing L10 and B18 mean values for all the indicated treatments; One-way Anova. (d) WB analysis showing SMC2 loading on B18 chromatin isolated at 0 and 300 min from addition of DNA to egg extract supplemented with topotecan (TPT) (+) or left untreated (−). MCM7 was used as loading control. A representative result is shown. (e) Modified EM protocol used to analyse BAC chromatin. Nuclei were isolated at different times from addition of L10 or B18 DNA to egg extracts and subjected to psoralen photo-crosslinking with 365 nm UV light. Chromatin was treated with 1 mg ml−1 Proteinase K for 5 min at 30 °C. DNA was isolated by phenol:chloroform extraction (25:5 v v−1) and spread in the presence of 1.5% formamide on carbon coated EM grids, which were subjected to rotary shadowing. (f) WB showing levels of SMC2 protein present on B18 derived centromeric chromatin following procedures for EM staining performed at low (5 min) and high (2 h) proteinase K treatment (see Methods). A typical result of is shown.
Supplementary information
Supplementary Information
Supplementary Information (PDF 1347 kb)
Supplementary Table 1
Total proteins identified by MS. (XLSX 2953 kb)
Supplementary Table 2
Proteins differentially represented on centromeric and non-centromeric chromatin in biological replicates. (XLSX 118 kb)
Supplementary Table 3
Proteins differentially represented on centromeric and non-centromeric chromatin considered for heatmap. (XLSX 12 kb)
Rights and permissions
About this article
Cite this article
Aze, A., Sannino, V., Soffientini, P. et al. Centromeric DNA replication reconstitution reveals DNA loops and ATR checkpoint suppression. Nat Cell Biol 18, 684–691 (2016). https://doi.org/10.1038/ncb3344
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3344
This article is cited by
-
FANCJ promotes PARP1 activity during DNA replication that is essential in BRCA1 deficient cells
Nature Communications (2024)
-
Dicentric chromosomes are resolved through breakage and repair at their centromeres
Chromosoma (2024)
-
Single-molecule imaging reveals replication fork coupled formation of G-quadruplex structures hinders local replication stress signaling
Nature Communications (2021)
-
The epigenetic regulator LSH maintains fork protection and genomic stability via MacroH2A deposition and RAD51 filament formation
Nature Communications (2021)
-
iDRP-PseAAC: Identification of DNA Replication Proteins Using General PseAAC and Position Dependent Features
International Journal of Peptide Research and Therapeutics (2021)