Abstract
Pancreatic ductal adenocarcinomas (PDACs) are highly metastatic with poor prognosis, mainly due to delayed detection. We hypothesized that intercellular communication is critical for metastatic progression. Here, we show that PDAC-derived exosomes induce liver pre-metastatic niche formation in naive mice and consequently increase liver metastatic burden. Uptake of PDAC-derived exosomes by Kupffer cells caused transforming growth factor β secretion and upregulation of fibronectin production by hepatic stellate cells. This fibrotic microenvironment enhanced recruitment of bone marrow-derived macrophages. We found that macrophage migration inhibitory factor (MIF) was highly expressed in PDAC-derived exosomes, and its blockade prevented liver pre-metastatic niche formation and metastasis. Compared with patients whose pancreatic tumours did not progress, MIF was markedly higher in exosomes from stage I PDAC patients who later developed liver metastasis. These findings suggest that exosomal MIF primes the liver for metastasis and may be a prognostic marker for the development of PDAC liver metastasis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Saif, M. W. Pancreatic neoplasm in 2011: an update. JOP 12, 316–321 (2011).
Chan, A., Diamandis, E. P. & Blasutig, I. M. Strategies for discovering novel pancreatic cancer biomarkers. J. Proteomics 81, 126–134 (2013).
Fesinmeyer, M. D., Austin, M. A., Li, C. I., De Roos, A. J. & Bowen, D. J. Differences in survival by histologic type of pancreatic cancer. Cancer Epidemiol. Biomarkers Prev. 14, 1766–1773 (2005).
Arscott, W. T. & Camphausen, K. A. EGFR isoforms in exosomes as a novel method for biomarker discovery in pancreatic cancer. Biomarkers Med. 5, 821 (2011).
Record, M., Carayon, K., Poirot, M. & Silvente-Poirot, S. Exosomes as new vesicular lipid transporters involved in cell–cell communication and various pathophysiologies. Biochim. Biophys. Acta 1841, 108–120 (2014).
El Andaloussi, S., Mager, I., Breakefield, X. O. & Wood, M. J. Extracellular vesicles: biology and emerging therapeutic opportunities. Nat. Rev. Drug Discov. 12, 347–357 (2013).
Choi, D. S., Kim, D. K., Kim, Y. K. & Gho, Y. S. Proteomics, transcriptomics and lipidomics of exosomes and ectosomes. Proteomics 13, 1554–1571 (2013).
Martins, V. R., Dias, M. S. & Hainaut, P. Tumor-cell-derived microvesicles as carriers of molecular information in cancer. Curr. Opin. Oncol. 25, 66–75 (2013).
Peinado, H., Lavotshkin, S. & Lyden, D. The secreted factors responsible for pre-metastatic niche formation: old sayings and new thoughts. Semin. Cancer Biol. 21, 139–146 (2011).
Thakur, B. K. et al. Double-stranded DNA in exosomes: a novel biomarker in cancer detection. Cell Res. 24, 766–769 (2014).
Tetta, C., Ghigo, E., Silengo, L., Deregibus, M. C. & Camussi, G. Extracellular vesicles as an emerging mechanism of cell-to-cell communication. Endocrine 44, 11–19 (2013).
Valadi, H. et al. Exosome-mediated transfer of mRNAs and microRNAs is a novel mechanism of genetic exchange between cells. Nat. Cell Biol. 9, 654–659 (2007).
Zoller, M. Pancreatic cancer diagnosis by free and exosomal miRNA. World J. Gastrointest. Pathophysiol. 4, 74–90 (2013).
Kaplan, R. N. et al. VEGFR1-positive haematopoietic bone marrow progenitors initiate the pre-metastatic niche. Nature 438, 820–827 (2005).
Sceneay, J., Smyth, M. J. & Moller, A. The pre-metastatic niche: finding common ground. Cancer Metastasis Rev. 32, 449–464 (2013).
Peinado, H. et al. Melanoma exosomes educate bone marrow progenitor cells toward a pro-metastatic phenotype through MET. Nat. Med. 18, 883–891 (2012).
Hood, J. L., San, R. S. & Wickline, S. A. Exosomes released by melanoma cells prepare sentinel lymph nodes for tumor metastasis. Cancer Res. 71, 3792–3801 (2011).
Corbett, T. H. et al. Induction and chemotherapeutic response of two transplantable ductal adenocarcinomas of the pancreas in C57BL/6 mice. Cancer Res. 44, 717–726 (1984).
Little, E. C. et al. Novel immunocompetent murine models representing advanced local and metastatic pancreatic cancer. J. Surg. Res. 176, 359–366 (2012).
Hingorani, S. R. et al. Trp53R172H and KrasG12D cooperate to promote chromosomal instability and widely metastatic pancreatic ductal adenocarcinoma in mice. Cancer Cell 7, 469–483 (2005).
Rhim, A. D. et al. EMT and dissemination precede pancreatic tumor formation. Cell 148, 349–361 (2012).
Achyut, B. R. & Yang, L. Transforming growth factor-β in the gastrointestinal and hepatic tumor microenvironment. Gastroenterology 141, 1167–1178 (2011).
Hayashi, H. & Sakai, T. Biological significance of local TGF-β activation in liver diseases. Front. Physiol. 3, 12 (2012).
Wight, T. N. & Potter-Perigo, S. The extracellular matrix: an active or passive player in fibrosis? Am. J. Physiol. Gastrointest. Liver Physiol. 301, G950–G955 (2011).
Gressner, A. M., Weiskirchen, R., Breitkopf, K. & Dooley, S. Roles of TGF-β in hepatic fibrosis. Front. Biosci. 7, d793–d807 (2002).
Cong, M., Iwaisako, K., Jiang, C. & Kisseleva, T. Cell signals influencing hepatic fibrosis. Int. J. Hepatol. 2012, 158547 (2012).
Kawelke, N. et al. Fibronectin protects from excessive liver fibrosis by modulating the availability of and responsiveness of stellate cells to active TGF-β. PLoS ONE 6, e28181 (2011).
Xu, G. et al. Gene expression and synthesis of fibronectin isoforms in rat hepatic stellate cells. Comparison with liver parenchymal cells and skin fibroblasts. J. Pathol. 183, 90–98 (1997).
Tojo, M. et al. The ALK-5 inhibitor A-83-01 inhibits Smad signaling and epithelial-to-mesenchymal transition by transforming growth factor-β. Cancer Sci. 96, 791–800 (2005).
Duffield, J. S. et al. Selective depletion of macrophages reveals distinct, opposing roles during liver injury and repair. J. Clin. Invest. 115, 56–65 (2005).
Lau, C. et al. Role of pancreatic cancer-derived exosomes in salivary biomarker development. J. Biol. Chem. 288, 26888–26897 (2013).
Heinrichs, D. et al. Macrophage migration inhibitory factor (MIF) exerts antifibrotic effects in experimental liver fibrosis via CD74. Proc. Natl Acad. Sci. USA 108, 17444–17449 (2011).
Barnes, M. A. et al. Macrophage migration inhibitory factor contributes to ethanol-induced liver injury by mediating cell injury, steatohepatitis, and steatosis. Hepatology 57, 1980–1991 (2013).
Funamizu, N. et al. Macrophage migration inhibitory factor induces epithelial to mesenchymal transition, enhances tumor aggressiveness and predicts clinical outcome in resected pancreatic ductal adenocarcinoma. Int. J. Cancer 132, 785–794 (2013).
Nanji, A. A. et al. Macrophage migration inhibitory factor expression in male and female ethanol-fed rats. J. Interferon Cytokine Res. 21, 1055–1062 (2001).
Shin, H. N., Moon, H. H. & Ku, J. L. Stromal cell-derived factor-1α and macrophage migration-inhibitory factor induce metastatic behavior in CXCR4-expressing colon cancer cells. Int. J. Mol. Med. 30, 1537–1543 (2012).
Zhang, H. Y. et al. Macrophage migration inhibitory factor expression correlates with inflammatory changes in human chronic hepatitis B infection. Liver Int. 25, 571–579 (2005).
Adamali, H. et al. Macrophage migration inhibitory factor enzymatic activity, lung inflammation, and cystic fibrosis. Am. J. Respir. Crit. Care Med. 186, 162–169 (2012).
Kobayashi, S., Nishihira, J., Watanabe, S. & Todo, S. Prevention of lethal acute hepatic failure by antimacrophage migration inhibitory factor antibody in mice treated with bacille Calmette-Guerin and lipopolysaccharide. Hepatology 29, 1752–1759 (1999).
Yaddanapudi, K. et al. Control of tumor-associated macrophage alternative activation by macrophage migration inhibitory factor. J. Immunol. 190, 2984–2993 (2013).
Chen, P. F. et al. ISO-1, a macrophage migration inhibitory factor antagonist, inhibits airway remodeling in a murine model of chronic asthma. Mol. Med. 16, 400–408 (2010).
Javle, M. et al. Biomarkers of TGF-β signaling pathway and prognosis of pancreatic cancer. PLoS ONE 9, e85942 (2014).
Ellermeier, J. et al. Therapeutic efficacy of bifunctional siRNA combining TGF-β1 silencing with RIG-I activation in pancreatic cancer. Cancer Res. 73, 1709–1720 (2013).
Gaspar, N. J. et al. Inhibition of transforming growth factor β signaling reduces pancreatic adenocarcinoma growth and invasiveness. Mol. Pharmacol. 72, 152–161 (2007).
Melisi, D. et al. LY2109761, a novel transforming growth factor β receptor type I and type II dual inhibitor, as a therapeutic approach to suppressing pancreatic cancer metastasis. Mol. Cancer Ther. 7, 829–840 (2008).
Pickup, M., Novitskiy, S. & Moses, H. L. The roles of TGFβ in the tumour microenvironment. Nat. Rev. Cancer 13, 788–799 (2013).
Ijichi, H. et al. Aggressive pancreatic ductal adenocarcinoma in mice caused by pancreas-specific blockade of transforming growth factor-β signaling in cooperation with active Kras expression. Genes Dev. 20, 3147–3160 (2006).
Hezel, A. F. et al. TGF-β and αvβ6 integrin act in a common pathway to suppress pancreatic cancer progression. Cancer Res. 72, 4840–4845 (2012).
Bayon, L. G. et al. Role of Kupffer cells in arresting circulating tumor cells and controlling metastatic growth in the liver. Hepatology 23, 1224–1231 (1996).
Kruse, J. et al. Macrophages promote tumour growth and liver metastasis in an orthotopic syngeneic mouse model of colon cancer. Int. J. Colorectal Dis. 28, 1337–1349 (2013).
Wen, S. W., Ager, E. I. & Christophi, C. Bimodal role of Kupffer cells during colorectal cancer liver metastasis. Cancer Biol. Ther. 14, 606–613 (2013).
Grzesiak, J. J. et al. Knockdown of the β(1) integrin subunit reduces primary tumor growth and inhibits pancreatic cancer metastasis. Int. J. Cancer 129, 2905–2915 (2011).
Saito, N. et al. Inhibition of hepatic metastasis in mice treated with cell-binding domain of human fibronectin and angiogenesis inhibitor TNP-470. Int. J. Clin. Oncol. 6, 215–220 (2001).
Zvibel, I., Halpern, Z. & Papa, M. Extracellular matrix modulates expression of growth factors and growth-factor receptors in liver-colonizing colon-cancer cell lines. Int. J. Cancer 77, 295–301 (1998).
Porembka, M. R. et al. Pancreatic adenocarcinoma induces bone marrow mobilization of myeloid-derived suppressor cells which promote primary tumor growth. Cancer Immunol. Immunother. 61, 1373–1385 (2012).
Yamamoto, M. et al. TSU68 prevents liver metastasis of colon cancer xenografts by modulating the premetastatic niche. Cancer Res. 68, 9754–9762 (2008).
Zhang, Y., Davis, C., Ryan, J., Janney, C. & Pena, M. M. Development and characterization of a reliable mouse model of colorectal cancer metastasis to the liver. Clin. Exp. Metastasis 30, 903–918 (2013).
Seubert, B. et al. Tissue inhibitor of metalloproteinases (TIMP)-1 creates a premetastatic niche in the liver through SDF-1/CXCR4-dependent neutrophil recruitment in mice. Hepatology 61, 238–248 (2015).
Kato, R. et al. A new type of antimetastatic peptide derived from fibronectin. Clin. Cancer Res. 8, 2455–2462 (2002).
Bissell, D. M. Therapy for hepatic fibrosis: revisiting the preclinical models. Clin. Res. Hepatol. Gastroenterol. 35, 521–525 (2011).
Korpal, M. & Kang, Y. Targeting the transforming growth factor-β signalling pathway in metastatic cancer. Eur. J. Cancer 46, 1232–1240 (2010).
Noy, R. & Pollard, J. W. Tumor-associated macrophages: from mechanisms to therapy. Immunity 41, 49–61 (2014).
Gu, G., Brown, J. R. & Melton, D. A. Direct lineage tracing reveals the ontogeny of pancreatic cell fates during mouse embryogenesis. Mech. Dev. 120, 35–43 (2003).
Zhong, S. et al. High throughput illuma strand-specific RNA sequencing library preparation. Cold Spring Harb. Protoc. 2011, 940–949 (2011).
Sakai, T. et al. Plasma fibronectin supports neuronal survival and reduces brain injury following transient focal cerebral ischemia but is not essential for skin-wound healing and hemostasis. Nat. Med. 7, 324–330 (2001).
Suemizu, H. et al. A versatile technique for the in vivo imaging of human tumor xenografts using near-infrared fluorochrome-conjugated macromolecule probes. PLoS ONE 8, e82708 (2013).
Morikawa, K., Walker, S. M., Jessup, J. M. & Fidler, I. J. In vivo selection of highly metastatic cells from surgical specimens of different primary human colon carcinomas implanted into nude mice. Cancer Res. 48, 1943–1948 (1988).
Acknowledgements
We thank D. L. Bajor (Vonderheide laboratory, University of Pennsylvania) for the gift of the R6560B cells. We thank L. Bojmar for carefully reviewing the paper. We thank S. Rudchenko and M. Barbu-Stevanovic at the Hospital for Special Surgery Fannie E. Rippel Foundation Flow Cytometry Core Facility for expert flow cytometry. We are supported by grants from the Children’s Cancer and Blood Foundation (H.P., D.L.), Manning Foundation (D.L.), Hartwell Foundation (D.L.), Champalimaud Foundation (D.L.), Fundacao para a Ciencia e a Tecnologia (D.L.), Nancy C and Daniel P Paduano Foundation (H.P., D.L.), Mary Kay Foundation (D.L.), Pediatric Oncology Experimental Therapeutic Investigator Consortium (D.L.), James Paduano Foundation (D.L., H.P.), Melanoma Research Alliance (H.P.), Sohn Conference Foundation (H.P.), Beth Tortolani Foundation (D.L., J.B.), Malcolm Hewitt Weiner Foundation (D.L.), Jose Carreras Leukemia Foundation (B.K.T.), Theodore Rapp Foundation (D.L.), American Hellenic Educational Progressive Association 5th District Cancer Research Foundation (D.L.), Charles and Marjorie Holloway Foundation (J.B.), Sussman Family Fund (J.B.), Lerner Foundation (J.B.), Breast Cancer Alliance (J.B.), and Manhasset Women’s Coalition Against Breast Cancer (J.B.).
Author information
Authors and Affiliations
Contributions
B.C-S. developed the hypothesis, designed the experimental approach, performed experiments, analysed the data, coordinated the project and wrote the manuscript. N.M.A. conducted experiments. S.S., G.R. and T-L.S. performed immunostaining. H.Z. and Y.H. extracted RNA. B.K.T. and A.B. performed western blots. A.H., T-M.T., C.W. and Y.A. maintained mouse colonies. M.T.M. and H.M. performed proteomic analysis. J.X. and T.Z. processed samples and analysed RNA sequencing data. G.G-S. and C.W. processed human samples. P.M.G., M.A.H., K.J.L., I.M.B.L., E.H.K., A.J.O., J.H., A.D., M.J., K.M., S.K.B. and W.R.J. collected patient samples and managed clinical records. S.H.E. contributed to writing the manuscript. R.E.S., I.M., H.P. and B.Z.S. contributed to hypothesis generation, experimental design and data interpretation. J.B. coordinated the project and interpreted data. D.L. conceived the hypothesis, led the project, interpreted data and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Morphological and functional characterization of pancreatic ductal adenocarcinoma exosomes.
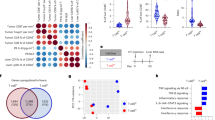
(a) A representative electron microscope image of exosomes isolated from PAN02 conditioned media. Scale bar, 100 nm. (b) Evaluation of liver metastasis by liver weight (grams) in mice pre-educated with exosomes isolated from a PKCY-mouse model tumor-derived primary cell line (PKCY) or a KPC-mouse model tumor-derived primary cell line (R6560B) before intra-splenic injection with PAN02 cells. Control tumor-bearing mice educated with PBS (TU); n = 4 (TU and PKCYexo + TU) and n = 3 (R6560Bexo + TU) mice from one experiment. ∗∗P < 0.01,∗P < 0.05, N.S. stands for not significant by ANOVA. Scale bar, 1 cm. (c) Flow cytometric quantification of the frequency of liver and lung cells incorporating PKH67-labeled exosomes. Exosomes were isolated from normal pancreas (NP), PAN02, PKCY, and R6560B cells; n = 4 (NP) and n = 5 (R6560B) mice from one experiment, and n = 7 (PAN02) and n = 8 (PKCY) mice pooled from two experiments. Statistical source data can be found in Supplementary Table 4. ∗∗P < 0.01 by ANOVA. (d) Percentage of PKH67-labeled exosome+ liver cells expressing CD11b and F480 markers (KCs). Exosomes were isolated from human (BxPC-3 and HPAF-II) and murine (R6560B) PDAC cell lines; n = 4 (BxPC-3 and HPAF-II) and n = 5 (R6560B) mice from one experiment. (e) Fluorescence microscopy analysis of PKH67-labeled exosomes (green) and αSMA+ cells, S100A4+ fibroblasts, CD31+ endothelial cells, or EpCAM+ epithelial cells (red). Arrows point to green fluorescent signal indicating exosome uptake. Scale bars, 50 μm. (f) Analysis of canonical pathways of genes upregulated by Kupffer cells following in vitro education with BxPC-3 exosomes or PBS treatment. The list is comprised of genes related to liver fibrosis. Data was obtained from one experiment performed in triplicate. Statistical source data can be found in Supplementary Table 2. All data are represented as mean ± s.e.m.
Supplementary Figure 2 Pancreatic ductal adenocarcinoma-derived exosomes induce αSMA and FN expression and increase F4/80+ cell frequency in the liver and migration of BM-derived macrophages to the liver.
(a) Analysis by fluorescence microscopy showing lack of co-localization of FN expression with CD31+ endothelial cells in the livers of mice educated with PAN02 exosomes. Scale bar, 50 μm. (b) Immunofluorescence quantification of αSMA and FN expression in arbitrary units (a.u.) in the livers of mice educated with PBS (CTL), normal pancreas (NP), PKCY, or R6560B exosomes; n = 4 (CTLαSMA; PKCY αSMA; R6560B αSMA and FN) and n = 5 (CTL FN; NP αSMA and FN; PKCY FN) mice from one experiment. Scale bars, 150 μm.∗∗∗P < 0.001; N.S. stands for not significant by ANOVA. (c) Immunofluorescence quantification of the frequency of F4/80+ cell recruited to the livers of mice educated with PBS (CTL), normal pancreas (NP), PKYC, or R6560B exosomes; n = 4 (NP and R6560B) and n = 5 (CTL and PKCY) mice from one experiment. Scale bars, 150 μm.∗∗P < 0.01; N.S. stands for not significant by ANOVA. All data are represented as mean ± s.e.m.
Supplementary Figure 4 Metastatic pancreatic cells localize adjacent to liver macrophages.
Fluorescence microscopy analysis of F4/80+ staining and mCherry+ PAN02 cell localization in early metastatic lesions of mice educated with PAN02 exosomes reveals close association between metastatic cells and liver macrophages. Scale bar, 150 μm.
Supplementary Figure 5 Evaluation of MIF knockdown exosome uptake by KCs in vivo and the functional consequences on hStC activation and early metastasis.
(a) Western blot analysis of MIF levels in cells and in exosomes (Exo) of non-infected parental PAN02 cells (WT), PAN02 cells infected with control shRNA (shCTL), MIF knockdown (shMIF), a second construct of MIF knockdown (shMIF(2)), and R6560B cells. Unprocessed scans of blots are presented in in Supplementary Fig. 6. (b) Left panel: flow cytometric quantification of PKH67-labeled exosome incorporation by CD11b+F4/80+ liver cells (KCs), represented as a percentage of all PKH67+ cells; n = 4 (WT and shMIF) and n = 5 (shCTL and shMIF(2)) mice from one experiment. Right panel: exosome protein quantification, represented as microgram (μg) per 106 exosomes; n = 3 (shCTL, shMIF, and shMIF(2)) and n = 4 (WT) independent exosome isolations from in vitro cell culture. N.S. stands for not significant by ANOVA. (c) Size distribution analysis of PAN02 exosomes by NanoSight. (d) Immunofluorescence quantification of mCherry+ PAN02 cells 24 hours after their intra-splenic injection into mice previously educated for three weeks with parental PAN02 exosomes (Exo) or PAN02shMIF exosomes (shMIF exo). Arrows In representative images indicate PAN02 cells. Control animals were educated with PBS (TU); n = 4 mice per cohort from one experiment. ∗∗P < 0.01 by ANOVA. Scale bars, 100 μm. All data are represented as mean ± s.e.m.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2049 kb)
Supplementary Table 1
Supplementary Information (XLSX 242 kb)
Supplementary Table 2
Supplementary Information (XLSX 140 kb)
Supplementary Table 3
Supplementary Information (XLSX 86 kb)
Supplementary Table 4
Supplementary Information (XLSX 44 kb)
Rights and permissions
About this article
Cite this article
Costa-Silva, B., Aiello, N., Ocean, A. et al. Pancreatic cancer exosomes initiate pre-metastatic niche formation in the liver. Nat Cell Biol 17, 816–826 (2015). https://doi.org/10.1038/ncb3169
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3169
This article is cited by
-
Tumor-derived small extracellular vesicles in cancer invasion and metastasis: molecular mechanisms, and clinical significance
Molecular Cancer (2024)
-
Exosome-derived tRNA fragments tRF-GluCTC-0005 promotes pancreatic cancer liver metastasis by activating hepatic stellate cells
Cell Death & Disease (2024)
-
Regulated secretion of mutant p53 negatively affects T lymphocytes in the tumor microenvironment
Oncogene (2024)
-
HDAC1/2 control mesothelium/ovarian cancer adhesive interactions impacting on Talin-1-α5β1-integrin-mediated actin cytoskeleton and extracellular matrix protein remodeling
Journal of Experimental & Clinical Cancer Research (2024)
-
Extracellular vesicles related gene HSPH1 exerts anti-tumor effects in prostate cancer via promoting the stress response of CD8 + T cells
Cellular Oncology (2024)