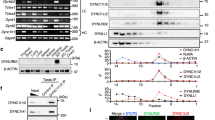
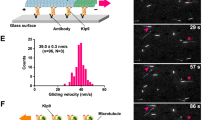
Abstract
Checkpoint controls ensure the completion of cell cycle events with high fidelity in the correct order. Here we show the existence of a novel checkpoint that ensures coupling of cell wall synthesis and mitosis. In response to a defect in cell wall synthesis, S. cerevisiae cells arrest the cell-cycle before spindle pole body separation. This arrest results from the regulation of the M-phase cyclin Clb2p at the transcriptional level through the transcription factor Fkh2p. Components of the dynactin complex are required to achieve the G2 arrest whilst keeping cells highly viable. Thus, the dynactin complex has a function in a checkpoint that monitors cell wall synthesis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Lew, D.J., Weinert, T. & Pringle, J.R. in The Molecular and Cellular Biology of the Yeast Saccharomyces (eds Pringle, J.R., Broach, J. & Jones, E.) 607–695 (Cold Spring Harbor Laboratory Press, New York, 1997).
Elledge, S.J. Cell cycle checkpoints: preventing an identity crisis. Science 274, 1664–1672 (1996).
Muhua, L., Adames, N.R., Murphy, M.D., Shields, C.R. & Cooper, J.A. A cytokinesis checkpoint requiring the yeast homologue of an APC-binding protein. Nature 393, 487–491 (1998).
Lew, D.J. & Reed, S.I. A cell cycle checkpoint monitors cell morphogenesis in budding yeast. J. Cell Biol. 129, 739–749 (1995).
McMillan, J.N., Sia, R.A.L. & Lew, D.J. A morphogenesis checkpoint monitors the actin cytoskeleton in yeast. J. Cell Biol. 142, 1487–1499 (1998).
Harvey, S.L. & Kellogg, D.R. Conservation of mechanisms controlling entry into mitosis: budding yeast wee1 delays entry into mitosis and is required for cell size control. Curr. Biol. 13, 264–275 (2003).
Orlean, P. in The Molecular and Cellular Biology of the Yeast Saccharomyces (eds Pringle, J.R., Broach, J. & Jones, E.) 229–362 (Cold Spring Harbor Laboratory Press, New York, 1997).
Cid, V.J. et al. Molecular basis of cell integrity and morphogenesis in Saccharomyces cerevisiae. Microbiol. Rev. 59, 345–386 (1995).
Cabib, E., Roberts, R. & Bowers, B. Synthesis of the yeast cell wall and its regulation. Annu. Rev. Biochem. 51, 763–793 (1982).
Lim, H.H., Goh, P.Y. & Surana, U. Spindle pole body separation in Saccharomyces cerevisiae requires dephosphorylation of the tyrosine 19 residue of Cdc28. Mol. Cell. Biol. 16, 6385–6397 (1996).
Douglas, C.M. et al. The Saccharomyces cerevisiae FKS 1(ETG1) gene encodes an integral membrane protein which is a subunit of 1,3-β-D-glucan synthase. Proc. Natl Acad. Sci. USA 91, 12907–12911 (1994).
Mazur, P. et al. Differential expression and function of two homologous subunits of yeast 1,3-β-D-glucan synthase. Mol. Cell. Biol. 15, 5671–5681 (1995).
Donaldson, A.D. & Kilmartin, J.V. Spc42p: a phosphorylated component of the S. cerevisiae spindle pole body (SPB) with an essential function during SPB duplication. J. Cell Biol. 132, 887–901 (1996).
Jacob, C.W., Adams, A.E.M., Szaniszlo, P.J. & Pringle, J.R. Functions of microtubules in the Saccharomyces cerevisiae cell cycle. J. Cell Biol. 107, 1409–1426 (1988).
Fitch, I. et al. Characterization of four B-type cyclin genes of the budding yeast Saccharomyces cerevisiae. Mol. Biol. Cell 3, 805–818 (1992).
Richardson, H., Lew, D.J., Henze, M., Sugimoto, K. & Reed, S.I. Cyclin-B homologs in Saccharomyces cerevisiae function in S phase and in G2 . Genes Dev. 6, 2021–2034 (1992).
Amon, A., Tyers, M., Futcher, B. & Nasmyth, K. Mechanisms that help the yeast cell cycle clock tick: G2 cyclins transcriptionally activate G2 cyclins and repress G1 cyclins. Cell 74, 993–1007 (1993).
Breeden, L.L. Cyclin transcription: Timing is everything. Curr. Biol. 10, R586–R588 (2000).
Muhua, L., Karpova, T.S. & Cooper, J.A. A yeast actin-related protein homologous to that in vertebrate dynactin complex is important for spindle orientation and nuclear migration. Cell 78, 669–679 (1994).
McMillan, J.N. & Tatchell, K. The JNM1 gene in the yeast Saccharomyces cerevisiae is required for nuclear migration and spindle orientation during the mitotic cell cycle. J. Cell Biol. 125, 143–158 (1994).
Geiser, J.R. et al. Saccharomyces cerevisiae genes required in the absence of the CIN8-encoded spindle motor act in functionally diverse mitotic pathways. Mol. Biol. Cell 8, 1035–1050 (1997).
Kahana, J.A. et al. The yeast dynactin complex is involved in partitioning the mitotic spindle between mother and daughter cells during anaphase B. Mol. Biol. Cell 9, 1741–1756 (1998).
Uetz, P. et al. A comprehensive analysis of protein–protein interactions in Saccharomyces cerevisiae. Nature 403, 623–627 (2000).
Ito, T. et al. A comprehensive two-hybrid analysis to explore the yeast protein interactome. Proc. Natl Acad. Sci. USA 98, 4569–4574 (2001).
Farkasovsky, M. & Kuntzel, H. Cortical Num1p interacts with the dynein intermediate chain Pac11p and cytoplasmic microtubules in budding yeast. J Cell Biol. 152, 251–262 (2001).
Lee, W.L., Oberle, J.R. & Cooper, J.A. The role of the lissencephaly protein Pac1 during nuclear migration in budding yeast. J. Cell Biol. 160, 335–364 (2003).
Sawistowska-Schroder, E.T., Kerridge, D. & Perry, H. Echinocandin inhibition of 1,3-β-D-glucan synthase from Candida albicans. FEBS Lett. 173, 134–138 (1984).
Mondesert, G. & Reed, S.I. BED1, a gene encoding a galactosyltransferase homologue, is required for polarized growth and efficient bud emergence in Saccharomyces cerevisiae. J. Cell Biol. 132, 137–151 (1996).
Gardner, R.D. & Burke, D.J. The spindle checkpoint: two transitions, two pathways. Trends Cell Biol. 10, 154–158 (2000).
Hoyt, M.A., Totis, L. & Roberts, B.T. S. cerevisiae genes required for cell cycle arrest in response to loss of microtubule function. Cell 66, 507–517 (1991).
Liu, J., Wang, H. & Balasubramanian, M.K. A checkpoint that monitors cytokinesis in Schizosaccharomyces pombe. J. Cell Sci. 113, 1223–1230 (2000).
Haase, S.B., Winey, M. & Reed, S.I. Multi-step control of spindle pole body duplication by cyclin-dependent kinase. Nature Cell Biol. 3, 38–42 (2001).
Surana, U. et al. The role of CDC28 and cyclins during mitosis in the budding yeast S. cerevisiae. Cell 65, 145–161 (1991).
Iyer, V.R. et al. Genomic binding sites of the yeast cell-cycle transcription factors SBF and MBF. Nature 409, 533–538 (2001).
Simon, I. et al. Serial regulation of transcriptional regulators in the yeast cell cycle. Cell 106, 697–708 (2001).
Gustin, M.C., Albertyn, J., Alexander, M. & Davenport, K. MAP kinase pathways in the yeast Saccharomyces cerevisiae. Microbiol. Mol. Biol. Rev. 62, 1264–1300 (1998).
Brown, J.L., North, S. & Bussey, H. SKN7, a yeast multicopy suppressor of a mutation affecting cell wall β-Glucan assembly, encodes a product with domains homologous to prokaryotic two-component regulators and to heat shock transcription factors. J. Bacteriol. 175, 6908–6915 (1993).
Harrison, J.C., Bardes, E.S., Ohya, Y. & Lew, D.J. A role for the Pkc1p/Mpk1p kinase cascade in the morphogenesis checkpoint. Nature Cell Biol. 3, 417–420 (2001).
Watanabe, Y., Takaesu, G., Hagiwara, M., Irie, K. & Matsumoto, K. Characterization of a serum response factor-like protein in Saccharomyces cerevisiae, Rlm1, which has transcriptional activity regulated by the Mpk1 (Slt2) mitogen-activated protein kinase pathway. Mol. Cell. Biol. 17, 2615–2623 (1997).
Madden, K., Sheu, Y.J., Baetz, K., Andrews, B. & Snyder, M. SBF cell cycle regulator as a target of the yeast PKC–MAP kinase pathway. Science 275, 1781–1784 (1997).
Adames, N.R. & Cooper, J.A. Microtubule interactions with the cell cortex causing nuclear movements in Saccharomyces cerevisiae. J. Cell Biol. 149, 863–874 (2000).
Vaughan, P.S., Miura, P., Henderson, M., Byrne, B. & Vaughan, K.T. A role for regulated binding of p150 (Glued) to microtubule plus ends in organelle transport. J. Cell Biol. 158, 305–319 (2002).
Sikorski, R.S. & Hieter, P. A system of shuttle vectors and yeast host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122, 19–27 (1989).
Schneider, B.L., Seufert, W., Steiner, B., Yang, Q.H. & Futcher, A.B. Use of polymerase chain reaction epitope tagging for protein tagging in Saccharomyces cerevisiae. Yeast 11, 1265–1274 (1995).
Ishihara, S., Hirata, A., Minemura, M., Nogami, S. & Ohya, Y. A mutation in SPC42, which encodes a component of the spindle pole body, results in production of two-spored asci in Saccharomyces cerevisiae. Mol. Genet. Genomics 265, 585–595 (2001).
Jones, J.S. & Prakash, L. Yeast Saccharomyces cerevisiae selectable markers in pUC18 polylinkers. Yeast 6, 363–366 (1990).
Sekiya-Kawasaki, M. et al. Dissection of upstream regulatory components of the Rho1p effector, 1,3-β-Glucan synthase, in Saccharomyces cerevisiae. Genetics 162, 663–676 (2002).
Rout, M.P. & Kilmartin, J.V. Components of the yeast spindle and spindle pole body. J. Cell Biol. 111, 1913–1927 (1990).
Ohtani, M., Sano, F., Saka, A., Ohya, Y. & Morishita, S. Development of image processing program for yeast cell morphology. J. Bioinfo. Comput. Biol. 1, 695–709 (2004).
Sun, G.H., Hirata, A., Oya, Y. & Anraku, Y. Mutation in yeast calmodulin cause defects in spindle pole body functions and nuclear integrity. J. Cell Biol. 119, 1625–1639 (1992).
Acknowledgements
We thank K. Morishita and S. Oshima for the initial work in this study; A. Hirata for electron microscopic analysis; M. Nishizawa, M. Fujino and D. Hirata for plasmids; T. Watanabe for Echinocandin B; K. Homma for critical reading of the manuscript; and M. Imanari and K. Shimane for preparation of the manuscript. Thanks also go to the members of the Laboratory of Signal Transduction for helpful discussions. This work was supported by a grant for Scientific Research from the Ministry of Education, Science, Sports and Culture of Japan, and by the Institute for Bioinformatics and Research and Development, of the Japan Science and Technology Corporation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Suzuki, M., Igarashi, R., Sekiya, M. et al. Dynactin is involved in a checkpoint to monitor cell wall synthesis in Saccharomyces cerevisiae. Nat Cell Biol 6, 861–871 (2004). https://doi.org/10.1038/ncb1162
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb1162
This article is cited by
-
Gene overexpression screen for chromosome instability in yeast primarily identifies cell cycle progression genes
Current Genetics (2019)
-
Conserved rules govern genetic interaction degree across species
Genome Biology (2012)
-
The ‘interactome’ of the Knr4/Smi1, a protein implicated in coordinating cell wall synthesis with bud emergence in Saccharomyces cerevisiae
Molecular Genetics and Genomics (2006)
-
A new checkpoint takes shape
Nature Cell Biology (2004)