Abstract
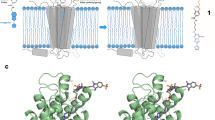
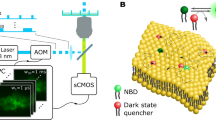
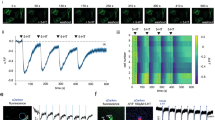
Chemical and biological labeling is fundamental for the elucidation of the function of proteins within biochemical cellular networks. In particular, fluorescent probes allow detection of molecular interactions, mobility and conformational changes of proteins in live cells with high temporal and spatial resolution1,2,3. We present a generic method to label proteins in vivo selectively, rapidly (seconds) and reversibly, with small molecular probes that can have a wide variety of properties. These probes comprise a chromophore and a metal-ion-chelating nitrilotriacetate (NTA) moiety, which binds reversibly and specifically to engineered oligohistidine sequences in proteins of interest4. We demonstrate the feasibility of the approach by binding NTA-chromophore conjugates5 to a representative ligand-gated ion channel and G protein–coupled receptor, each containing a polyhistidine sequence. We investigated the ionotropic 5HT3 serotonin receptor by fluorescence measurements to characterize in vivo the probe-receptor interactions, yielding information on structure and plasma membrane distribution of the receptor.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cha, A., Snyder, G.E., Selvin, P.R. & Bezanilla, F. Atomic scale movement of the voltage-sensing region in a potassium channel measured via spectroscopy. Nature 402, 809–813 (1999).
Schwille, P., Haupts, U., Maiti, S. & Webb, W.W. Molecular dynamics in living cells observed by fluorescence correlation spectroscopy with one- and two-photon excitation. Biophys. J. 77, 2251–2265 (1999).
Chamberlain, C. & Hahn, K.M. Watching proteins in the wild: fluorescence methods to study protein dynamics in living cells. Traffic 1, 755–762 (2000).
Hochuli, E., Dobeli, H. & Schacher, A. New metal chelate absorbant selective for proteins and peptides containing neighbouring histidine residues. J. Chromatography 411, 177–184 (1987).
Stora, T., Hovius, R., Dienes, Z., Pachoud, M. & Vogel, H. Metal ion trace detection by a chelator-modified gold electrode: a comparison of surface to bulk affinity. Langmuir 13, 5211–5214 (1997).
Holmes, K.L. & Lantz, L.M. Protein labeling with fluorescent probes. Methods Cell Biol. 63, 185–204 (2001).
Tsien, R.Y. The green fluorescent protein. Annu. Rev. Biochem. 67, 509–544 (1998).
Keppler, A. et al. A general method for the covalent labeling of fusion proteins with small molecules in vivo. Nat. Biotechnol. 21, 86–89 (2003).
Mendel, D., Cornish, V.W. & Schultz, P.G. Site-directed mutagenesis with an expanded genetic code. Annu. Rev. Biophys. Biomol. Struct. 24, 435–462 (1995).
Ilegems, E., Pick, H.M. & Vogel, H. Monitoring mis-acylated tRNA suppression efficiency in mammalian cells via EGFP fluorescence recovery. Nucleic Acids Res. 30, e128 (2002).
Griffin, B.A., Adams, S.R. & Tsien, R.Y. Specific covalent labeling of recombinant protein molecules inside live cells. Science 281, 269–272 (1998).
Giriat, I. & Muir, T.W. Protein semi-synthesis in living cells. J. Am. Chem. Soc. 215, 7180–7181 (2003).
McMahan, S.A. & Burgess, R.R. Single-step synthesis and characterization of biotinylated nitrilotriaceticacid, a unique reagent for the detection of histidine-tagged proteins immobilized on nitrocellulose. Anal. Biochem. 236, 101–106 (1996).
Kapanidis, A.N., Ebright, Y.W. & Ebright, R.H. Site-specific incorporation of fluorescent probes into protein: hexahistidine-tag-mediated fluorescent labeling with (Ni(2+):nitrilotriacetic acid (n)-fluorochrome conjugates. J. Am. Chem. Soc. 123, 12123–12125 (2001).
Keller, T.A. et al. Reversible oriented immobilization of histidine-tagged proteins on gold surfaces using a chelator thioalkane. Supramolecular Science 2, 155–160 (1995).
Reeves, D.C. & Lummis, S.C. The molecular basis of the structure and function of the 5-HT3 receptor: a model ligand-gated ion channel. Mol. Membr. Biol. 19, 11–26 (2002).
Tairi, A.P. et al. Ligand binding to the serotonin 5HT3 receptor studied with a novel fluorescent ligand. Biochemistry 37, 15850–15864 (1998).
Schreiter, C. et al. Characterization of the ligand-binding site of the serotonin 5-HT3 receptor: the role of glutamate residues 97, 224, and 235. J. Biol. Chem. 278, 22709–22716 (2003).
White, S.H. & Wimley, W.C. Membrane protein folding and stability: physical principles. Annu. Rev. Biophys. Biomol. Struct. 28, 319–365 (1999).
Maricq, A.V., Peterson, A.S., Brake, A.J., Myers, R.M. & Julius, D. Primary structure and functional expression of the 5HT3 receptor, a serotonin-gated ion channel. Science 254, 432–437 (1991).
Sprong, H., Van der Sluijs, P. & Van Meer, G. How proteins move lipids and lipids move proteins. Nat. Rev. Mol. Cell Biol. 2, 504–513 (2001).
Miyazawa, A., Fujiyoshi, Y. & Unwin, N. Structure and gating mechanism of the acetylcholine receptor pore. Nature 423, 949–955 (2003).
Jares-Erijman, E.A. & Jovin, T.M. FRET imaging. Nat. Biotech. 21, 1387–1395 (2003).
Maggi, C.A. & Schwartz, T.W. The dual nature of the tachykinin NK1 receptor. Trends Pharmacol. Sci. 18, 351–355 (1997).
Pierce, K.L., Premont, R.T. & Lefkowitz, R.J. Seven-transmembrane receptors. Nat. Rev. Mol. Cell. Biol. 3, 639–650 (2002).
Pick, H. et al. Monitoring expression and clustering of the ionotropic 5HT3 receptor in plasma membranes of live biological cell. Biochemistry 42, 877–884 (2003).
Van der Meer, B.W., Coker, G. & Chen, S.Y.S. Resonance energy transfer: theory and data (VCH Publishers, New York, 1994).
Boess, F.G., Beroukhim, R. & Martin, I.L. Ultrastructure of the 5-hydroxytryptamine3 receptor. J. Neurochem. 64, 1401–1405 (1995).
Wohland, T., Friedrich, K., Hovius, R. & Vogel, H. Study of ligand-receptor interactions by fluorescence correlation spectroscopy with different fluorophores: evidence that the homopentameric 5-hydroxytryptamine type 3As receptor binds only one ligand. Biochemistry 38, 8671–8681 (1999).
Vallotton, P. et al. Mapping the antagonist binding site of the serotonin type 3 receptor by fluorescence resonance energy transfer. Biochemistry 40, 12237–12242 (2001).
Acknowledgements
We are grateful to several colleagues: Horst Blasey for the stable cell line expressing 5HT3R-C-His6; Bruno Meyer for construction and functional expression of GFP-NK1-His10; David Lovinger, Horst Pick, Erwin Ilegems, Cédric Deluz and Kenneth Lundstrom for plasmids encoding 5HT3R, 5HT3R-Loop-His6, 5HT3R-N-His6, 5HT3R-C-His10 and NK1R; Andreas Heusler for synthesizing NTA-Rho; Annmarie Surprenant and Christoph Schreiter for the electrophysiological characterization of the oligohistidine-tagged 5HT3Rs; Martha Liley and Daniel Abankwa for carefully reading the manuscript. This work was financially supported by the Swiss National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Guignet, E., Hovius, R. & Vogel, H. Reversible site-selective labeling of membrane proteins in live cells. Nat Biotechnol 22, 440–444 (2004). https://doi.org/10.1038/nbt954
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt954
This article is cited by
-
Maturation strategy influences expression levels and cofactor occupancy in Fe–S proteins
JBIC Journal of Biological Inorganic Chemistry (2022)
-
A comprehensive comparison between camelid nanobodies and single chain variable fragments
Biomarker Research (2021)
-
Ultrafast in-gel detection by fluorescent super-chelator probes with HisQuick-PAGE
Communications Biology (2020)
-
Protein recognition by a pattern-generating fluorescent molecular probe
Nature Nanotechnology (2017)
-
Chemical labelling for visualizing native AMPA receptors in live neurons
Nature Communications (2017)