Abstract
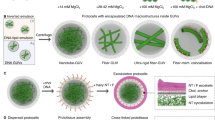
Two complementary strategies can be used in the fabrication of molecular biomaterials. In the 'top-down' approach, biomaterials are generated by stripping down a complex entity into its component parts (for example, paring a virus particle down to its capsid to form a viral cage). This contrasts with the 'bottom-up' approach, in which materials are assembled molecule by molecule (and in some cases even atom by atom) to produce novel supramolecular architectures. The latter approach is likely to become an integral part of nanomaterials manufacture and requires a deep understanding of individual molecular building blocks and their structures, assembly properties and dynamic behaviors. Two key elements in molecular fabrication are chemical complementarity and structural compatibility, both of which confer the weak and noncovalent interactions that bind building blocks together during self-assembly. Using natural processes as a guide, substantial advances have been achieved at the interface of nanomaterials and biology, including the fabrication of nanofiber materials for three-dimensional cell culture and tissue engineering, the assembly of peptide or protein nanotubes and helical ribbons, the creation of living microlenses, the synthesis of metal nanowires on DNA templates, the fabrication of peptide, protein and lipid scaffolds, the assembly of electronic materials by bacterial phage selection, and the use of radiofrequency to regulate molecular behaviors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Branden, C.-I. & Tooze, J. Introduction to Protein Structure edn. 2 (Garland, New York, 1999).
Gross, M. Travels to the Nanoworld: Miniature Machinery in Nature and Technology (Plenum, New York, 1999).
Benyus, J.M. Biomimicry: Innovation Inspired by Nature (Quill–William Morrow, New York, 1997).
Sarikaya, M., Tamerler, C., Jen, A.K.-Y., Schulten, K., & Baneyx, F. Molecular biomimetics: nanotechnology through biology. Nat. Materials 2, 577–585 (2003).
Lehn, J.-M. Supramolecular Chemistry: Concepts and Perspectives (John Wiley, New York, 1995).
Roukes, M.L. et al. Understanding Nanotechnology (Warner Books, New York, 2002).
Ratner, M. & Ratner, D. Nanotechnology: A Gentle Introduction to the Next Big Idea (Prentice Hall, Upper Saddle River, New Jersey, USA, 2003)
Freitas, R. Nanomedicine, Volume I: Basic Capabilities (Landes Bioscience, Austin, Texas, USA, 1999).
Zhang, S. Molecular self-assembly. in The Encyclopedia of Materials: Science & Technology (eds. Buschon, K.H. et al.) pp. 5822–5829 (Elsevier Science, Oxford, 2001).
Zhang, S., Marini, D., Hwang, W. & Santoso, S. Designing nanobiological materials through self-assembly of peptide & proteins. Curr. Opin. Chem. Biol. 6, 865–871 (2002).
Zhang, S. Building from bottom-up. Mater. Today 6, 20–27 (2003).
Gajdusek, D.C. Unconventional viruses and the origin and disappearance of kuru. Science 197, 943–960 (1977).
Fandrich, M., Fletcher, M.A. & Dobson, C.M. Amyloid fibrils from muscle myoglobin. Nature 410, 165–166 (2001).
Scheibel, T., Kowal, A.S., Bloom, J.D. & Lindquist, S.L. Bidirectional amyloid fiber growth for a yeast prion determinant. Curr. Biol. 11, 366–369 (2001).
Lynn, D.G. & Meredith, S.C. Review: model peptides and the physicochemical approach to β-amyloids. J. Struct. Biol. 130, 153–173 (2000).
West, M.W., Wang, W., Mancias, J.D., Beasley, J.R. & Hecht, M.H. De novo amyloid proteins from designed combinatorial libraries. Proc. Natl. Acad. Sci. USA 96, 11211–11216 (1999).
Hammarstrom, P., Schneider, F. & Kelly, J.W. Trans-suppression of misfolding in an amyloid disease. Science 293, 2459–2462 (2001).
Shtilerman, M.D., Ding, T.T. & Lansbury, P.T. Jr. Molecular crowding accelerates fibrillization of α-synuclein: could an increase in the cytoplasmic protein concentration induce Parkinson's disease? Biochemistry 41, 3855–3860 (2002).
Reches, M., Porat, Y. & Gazit, E. Amyloid fibril formation by pentapeptide and tetrapeptide fragments of human calcitonin. J. Biol. Chem. 277, 35475–35480 (2002).
Perutz, M.F., Finch, J.T., Berriman, J. & Lesk, A. Amyloid fibers are water-filled nanotubes. Proc. Natl. Acad. Sci. USA 99, 5591–5595 (2002).
Jimenez, J.L., Tennent, G., Pepys, M. & Saibil, H.R. Structural diversity of ex vivo amyloid fibrils studied by cryo-electron microscopy. J. Mol. Biol. 311, 241–247 (2001).
Lindquist, S.L. & Henikoff, S. Self-perpetuating structural states in biology, disease, and genetics. Proc. Natl. Acad. Sci. USA 99 (Suppl. 4), 16377 (2002).
Dobson, C.M. Protein folding and its links with human disease. Biochem. Soc. Symp. 68, 1–26 (2001).
Hwang, W., Marini, D., Kamm, R. & Zhang, S. Supramolecular structure of helical ribbons self-assembled from a β-sheet peptide. J. Chem. Physics 118, 389–397 (2003).
Ryadnov, M.G. & Woolfson, D.N. Engineering the morphology of a self-assembling protein fibre. Nat. Mater. 2, 329–332 (2003).
Lee, S. & Eisenberg, D. Seeded conversion of recombinant prion protein to a disulfide-bonded oligomer by a reduction-oxidation process. Nat. Struct. Biol. 10, 725–730 (2003).
Chiti, F., Stefani, M., Taddei, N., Ramponi, G. & Dobson, C.M. Rationalization of the effects of mutations on peptide and protein aggregation rates. Nature 424, 805–808 (2003).
Israelachvili, J.N., Mitchell, D.J. & Ninham, B.W. J. Chem. Soc. Faraday Trans. II 72, 1525–1568 (1976).
Singh, A., Price, R., Schoen, P.E., Yager, P. & Schnur, J.M. Tubule formation by hetero bifunctional polymerizable lipids: synthesis and characterization. Polymer Preprints 27, 393–394 (1986).
Schnur, J.M. et al. Lipid based tubule microstructures. Thin Solid Films 152, 181–206 (1987).
Schnur, J.M. Lipid tubules: a paradigm for molecular engineered structures. Science 262, 1669–1676 (1993).
Rudolph, A.S., Calvert, J.M., Schoen, P.E. & Schnur, J.M. Technological development of lipid based tubule microstructures. Adv. Exp. Med. Biol. 238, 305–320 (1988).
Vauthey, S. Santoso, S., Gong, H., Watson, N. & Zhang, S. Molecular self-assembly of surfactant-like peptides to form nanotubes and nanovesicles. Proc. Natl. Acad. Sci. USA 99, 5355–5360 (2002).
Santoso, S., Hwang, W., Hartman, H. & Zhang, S. Self-assembly of surfactant-like peptides with variable glycine tails to form nanotubes and nanovesicles. Nano Lett. 2, 687–691 (2002).
von Maltzahn, G., Vauthey, S., Santoso, S. & Zhang, S. Positively charged surfactant-like peptides self-assemble into nanostructures. Langmuir 19, 4332–4337 (2003).
Perutz, M.F. Glutamine repeats and neurodegenerative diseases. Brain Res. Bull. 50, 467 (1999).
Perutz, M.F., Pope, B.J., Owen, D., Wanker, E.E. & Scherzinger, E. Aggregation of proteins with expanded glutamine and alanine repeats of the glutamine-rich and asparagine-rich domains of Sup35 and of the amyloid β-peptide of amyloid plaques. Proc. Natl. Acad. Sci. USA 99, 5596–5600 (2002).
Perutz, M.F. & Windle, A.H. Cause of neural death in neurodegenerative diseases attributable to expansion of glutamine repeats. Nature 412, 143–144 (2001).
Whitesides, G.M., Mathias, J.P. & Seto, C.T. Molecular self-assembly and nanochemistry: a chemical strategy for the synthesis of nanostructures. Science 254, 1312–1319 (1991).
Mrksich, M. & Whitesides, G.M. Using self-assembled monolayers to understand the interactions of man-made surfaces with proteins and cells. Annu. Rev. Biophys. Biomol. Struct. 25, 55–78 (1996).
Chen, C.S., Mrksich, M., Huang, S., Whitesides, G.M. & Ingber, D.E. Geometric control of cell life and death. Science 276, 1425–1428 (1997).
Piner, R.D., Zhu, J., Xu, F., Hong, S., & Mirkin, C.A. Dip-pen nanolithography. Science 283, 661–664 (1999).
Lee, K-B., Park, S-J., Mirkin, C.A., Smith, J.C., & Mrksich, M. Protein nanoarrays generated by dip-pen nanolithography. Science 295, 1702–1705 (2002).
Demers, L.M. et al. Direct patterning of modified oligonucleotides on metals andinsulators by dip-pen nanolithography. Science 296, 1836–1838 (2002).
Dillo, A.K., Ochsenhirt, S.E., McCarthy, J.B., Fields, G.B. & Tirrell, M. Adhesion of α5β1 receptors to biomimetic substrates constructed from peptide amphiphiles. Biomaterials 22, 1493–1505 (2001).
Leufgen, K., Mutter, M., Vogel, H. & Szymczak, W. Orientation modulation of a synthetic polypeptide in self-assembled monolayers: a TOF-SIMS study. J. Am. Chem. Soc. 125, 8911–8915 (2003).
Zhang, S. et al. Biological surface engineering: a simple system for cell pattern formation. Biomaterials 20, 1213–1220 (1999).
Zhang, S., Holmes, T., Lockshin, C. & Rich, A. Spontaneous assembly of a self-complementary oligopeptide to form a stable macroscopic membrane. Proc. Natl. Acad. Sci. USA 90, 3334–3338 (1993).
Zhang, S. et al. Self-complementary oligopeptide matrices support mammalian cell attachment. Biomaterials 16, 1385–1393 (1995).
Holmes, T., Delacalle, S., Su, X., Rich, A. & Zhang, S. Extensive neurite outgrowth and active neuronal synapses on peptide scaffolds. Proc. Natl. Acad. Sci. USA 97, 6728–6733 (2000).
Caplan, M., Schwartzfarb, E., Zhang, S., Kamm, R. & Lauffenburger, D. Control of self-assembling oligopeptide matrix formation through systematic variation of amino acid sequence. Biomaterials 23, 219–227 (2002).
Marini, D., Hwang, W., Lauffenburger, D.A., Zhang, S. & Kamm, R.D. Left-handed helical ribbon intermediates in the self-assembly of a β-sheet peptide. Nano Lett. 2, 295–299 (2002).
Kisiday, J. et al. Self-assembling peptide hydrogel fosters chondrocyte extracellular matrix production and cell division: implications for cartilage tissue repair. Proc. Natl. Acad. Sci. USA 99, 9996–10001 (2002).
Petka, W.A., Harden, J.L., McGrath, K.P., Wirtz, D. & Tirrell, D.A. Reversible hydrogels from self-assembling artificial proteins. Science 281, 389–392 (1998).
Welsh, E.R. & Tirrell, D.A. Engineering the extracellular matrix: a novel approach to polymeric biomaterials. I. Control of the physical properties of artificial protein matrices designed to support adhesion of vascular endothelial cells. Biomacromolecules 1, 23–30 (2000).
Nowak, A.P. et al. Rapidly recovering hydrogel scaffolds from self-assembling diblock copolypeptide amphiphiles. Nature 417, 424–428 (2002).
Schneider, J.P. et al. Responsive hydrogels from the intramolecular folding and self-assembly of a designed peptide. J. Am. Chem. Soc. 124, 15030–15037 (2002).
Pochan, D. et al. Thermally reversible hydrogels via intramolecular folding and consequent self-assembly of a de novo designed peptide. J. Am. Chem. Soc. 125, in the press (2003).
Hartgerink, J.D., Beniash, E. & Stupp, S.I. Self-assembly and mineralization of peptide-amphiphile nanofibers. Science 294, 1684–1688 (2001).
Hartgerink, J.D., Beniash, E. & Stupp, S.I. Peptide-amphiphile nanofibers: a versatile scaffold for the preparation of self-assembling materials. Proc. Natl. Acad. Sci. USA 99, 5133–5138 (2002).
Semino, C.E., Merok, J.R., Crane, G., Panagiotakos, G. & Zhang, S. Functional differentiation of hepatocyte-like spheroid structures from putative liver progenitor cells in three-dimensional peptide scaffolds. Differentiation 71, 262–270 (2003).
Semino, C.E., Kasahara, J., Hayashi, Y. & Zhang, S. (2003). Entrapment of hippocampal neural cells in self-assembling peptide scaffold. Tissue Eng. in the press (2003).
Djalali, R., Chen, Y.F. & Matsui, H. Au nanowire fabrication from sequenced histidine-rich peptide. J. Am. Chem. Soc. 124, 13660–13661 (2002).
Matsui, H., Porrata, P. & Douberly, G.E. Protein tubule immobilization on self-assembled monolayers on Au substrates. Nano Lett. 1, 461–464 (2001).
Lvov, Y.M. et al. Imaging nanoscale patterns on biologically derived microstructures. Langmuir 16, 5932–5935 (2000).
Reches, M. & Gazit, E. Casting metal nanowires within discrete self-assembled peptide nanotubes. Science 300, 625–627 (2003).
Scheibel, T. et al. Conducting nanowires built by controlled self-assembly of amyloid fibers and selective metal deposition. Proc. Natl. Acad. Sci. USA 100, 4527–4532 (2003).
Whaley, S.R., English, D.S., Hu, E.L., Barbara, P.F. & Belcher, A.M. Selection of peptides with semiconductor binding specificity for directed nanocrystal assembly. Nature 405, 665–668 (2000).
Lee, S.W., Mao, C., Flynn, C.E. & Belcher, A.M. Ordering of quantum dots using genetically engineered viruses. Science 296, 892–895 (2002).
Mao, C. et al. Viral assembly of oriented quantum dot nanowires. Proc. Natl. Acad. Sci. USA 100, 6946–6951 (2003).
Aizenberg, J., Tkachenko, A., Weiner, S., Addadi, L. & Hendler, G. Calcitic microlenses as part of the photoreceptor system in brittlestars. Nature 412, 819–822 (2001).
Sundar, V.C., Yablon, A.D., Grazul, J.L., Ilan, M. & Aizenberg, J. Fibre-optical features of a glass sponge. Nature 424, 899–900 (2003).
Aizenberg, J., Muller, D.A., Grazul, J.L. & Hamann, D.R. Direct fabrication of large micropatterned single crystals. Science 299, 1205–1208 (2003).
Bach, K. & Burkhardt, B. (eds.). Diatoms I: Shells in Nature and Technics (Karl Kramer Verlag, Stuttgart, Germany, 1984).
The Ribosome. Cold Spring Harbor Symposium Quantitative Biology vol. LXVI (Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, USA, 2001).
Seeman, N.C. & Belcher, A.M. Emulating biology: building nanostructures from the bottom up. Proc. Natl. Acad. Sci. USA 99 (Suppl. 2), 6451–6455 (2002).
Furka, A. Combinatorial chemistry: 20 years on.... Drug Discov. Today 7, 1–4 (2002).
Blondelle, S.E., Takahashi, E., Houghten, R.A. & Perez-Paya, E. Rapid identification of compounds with enhanced antimicrobial activity by using conformationally defined combinatorial libraries. Biochem. J. 313, 141–147 (1996).
Moffet, D.A. & Hecht, M.H. De novo proteins from combinatorial libraries. Chem. Rev. 101, 3191–3203 (2001).
Feldhaus, M.J. et al. Flow-cytometric isolation of human antibodies from a nonimmune Saccharomyces cerevisiae surface display library. Nat. Biotechnol. 21, 163–170 (2003).
Leaky, R. & Lewin, R. The Sixth Extinction: Patterns of Life and the Future of Humankind (Doubleday, New York, 1995).
Hamad-Schifferli, K., Schwartz, J., Santos, A., Zhang, S. & Jacobson, J. Remote electronic control of DNA hybridization through inductive coupling to an attached metal nanocrystal antenna. Nature 415, 152–155 (2002).
Lindquist, S. Presentation at the Third Multidisciplinary Workshop: Self-assembly of Peptides, Proteins in Biology, Engineering and Medicine, Crete, Greece, August 1–5, 2003.
Selinger, J.V. & Schnur, J.M. Theory of chiral lipid tubules. Phys. Rev. Lett. 71, 4091–4094 (1993).
Aggeli, A. et al. Hierarchical self-assembly of chiral rod-like molecules as a model for peptide β-sheet tapes, ribbons, fibrils, and fibers. Proc. Natl. Acad. Sci. USA 98, 11857–11862 (2001).
Dobson, C.M. Protein misfolding, evolution and disease. Trends Biochem. Sci. 24329–24332 (1999).
Altman, M., Lee, P., Rich, A. & Zhang, S. Conformational behavior of ionic self-complementary peptides. Protein Sci. 9, 1095–1105 (2000).
Acknowledgements
I thank Hidenori Yokoi for helping to organize the figures and Steve Santoso for critically reading the manuscript. I would also like to thank members of my laboratory, past and present, for making discoveries and conducting exciting research. I gratefully acknowledge the support by grants from the US Army Research Office, Office of Naval Research, Defense Advanced Research Project Agency (BioComputing), DARPA/Naval Research Labs; NSF-MIT BPEC and NSF CCR-0122419 to the MIT Media Lab's Center for Bits & Atoms; the US National Institutes of Health; the Whitaker Foundation; the DuPont–MIT Alliance; and Menicon, Ltd., Japan. The author also acknowledges the Intel Corp. for its educational donation of a computing cluster to the Center for Biomedical Engineering at MIT.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Zhang, S. Fabrication of novel biomaterials through molecular self-assembly. Nat Biotechnol 21, 1171–1178 (2003). https://doi.org/10.1038/nbt874
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt874
This article is cited by
-
Biogenically synthesized nanoparticles in wastewater treatment; a greener approach: a review
Clean Technologies and Environmental Policy (2024)
-
Hierarchical Self-Assembly of Injectable Alginate Supramolecular Nanofibril Hydrogels for Hemostasis In Vivo
Advanced Fiber Materials (2024)
-
Amyloid-polysaccharide interfacial coacervates as therapeutic materials
Nature Communications (2023)
-
Self-assembled photonic cavities with atomic-scale confinement
Nature (2023)
-
Bioinspired chiral inorganic nanomaterials
Nature Reviews Bioengineering (2023)