Abstract
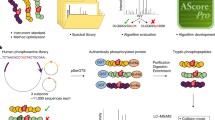
With the recent exponential increase in protein phosphorylation sites identified by mass spectrometry, a unique opportunity has arisen to understand the motifs surrounding such sites. Here we present an algorithm designed to extract motifs from large data sets of naturally occurring phosphorylation sites. The methodology relies on the intrinsic alignment of phospho-residues and the extraction of motifs through iterative comparison to a dynamic statistical background. Results show the identification of dozens of novel and known phosphorylation motifs from recently published serine, threonine and tyrosine phosphorylation studies. When applied to a linguistic data set to test the versatility of the approach, the algorithm successfully extracted hundreds of language motifs. This method, in addition to shedding light on the consensus sequences of identified and as yet unidentified kinases and modular protein domains, may also eventually be used as a tool to determine potential phosphorylation sites in proteins of interest.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
References
Schlessinger, J. & Lemmon, M.A. SH2 and PTB domains in tyrosine kinase signaling. Sci. STKE 2003, RE12 (2003).
Ang, X.L. & Wade Harper, J. SCF-mediated protein degradation and cell cycle control. Oncogene 24, 2860–2870 (2005).
Pawson, T. & Scott, J.D. Protein phosphorylation in signaling—50 years and counting. Trends Biochem. Sci. 30, 286–290 (2005).
Obenauer, J.C., Cantley, L.C. & Yaffe, M.B. Scansite 2.0: Proteome-wide prediction of cell signaling interactions using short sequence motifs. Nucleic Acids Res. 31, 3635–3641 (2003).
Manning, B.D. & Cantley, L.C. Hitting the target: emerging technologies in the search for kinase substrates. Sci. STKE 2002, PE49 (2002).
Rush, J. et al. Immunoaffinity profiling of tyrosine phosphorylation in cancer cells. Nat. Biotechnol. 23, 94–101 (2005).
Ficarro, S.B. et al. Phosphoproteome analysis by mass spectrometry and its application to Saccharomyces cerevisiae. Nat. Biotechnol. 20, 301–305 (2002).
Beausoleil, S.A. et al. Large-scale characterization of HeLa cell nuclear phosphoproteins. Proc. Natl. Acad. Sci. USA 101, 12130–12135 (2004).
Collins, M.O. et al. Proteomic analysis of in vivo phosphorylated synaptic proteins. J. Biol. Chem. 280, 5972–5982 (2005).
Ballif, B.A., Villen, J., Beausoleil, S.A., Schwartz, D. & Gygi, S.P. Phosphoproteomic analysis of the developing mouse brain. Mol. Cell. Proteomics 3, 1093–1101 (2004).
Gruhler, A. et al. Quantitative phosphoproteomics applied to the yeast pheromone signaling pathway. Mol. Cell. Proteomics 4, 310–327 (2005).
Nuhse, T.S., Stensballe, A., Jensen, O.N. & Peck, S.C. Phosphoproteomics of the Arabidopsis plasma membrane and a new phosphorylation site database. Plant Cell 16, 2394–2405 (2004).
Loyet, K.M., Stults, J.T. & Arnott, D. Mass spectrometric contributions to the practice of phosphorylation site mapping through 2003: a literature review. Mol. Cell. Proteomics 4, 235–245 (2005).
Bussemaker, H.J., Li, H. & Siggia, E.D. Building a dictionary for genomes: identification of presumptive regulatory sites by statistical analysis. Proc. Natl. Acad. Sci. USA 97, 10096–10100 (2000).
Melville, H. Moby-Dick, or, The whale (Signet Classic, New York, 1998).
Diella, F. et al. Phospho.ELM: a database of experimentally verified phosphorylation sites in eukaryotic proteins. BMC Bioinformatics 5, 79 (2004).
Rigoutsos, I. & Floratos, A. Combinatorial pattern discovery in biological sequences: The TEIRESIAS algorithm. Bioinformatics 14, 55–67 (1998).
Jonassen, I., Collins, J.F. & Higgins, D.G. Finding flexible patterns in unaligned protein sequences. Protein Sci. 4, 1587–1595 (1995).
Thompson, W., Rouchka, E.C. & Lawrence, C.E. Gibbs Recursive Sampler: finding transcription factor binding sites. Nucleic Acids Res. 31, 3580–3585 (2003).
Nevill-Manning, C.G., Wu, T.D. & Brutlag, D.L. Highly specific protein sequence motifs for genome analysis. Proc. Natl. Acad. Sci. USA 95, 5865–5871 (1998).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Crooks, G.E., Hon, G., Chandonia, J.M. & Brenner, S.E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004).
Boucher, L., Ouzounis, C.A., Enright, A.J. & Blencowe, B.J. A genome-wide survey of RS domain proteins. RNA 7, 1693–1701 (2001).
Fujimoto, J. et al. Characterization of the transforming activity of p80, a hyperphosphorylated protein in a Ki-1 lymphoma cell line with chromosomal translocation t(2;5). Proc. Natl. Acad. Sci. USA 93, 4181–4186 (1996).
Iuchi, S. Three classes of C2H2 zinc finger proteins. Cell. Mol. Life Sci. 58, 625–635 (2001).
Songyang, Z. & Cantley, L.C. Recognition and specificity in protein tyrosine kinase-mediated signalling. Trends Biochem. Sci. 20, 470–475 (1995).
Branch, D.R. & Mills, G.B. pp60c-src expression is induced by activation of normal human T lymphocytes. J. Immunol. 154, 3678–3685 (1995).
Shin, N.Y. et al. Subsets of the major tyrosine phosphorylation sites in Crk-associated substrate (CAS) are sufficient to promote cell migration. J. Biol. Chem. 279, 38331–38337 (2004).
Yates, J.R. III, Eng, J.K. & McCormack, A.L. Mining genomes: correlating tandem mass spectra of modified and unmodified peptides to sequences in nucleotide databases. Anal. Chem. 67, 3202–3210 (1995).
Acknowledgements
The authors thank John Rush and Cell Signaling Technology for providing access to the tyrosine phosphorylation data sets prior to their publication. Additionally, D.S. wishes to thank Michael Chou for assistance with the Moby Dick analysis as well as numerous stimulating conversations regarding the algorithm and critical reading of the manuscript. This work was supported in part by National Institutes of Health grant HG03456 (S.P.G.).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
About this article
Cite this article
Schwartz, D., Gygi, S. An iterative statistical approach to the identification of protein phosphorylation motifs from large-scale data sets. Nat Biotechnol 23, 1391–1398 (2005). https://doi.org/10.1038/nbt1146
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1146
This article is cited by
-
Broad phosphorylation mediated by testis-specific serine/threonine kinases contributes to spermiogenesis and male fertility
Nature Communications (2023)
-
Integrative proteogenomic characterization of early esophageal cancer
Nature Communications (2023)
-
Proteogenomic insights into the biology and treatment of pancreatic ductal adenocarcinoma
Journal of Hematology & Oncology (2022)
-
Linking post-translational modifications and protein turnover by site-resolved protein turnover profiling
Nature Communications (2022)
-
Integrated multi-omics analysis of adverse cardiac remodeling and metabolic inflexibility upon ErbB2 and ERRα deficiency
Communications Biology (2022)