Abstract
Mass-spectrometry-based phosphoproteomics has become indispensable for understanding cellular signaling in complex biological systems. Despite the central role of protein phosphorylation, the field still lacks inexpensive, regenerable, and diverse phosphopeptides with ground-truth phosphorylation positions. Here, we present Iterative Synthetically Phosphorylated Isomers (iSPI), a proteome-scale library of human-derived phosphoserine-containing phosphopeptides that is inexpensive, regenerable, and diverse, with precisely known positions of phosphorylation. We demonstrate possible uses of iSPI, including use as a phosphopeptide standard, a tool to evaluate and optimize phosphorylation-site localization algorithms, and a benchmark to compare performance across data analysis pipelines. We also present AScorePro, an updated version of the AScore algorithm specifically optimized for phosphorylation-site localization in higher energy fragmentation spectra, and the FLR viewer, a web tool for phosphorylation-site localization, to enable community use of the iSPI resource. iSPI and its associated data constitute a useful, multi-purpose resource for the phosphoproteomics community.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout


Similar content being viewed by others
Data Availability
The mass spectrometry data generated in this work have been deposited to the ProteomeXchange Consortium with the dataset identifier PXD031171 (Fig. 2, Extended Data Fig. 1 and Supplementary Figs. 2, 4, 5, and 7). Previously published datasets re-analyzed in this work (Supplementary Fig. 6) are also available through ProteomeXchange via their dataset identifiers listed in Supplementary Table 5. Publicly available databases used include EcoCyc v17 (https://biocyc.org/organism-summary?object=ECOLI) and the Uniprot Human Database (11/2018 release, https://ftp.uniprot.org/pub/databases/uniprot/previous_releases/release-2018_11/).
Code Availability
The AScorePro software package is available as a standalone application from Github at https://github.com/gygilab/MPToolKit. The FLR viewer for phosphorylation site localization is available as a web application at http://wren.hms.harvard.edu/iSPI/. The source code for the FLR viewer is available from Github at https://github.com/gygilab/iSPI_Viewer.
References
Hornbeck, P. V. et al. PhosphoSitePlus, 2014: mutations, PTMs and recalibrations. Nucleic Acids Res. 43, D512–D520 (2015).
Yu, K. et al. QPhos: a database of protein phosphorylation dynamics in humans. Nucleic Acids Res. 47, D451–D458 (2019).
Krug, K. et al. A curated resource for phosphosite-specific signature analysis. Mol. Cell. Proteom. 18, 576–593 (2019).
Ochoa, D. et al. The functional landscape of the human phosphoproteome. Nat. Biotechnol. 38, 365–373 (2020).
Kalyuzhnyy, A. et al. Profiling the human phosphoproteome to estimate the true extent of protein phosphorylation. J. Proteome Res. 21, 1510–1524 (2022).
Marx, H. et al. A large synthetic peptide and phosphopeptide reference library for mass spectrometry-based proteomics. Nat. Biotechnol. 31, 557–564 (2013).
Ferries, S. et al. Evaluation of parameters for confident phosphorylation site localization using an orbitrap fusion tribrid mass spectrometer. J. Proteome Res. 16, 3448–3459 (2017).
Cui, L. & Reid, G. E. Examining factors that influence erroneous phosphorylation site localization via competing fragmentation and rearrangement reactions during ion trap CID-MS/MS and -MS(3.). Proteomics 13, 964–973 (2013).
Wiese, H. et al. Comparison of alternative MS/MS and bioinformatics approaches for confident phosphorylation site localization. J. Proteome Res. 13, 1128–1137 (2014).
Suni, V. et al. SimPhospho: a software tool enabling confident phosphosite assignment. Bioinformatics 34, 2690–2692 (2018).
Sharma, K. et al. Ultradeep human phosphoproteome reveals a distinct regulatory nature of Tyr and Ser/Thr-based signaling. Cell Rep. 8, 1583–1594 (2014).
Ramsbottom, K. A. et al. Method for independent estimation of the false localization rate for phosphoproteomics. J. Proteome Res. 21, 1603–1615 (2022).
Jiang, W. et al. Deep-learning-derived evaluation metrics enable effective benchmarking of computational tools for phosphopeptide identification. Mol. Cell. Proteom. 20, 100171 (2021).
Pirman, N. L. et al. A flexible codon in genomically recoded Escherichia coli permits programmable protein phosphorylation. Nat. Commun. 6, 8130 (2015).
Mohler, K., Moen, J., Rogulina, S. & Rinehart, J. Principles for systematic optimization of an orthogonal translation system with enhanced biological tolerance. Preprint at bioRxiv https://doi.org/10.1101/2021.05.20.444985 (2021).
Barber, K. W. et al. Encoding human serine phosphopeptides in bacteria for proteome-wide identification of phosphorylation-dependent interactions. Nat. Biotechnol. 36, 638–644 (2018).
Schroeder, M. J., Shabanowitz, J., Schwartz, J. C., Hunt, D. F. & Coon, J. J. A neutral loss activation method for improved phosphopeptide sequence analysis by quadrupole ion trap mass spectrometry. Anal. Chem. 76, 3590–3598 (2004).
Beausoleil, S. A., Villén, J., Gerber, S. A., Rush, J. & Gygi, S. P. A probability-based approach for high-throughput protein phosphorylation analysis and site localization. Nat. Biotechnol. 24, 1285–1292 (2006).
Mintseris, J. & Gygi, S. P. High-density chemical cross-linking for modeling protein interactions. Proc. Natl Acad. Sci. USA 117, 93–102 (2020).
Pedrioli, P. G. A. et al. A common open representation of mass spectrometry data and its application to proteomics research. Nat. Biotechnol. 22, 1459–1466 (2004).
Martens, L. et al. mzML — a community standard for mass spectrometry data. Mol. Cell. Proteomics 10, R110.000133 (2011).
Taus, T. et al. Universal and confident phosphorylation site localization using phosphoRS. J. Proteome Res. 10, 5354–5362 (2011).
Hogrebe, A. et al. Benchmarking common quantification strategies for large-scale phosphoproteomics. Nat. Commun. 9, 1–13 (2018).
Potel, C. M., Lemeer, S. & Heck, A. J. R. Phosphopeptide fragmentation and site localization by mass spectrometry: an update. Anal. Chem. 91, 126–141 (2019).
Verheggen, K. et al. Anatomy and evolution of database search engines—a central component of mass spectrometry based proteomic workflows. Mass Spectrom. Rev. 39, 292–306 (2020).
Locard-Paulet, M., Bouyssié, D., Froment, C., Burlet-Schiltz, O. & Jensen, L. J. Comparing 22 popular phosphoproteomics pipelines for peptide identification and site localization. J. Proteome Res. 19, 1338–1345 (2020).
Eng, J. K., Jahan, T. A. & Hoopmann, M. R. Comet: an open-source MS/MS sequence database search tool. Proteomics 13, 22–24 (2013).
Cox, J. et al. Andromeda: a peptide search engine integrated into the MaxQuant environment. J. Proteome Res. 10, 1794–1805 (2011).
Tabb, D. L. The SEQUEST family tree. J. Am. Soc. Mass. Spectrom. 26, 1814–1819 (2015).
Dorfer, V. et al. MS Amanda, a universal identification algorithm optimized for high accuracy tandem mass spectra. J. Proteome Res. 13, 3679–3684 (2014).
Kong, A. T., Leprevost, F. V., Avtonomov, D. M., Mellacheruvu, D. & Nesvizhskii, A. I. MSFragger: ultrafast and comprehensive peptide identification in mass spectrometry-based proteomics. Nat. Methods 14, 513–520 (2017).
Yu, F. et al. Identification of modified peptides using localization-aware open search. Nat. Commun. 11, 1–9 (2020).
Geiszler, D. J. et al. PTM-shepherd: Analysis and summarization of post-translational and chemical modifications from open search results. Mol. Cell. Proteomics 20, 100018 (2021).
Amiram, M. et al. Evolution of translation machinery in recoded bacteria enables multi-site incorporation of nonstandard amino acids. Nat. Biotechnol. 33, 1272–1279 (2015).
Muehlbauer, L. K., Hebert, A. S., Westphall, M. S., Shishkova, E. & Coon, J. J. Global phosphoproteome analysis using high-field asymmetric waveform ion mobility spectrometry on a hybrid orbitrap mass spectrometer. Anal. Chem. 92, 15959–15967 (2020).
Rad, R. et al. Improved monoisotopic mass estimation for deeper proteome coverage. J. Proteome Res. 20, 591–598 (2021).
Keseler, I. M. et al. The EcoCyc database: reflecting new knowledge about Escherichia coli K-12. Nucleic Acids Res. 45, D543–D550 (2017).
Elias, J. E. & Gygi, S. P. Target-decoy search strategy for increased confidence in large-scale protein identifications by mass spectrometry. Nat. Methods 4, 207–214 (2007).
Huttlin, E. L. et al. A tissue-specific atlas of mouse protein phosphorylation and expression. Cell 143, 1174–1189 (2010).
Savitski, M. M., Wilhelm, M., Hahne, H., Kuster, B. & Bantscheff, M. A scalable approach for protein false discovery rate estimation in large proteomic data sets. Mol. Cell. Proteom. 14, 2394–2404 (2015).
Li, J. et al. TMTpro-18plex: the expanded and complete set of TMTpro reagents for sample multiplexing. J. Proteome Res. 20, 2964–2972 (2021).
Li, J. et al. TMTpro reagents: a set of isobaric labeling mass tags enables simultaneous proteome-wide measurements across 16 samples. Nat. Methods 17, 399–404 (2020).
Li, J., Paulo, J. A., Nusinow, D. P., Huttlin, E. L. & Gygi, S. P. Investigation of proteomic and phosphoproteomic responses to signaling network perturbations reveals functional pathway organizations in yeast. Cell Rep. 29, 2092–2104.e4 (2019).
Popow, O., Liu, X., Haigis, K. M., Gygi, S. P. & Paulo, J. A. A compendium of murine (phospho)peptides encompassing different isobaric labeling and data acquisition strategies. J. Proteome Res. 20, 3678–3688 (2021).
Acknowledgements
We would like to thank members of the Gygi Lab at Harvard Medical School for productive discussions, and S. Rogulina for technical assistance. This work was funded in part NIH/NIGMS grants GM132129 (J. A. P.), GM117230 (J. R.) and GM67945 (S. P. G). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
B. M. G., J.R. and S. P. G. conceived the study. J. R. and K. M. provided reagents. B. M. G. and J. A. P. performed experiments. B. M. G. and J. L. analyzed data. J. L. R. R. and J. M. conceived and provided computational tools. T. L., M. A. and S. A. B. provided advice on data analysis. E. L. H. and S. P. G. oversaw this work. B. M. G., J. L. and S. P. G. wrote the manuscript with input and editing from all authors.
Corresponding author
Ethics declarations
Competing interests
The iSPI library is covered under patent EP3755798A4 (pending, inventor: J. R., assignee: Yale University, Agilent Technologies Inc), and pSerOTS is covered under patent US7723069B2 (active, inventor: J. R., assignee: Yale University). The other authors declare no competing interests.
Peer review
Peer review information
Nature Methods thanks Tiannan Guo, Martin Larsen, and the other, anonymous, reviewer(s) for their contribution to the peer review of this work. Primary Handling Editor: Arunima Singh, in collaboration with the Nature Methods team. Peer reviewer reports are available.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 Using the iSPI to compare phosphoproteomics pipelines.
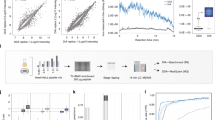
The datasets for the three subpools were analyzed using four different pipelines. Each pipeline included a database searching algorithm and a site localization tool. a) Similar to Fig. 2, receiver operating curves are shown displaying the number of combined sites from the three pools on the y-axis as a function of the FLR on the x-axis. Vertical grey dotted lines represent empirical FLR’s of 0.01 and 0.05 respectively. Labelled points represent commonly used localization cutoffs: 13 for AScore and AScorePro, 0.75 localization probability for MaxQuant, and 0.95 site probability for proteome discoverer. At an empirical FLR of 0.05, differences are apparent. b) The number of phosphorylation sites per pool is shown for label free peptides at an empirical 5% FLR. c) The number of phosphorylation sites per pool is shown for TMTpro-labeled peptides at an empirical 5% FLR. Bar represents the mean. Error bars represent the standard error of the mean for n = 3 subpools.
Supplementary information
Supplementary Information
Supplementary Notes 1–3 and Supplementary Figures 1–8
Supplementary Table 1
iSPI amino acid and oligonucleotide sequences
Supplementary Table 2
Combined identified and unique iSPI phosphopeptides
Supplementary Table 3
iSPi mass spectrometry file metadata
Supplementary Table 4
iSPI phosphosite identifications and FLR at various score cutoffs
Supplementary Table 5
Biological data re-analysis metadata
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Gassaway, B.M., Li, J., Rad, R. et al. A multi-purpose, regenerable, proteome-scale, human phosphoserine resource for phosphoproteomics. Nat Methods 19, 1371–1375 (2022). https://doi.org/10.1038/s41592-022-01638-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41592-022-01638-5
This article is cited by
-
Covalent inhibition of pro-apoptotic BAX
Nature Chemical Biology (2024)
-
Spatial probabilistic mapping of metabolite ensembles in mass spectrometry imaging
Nature Communications (2023)
-
DeepFLR facilitates false localization rate control in phosphoproteomics
Nature Communications (2023)