Abstract
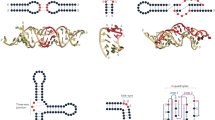
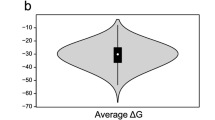
In contrast to the fairly reliable and complete annotation of the protein coding genes in the human genome, comparable information is lacking for noncoding RNAs (ncRNAs). We present a comparative screen of vertebrate genomes for structural noncoding RNAs, which evaluates conserved genomic DNA sequences for signatures of structural conservation of base-pairing patterns and exceptional thermodynamic stability. We predict more than 30,000 structured RNA elements in the human genome, almost 1,000 of which are conserved across all vertebrates. Roughly a third are found in introns of known genes, a sixth are potential regulatory elements in untranslated regions of protein-coding mRNAs and about half are located far away from any known gene. Only a small fraction of these sequences has been described previously. A comparison with recent tiling array data shows that more than 40% of the predicted structured RNAs overlap with experimentally detected sites of transcription. The widespread conservation of secondary structure points to a large number of functional ncRNAs and cis-acting mRNA structures in the human genome.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
The Human Genome Sequencing Consortium. Finishing the euchromatic sequence of the human genome. Nature 431, 931–945 (2004).
Bertone, P. et al. Global identification of human transcribed sequences with genome tiling arrays. Science 306, 2242–2246 (2004).
Kampa, D. et al. Novel RNAs identified from an in-depth analysis of the transcriptome of human chromosomes 21 and 22. Genome Res. 14, 331–342 (2004).
Johnson, J.M., Edwards, S., Shoemaker, D. & Schadt, E.E. Dark matter in the genome: evidence of widespread transcription detected by microarray tiling experiments. Trends Genet. 21, 93–102 (2005).
Cheng, J. et al. Transcriptional maps of 10 human chromosomes at 5-nucleotide resolution. Science 308, 1149–1154 (2005).
Okazaki, Y. et al. Analysis of the mouse transcriptome based on functional annotation of 60,770 full-length cDNAs. Nature 420, 563–573 (2002).
Imanishi, T. et al. Integrative annotation of 21,037 human genes validated by full-length cDNA clones. PLoS Biology 2, 0856–0875 (2004).
Cawley, S. et al. Unbiased mapping of transcription factor binding sites along human chromosomes 21 and 22 points to widespread regulation of noncoding RNAs. Cell 116, 499–509 (2004).
Hüttenhofer, A., Schattner, P. & Polacek, N. Non-coding RNAs: hope or hype? Trends Genet. 21, 289–297 (2005).
Hofacker, I.L. et al. Automatic detection of conserved RNA structure elements in complete RNA virus genomes. Nucleic Acids Res. 26, 3825–3836 (1998).
Rivas, E., Klein, R.J., Jones, T.A. & Eddy, S.R. Computational identification of noncoding RNAs in E. coli by comparative genomics. Curr. Biol. 11, 1369–1373 (2001).
Washietl, S., Hofacker, I.L. & Stadler, P.F. Fast and reliable prediction of noncoding RNAs. Proc. Natl. Acad. Sci. USA 102, 2454–2459 (2005).
Moulton, V. Tracking down noncoding RNAs. Proc. Natl. Acad. Sci. USA 102, 2269–2270 (2005).
Shabalina, S.A. & Kondrashov, A.S. Pattern of selective constraint in C. elegans and C. briggsae genomes. Genet. Res. 74, 23–30 (1999).
Shabalina, S.A., Ogurtsov, A.Y., Kondrashov, V.A. & Kondrashov, A.S. Selective constraint in intergenic regions of human and mouse genomes. Trends Genet. 17, 373–376 (2001).
Margulies, E.H., Blanchette, M., Haussler, D. & Green, E.D. Identification and characterization of multi-species conserved sequences. Genome Res. 13, 2507–2518 (2003).
Dermitzakis, E.T. et al. Evolutionary discrimination of mammalian conserved non-genic sequences (CNGs). Science 302, 1033–1035 (2003).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
International Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002).
Cooper, G.M. et al. Characterization of evolutionary rates and constraints in three mammalian genomes. Genome Res. 14, 539–548 (2004).
Le, S.V., Chen, J.H., Currey, K.M. & Maizel, J.V., Jr. A program for predicting significant RNA secondary structures. Comput. Appl. Biosci. 4, 153–159 (1988).
Washietl, S. & Hofacker, I.L. Consensus folding of aligned sequences as a new measure for the detection of functional RNAs by comparative genomics. J. Mol. Biol. 342, 19–30 (2004).
Hofacker, I.L., Fekete, M. & Stadler, P.F. Secondary structure prediction for aligned RNA sequences. J. Mol. Biol. 319, 1059–1066 (2002).
Griffiths-Jones, S. et al. Rfam: annotating non-coding RNAs in complete genomes. Nucleic Acids Res. 33, D121–D124 (2005).
Accardo, M.C. et al. A computational search for box C/D snoRNA genes in the D. melanogaster genome. Bioinformatics 20, 3293–3301 (2004).
Childs, J.L., Poole, A.W. & Turner, D.H. Inhibition of Escherichia coli RNase P by oligonucleotide directed misfolding of RNA. RNA 9, 1437–1445 (2003).
Lin, J. et al. A universal telomerase RNA core structure includes structured motifs required for binding the telomerase reverse transcriptase protein. Proc. Natl. Acad. Sci. USA 101, 14713–14718 (2004).
Avner, P. & Heard, E. X-chromosome inactivation: counting, choice, and initiation. Nat. Rev. Genet. 2, 59–67 (2001).
Rougeulle, C. & Heard, E. Antisense RNA in imprinting: spreading silence through Air. Trends Genet. 18, 434–437 (2002).
Pang, K.C. et al. RNAdb — comprehensive mammalian noncoding RNA database. Nucleic Acids Res. Database issue. 33, D125–D130 (2005).
Hüttenhofer, A. et al. RNomics: an experimental approach that identifies 201 candidates for novel, small, non-messenger RNAs in mouse. EMBO J. 20, 2943–2953 (2001).
Bachellerie, J.-P., Cavaillé, J. & Hüttenhofer, A. The expanding snoRNA world. Biochimie 84, 775–790 (2002).
Berezikov, E. et al. Phylogenetic shadowing and computational identification of human microRNA genes. Cell 120, 21–24 (2005).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Mattick, J.S. RNA regulation: a new genetics? Nat. Rev. Genet. 5, 316–323 (2004).
Glazov, E.A., Pheasant, M., McGraw, E.A., Bejerano, G. & Mattick, J.S. Ultraconserved elements in insect genomes: a highly conserved intronic sequence implicated in the control of homothorax mrna splicing. Genome Res. 15, 800–808 (2005).
Doudna, J.A. Structural genomics of RNA. Nat. Struct. Biol. 7, 954–956 (2000).
Hartig, J.S., Grüne, I., Najafi-Shoushtari, S.H. & Famulok, M. Sequence-specific detection of microRNAs by signal-amplifying ribozymes. J. Am. Chem. Soc. 126, 722–723 (2004).
Missal, K., Rose, D. & Stadler, P.F. Non-coding RNAs in Ciona intestinalis. Bioinformatics 21, Suppl 2, ii77–ii78 (2005).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Liu, C. et al. NONCODE: an integrated knowledge database of non-coding RNAs. Nucleic Acids Res. Database issue. 33, D112–D115 (2005).
Pesole, G. et al. UTRdb and UTRSite: specialized databases of sequences and functional elements of 5′ and 3′ untranslated regions of eukaryotic mRNAs. Update 2002. Nucleic Acids Res. 30, 335–340 (2002).
Scherer, S.W. et al. Human chromosome 7: DNA sequence and biology. Science 300, 767–772 (2003).
Acknowledgements
This work was supported in part by the Austrian Fonds zur Förderung der Wissenschaftlichen Forschung, Project No. P15893, by the German DFG Bioinformatics Initiative BIZ-6/1-2, and by the Austrian Gen-AU bioinformatics integration network sponsored by bm:bwk.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Fig. 1
Northern Blot analysis of five H/ACA snoRNA candidates. (PDF 30 kb)
Supplementary Table 1
Detailed results of the native screen and the random control screen. (PDF 12 kb)
Supplementary Table 2
MicroRNAs missing from the input set. (PDF 11 kb)
Supplementary Table 3
H/ACA snoRNAs missing from the input set. (PDF 10 kb)
Supplementary Table 4
Selected ncRNAs from literature with conserved RNA secondary structures detected in our screen. (PDF 11 kb)
Supplementary Table 5
50 Selected RNAz Hits in intergenic regions overlapping with 'transfrag' transcriptional map. (PDF 347 kb)
Rights and permissions
About this article
Cite this article
Washietl, S., Hofacker, I., Lukasser, M. et al. Mapping of conserved RNA secondary structures predicts thousands of functional noncoding RNAs in the human genome. Nat Biotechnol 23, 1383–1390 (2005). https://doi.org/10.1038/nbt1144
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt1144
This article is cited by
-
Functional and structural basis of extreme conservation in vertebrate 5′ untranslated regions
Nature Genetics (2021)
-
LncRNA LncHrt preserves cardiac metabolic homeostasis and heart function by modulating the LKB1-AMPK signaling pathway
Basic Research in Cardiology (2021)
-
Long noncoding RNAs coordinate functions between mitochondria and the nucleus
Epigenetics & Chromatin (2017)
-
Comprehensive prediction of lncRNA–RNA interactions in human transcriptome
BMC Genomics (2016)
-
Mycoplasma non-coding RNA: identification of small RNAs and targets
BMC Genomics (2016)