Abstract
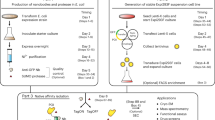
In the affinity purification of recombinant fusion proteins, the rate-limiting step is usually the efficient proteolytic cleavage and removal of the affinity tail and the protease from the purified recombinant protein. We have developed a rapid, convenient and efficient method of affinity purification which can overcome this limitation. In one example of the method, the protease 3C from a picornavirus (3Cpro), which cleaves specific sequences containing a minimum of 6–7 amino acids, has been expressed as a fusion with glutathione S-transferase. The resultant recombinant ‘fusion protease’ cleaves fusion proteins bearing (from the amino-terminus) the same affinity tail as the fusion protease, a 3Cpro cleavage recognition site, and the recombinant protein of interest. The recombinant protein is purified in a single chromatographic step which removes both the affinity tail and the fusion protease. The advantages over existing methods include much improved specificity of proteolytic cleavage, complete removal of the protease and the affinity tail in one step, and the option of adding any desired amount of fusion protease to ensure efficient cleavage. The potential flexibility of the method is shown by the use of various affinity tails and alternative fusion proteases.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Uhlen, M., Forsberg, G., Moks, T., Hartmanis, M. and Nilsson, B. 1992. Fusion proteins in biotechnology. Curr. Opin. Biotechnol. 3: 363–369.
Ford, C.F., Suominen, I. and Glatz, C.E. 1991. Fusion tails for the recovery and purification of recombinant proteins. Prot. Exp. Pur. 2: 95–107.
Smith, D.B. and Johnson, K.S. 1988. Single-step purification of polypeptides expressed in Escherichia coli as fusions with glutathione S-transferase. Gene 67: 31–40.
Arnold, F.H. 1991. Metal-affinity separations: a new dimension in protein processing Bio/Technology. 9: 151–156.
Van Dyke, M.W., Sirito, M. and Sawadogo, M. 1992. Single-step purification of bacterially expressed polypeptides containing an oligo-histidine domain. Gene 111: 99–104.
di Guan, C., Li, P., Riggs, P.D. and Inouye, H. 1988. Vectors that facilitate the expression and purification of foreign peptides in Escherichia coli by fusion to maltose-binding protein. Gene 67: 21–30.
Taylor, M.E. and Drickamer, K. 1991. Carbohydrate-recognition domains as tools for rapid purification of recombinant eukaryotic proteins. Biochem. J. 274: 575–580.
Robben, J., Massie, G., Bosmans, E., Wellens, B. and Volckaert, G. 1993. An Escherichia coli plasmid vector system for high-level production and purification of heterologous peptides fused to active chloramphenicol acetyltransferase. Gene 126: 109–113.
Hopp, T.P., Prickett, K.S., Price, V.L., Libby, R.L., March, C.J., Cerretti, D.P., Urdal, D.L. and Conlon, P.J. 1988. A short polypeptide marker sequence useful for recombinant protein identification and purification. Bio/Technology 6: 1204–1210.
Löwenadler, B., Jansson, B., Paleus, S., Holmgren, E., Nilsson, B., Moks, T., Palm, G., Josephson, S., Philipson, L. and Uhlen, M. 1987. A gene fusion system for generating antibodies against short peptides. Gene 58: 87–97.
Chang, J.-Y. 1985. Thrombin specificity. Requirement for apolar amino acids adjacent to the thrombin cleavage site of polypeptide substrate. Eur. J. Biochem. 151: 217–224.
Porter, A.G. 1993. Picornavirus nonstructural proteins: emerging roles in virus replication and inhibition of host cell functions. J. Virol. 67: 6917–6921.
Leong, L.E.-C., Walker, P.A. and Porter, A.G. 1992. Efficient expression and purification of a protease from the common cold virus, human rhinovirus type 14. J. Crystal Growth. 122: 246–252.
Palmenberg, A.C. 1990. Proteolytic processing of picornaviral polyprotein. Annu. Rev. Microbiol. 44: 603–623.
Cordingley, M.G., Callahan, P.L., Sardana, V.V., Garsky, V.M. and Colonno, R.J. 1990. Substrate requirements of human rhinovirus 3C protease for peptide cleavage in vitro. J. Biol. Chem. 265: 9062–9065.
Sommergrüber, W., Ahorn, H., Zöphel, A., Maurer-Fogy, I., Fessl, F., Schnorrenberg, G., Liebig, H.-D., Blaas, D., Kuechler, E. and Skern, T. 1992. Cleavage specificity on synthetic peptide substrates of human rhinovirus 2 proteinase 2A. J. Biol. Chem. 267: 22639–22644.
Stanway, G., Hughes, P.J., Mountford, R.C., Minor, P.D. and Almond, J.W. 1984. The complete nucleotide sequence of a common cold virus: human rhinovirus 14. Nucl. Acids Res. 12: 7859–7875.
Pearson, J.D., DeWald, D.B., Mathews, W.R., Mozier, N.M., Zürcher-Neely, H.A., Heinrikson, R.L., Morris, M.A., McCubbin, W.D., McDonald, J.R., Fraser, E.D., Vogel, H.J., Kay, C.M. and Walsh, M.P. 1990. Amino acid sequence and characterization of a protein inhibitor of protein kinase C. J. Biol. Chem. 265: 4583–4591.
Smith, C.A., Davis, T., Anderson, D., Solam, L., Beckmann, M.P., Jerzy, R., Dower, S.K., Cosman, D. and Goodwin, R.G. 1990. A receptor for tumor necrosis factor defines an unusual family of cellular and viral proteins. Science. 248: 1019–1023.
Phelps, W.C. and Howley, P.M. 1987. Transcriptional trans-activation by the human papillomavirus type 16 E2 gene product. J. Virol. 61: 1630–1638.
Leong, L.E.C., Walker, P.A. and Porter, A.G. 1993. Human rhinovirus-14 protease 3C (3Cpro) binds specifically to the 5′-noncoding region of the viral RNA. J. Biol. Chem. 268: 25735–25739.
Urzainqui, A. and Carrasco, L. 1989. Degradation of cellular proteins during poliovirus infection: studies by two-dimensional gel electrophoresis. J. Virol. 63: 4729–4735.
Brewer, S.J. and Sassenfeld, H.M. 1985. The purification of recombinant proteins using C-terminal polyarginine fusions. Trends Biotechnol. 3: 119–122.
Taylor, M.E. and Drickamer, K. 1992. Expression and purification of the cytoplasmic tail of an endocytic receptor by fusion to a carbohydrate-recognition domain. Prot. Exp. Pur. 3: 308–312.
Van Heeke, G., Shaw, R., Schnizer, R., Couton, J.M., Schuster, S.M. and Wagner, F.W. 1993. Gene fusions with human carbonic anhydrase II for efficient expression and rapid single-step recovery of recombinant proteins: expression of the Escherichia coli F1-ATPase E subunit. Prot. Exp. Pur. 4: 265–274.
Ullmann, A. 1984. One-step purification of hybrid proteins which have β-galactosidase activity. Gene 29: 27–31.
Sverdrup, F. and Kaleem, S.A. 1994. Replication of human papillomavirus (HPV) DNAs supported by the HPV type 18 E1 and E2 proteins. J. Virol. 68: 505–509.
Sambrook, J.L., Fritsch, E.F. and Maniatis, T. 1989. Molecular Cloning: A Laboratory Manual, 2nd ed. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
Cheah, K.C., Sankar, S. and Porter, A.G. 1988. Expression and processing of human rhinovirus type 14 polypeptide precursors in E. coli maxicells. Gene 69: 265–274.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Walker, P., Leong, LC., Ng, P. et al. Efficient and Rapid Affinity Purification of Proteins Using Recombinant Fusion Proteases. Nat Biotechnol 12, 601–605 (1994). https://doi.org/10.1038/nbt0694-601
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nbt0694-601
This article is cited by
-
The Center for Optimized Structural Studies (COSS) platform for automation in cloning, expression, and purification of single proteins and protein–protein complexes
Amino Acids (2014)
-
A simplified counter-selection recombineering protocol for creating fluorescent protein reporter constructs directly from C. elegans fosmid genomic clones
BMC Biotechnology (2013)
-
Immobilized metal ion affinity chromatography: a review on its applications
Applied Microbiology and Biotechnology (2012)
-
Biochemical characterization of trans-sialidase TS1 variants from Trypanosoma congolense
BMC Biochemistry (2011)
-
Optimal use of tandem biotin and V5 tags in ChIP assays
BMC Molecular Biology (2009)