Abstract
We present an in silico approach to identifying neoepitopes derived from intron retention events in tumor transcriptomes. Using mass spectrometry immunopeptidome analysis, we show that retained intron neoepitopes are processed and presented on MHC I on the surface of cancer cell lines. RNA-derived neoepitopes should be considered for prospective personalized cancer vaccine development.
Similar content being viewed by others
Main
Personalized cancer vaccines comprising neoepitope peptides generated from somatic mutations have shown potential as targeted immunotherapies1,2,3. Other types of aberrant peptides, including cancer germline antigens generated from genes that are transcriptionally silent in adult tissues, have been shown to act as tumor neoepitopes in immune rejection4,5. Dysregulation of RNA splicing through intron retention, which is common in tumor transcriptomes6,7, represents another potential source of tumor neoepitopes, but has not been previously explored. Intron retention is caused by splicing errors that lead to inclusion of an intron in the final mRNA transcript. Retained intron (RI) transcripts are translated and degraded by the nonsense-mediated decay pathway, which generates peptides for endogenous processing, proteolytic cleavage and presentation on MHC type I8,9,10.
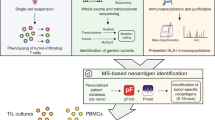
We developed a computational approach to detecting intron retention events from tumor RNA-seq data (Fig. 1a and Online Methods). Intron fragments likely to be translated on the basis of their position downstream of a translated exon and upstream of an in-frame stop codon were identified. Predicted binding affinities between RI peptide sequences and the products of sample-specific HLA class I alleles were calculated to identify candidate RI neoepitopes. We filtered and thresholded preliminary results to exclude artifacts. This process (Online Methods) generated a robust list of putative RI neoepitopes for each sample.
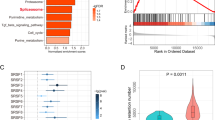
(a) An in silico pipeline detects intron retention events from transcriptome sequencing, determines open reading frames extending into introns, and identifies putative HLA-specific neoepitopes. ORF, open reading frame; WES, whole exome sequencing. (b) Distribution of total RI load, neoepitope-yielding RI load, and RI neoepitope load in patient cohorts (n = 27 Hugo samples, n = 21 Snyder samples). Box plots show the median, first and third quartiles, whiskers extend to 1.5 × the interquartile range, and outlying points are plotted individually. (c) Somatic and RI neoepitope load by patient. Within each cohort, patients are sorted by total neoepitope load. Neoepitope counts (y-axis values) are represented in natural log format.
We applied this method to tumor sequencing data from two cohorts of melanoma patients treated with checkpoint inhibitors11,12 to identify putative RI neoepitopes (n = 48 melanomas; Supplementary Tables 1 and 2). Apart from one outlier, both cohorts had comparable levels of intron retention and predicted RI neoepitopes (Fig. 1b). Slight variation in RI neoepitope load between cohorts was expected given differences in RNA sequencing run, depth, and quality13. The total predicted neoepitope load included RI neoepitopes, as well as somatic mutation neoepitopes derived computationally using published methods (Supplementary Fig. 1, Supplementary Table 1 and Online Methods). Most patients showed substantially augmented total neoepitope loads with the additional consideration of RI neoepitopes. Mean somatic neoepitope load was 2,218 and mean RI neoepitope load was 1,515, yielding a ∼0.7-fold increase in mean total neoepitope load with the addition of RI neoepitopes (Fig. 1c). Excluding one outlier sample with a vastly higher level of somatic neoepitopes than the rest, incorporation of RI neoepitopes roughly doubled the total neoepitope load. There was no significant correlation between somatic neoepitope load and RI neoepitope load (ordinary linear regression P = 0.63; Supplementary Fig. 2).
To demonstrate that RI neoepitopes are processed and presented on MHC I, we predicted RI neoepitopes from six human tumor cell lines and detected neoepitopes that were complexed to MHC I by mass spectrometry (Supplementary Table 3). In melanoma cell line MeWo, the predicted RI neoepitopes EVYAAGKYV and YAAGKYVSF from KCNAB2 (chr1:6142308–6145287) were experimentally discovered in complex with MHC I via mass spectrometry with high confidence (Fig. 2a). We identified RI neoepitopes in another melanoma cell line, SK-MEL-5 (AMSDVSHPK and LAMSDVSHPK from SMARCD1), in B cell lymphoma cell lines CA46 (FRYVAQAGL from LRSAM1) and DOHH-2 (TLFLLSLPL and FLLSLPLPV from CYB561A3), and in leukemia cell lines HL-60 (SVLDDVRGW from TAF1) and THP-1 (LTSQGKSAF from ZCCHC6) (Fig. 2b and Supplementary Fig. 3). Applying this method to somatic mutation–derived neoepitopes, a comparable percentage of predicted neoepitopes were detected by mass spectrometry (Supplementary Table 4). The discovery of peptides in complex with MHC I in cell lines using mass spectrometry with RI neoepitope sequences predicted computationally with our pipeline provides direct evidence of the processing and presentation of RI neoepitopes through the MHC I pathway.
(a) Two RI neoepitopes identified in the MeWo cell line originating from gene KCNAB2 were both predicted in silico and found by mass spectrometry in the MeWo immunopeptidome. Integrative Genomics Viewer (IGV) sashimi plot indicating RNA-seq read depth (RI expression in TPM = 5.13, percent-spliced-in (PSI) value = 1.07%) and mass spectra. Experiments were repeated five times with independent measurements for cell line MeWo. Neoepitopes shown had one peptide-to-spectrum match (PSM) and were identified in one replicate within a 1% false discovery rate. CCLE, Cancer Cell Line Encyclopedia. (b) Predicted RI neoepitopes were found to have mass spectrometric evidence supporting their presentation in complex with MHC I using the same methodology in additional tumor cell lines: SK-MEL-5, CA46, DOHH-2, HL-60 and THP-1.
Given that somatic neoepitope burden is a known correlate of checkpoint inhibitor response in melanoma14, we next examined whether RI neoepitope load might be similarly associated with response. However, there was no association between RI neoepitope load and clinical benefit from checkpoint inhibitor therapy, nor was there correlation with expression of the canonical markers of immune cytolytic activity CD8A, GZMA or PRF115, or clinical covariates (Pearson correlation P > 0.05 for all; Supplementary Figs. 4, 5, 6). Rather, there was a nonsignificant trend toward association between high RI neoepitope load and lack of benefit (two-sided Mann–Whitney U, P = 0.29 Snyder12 cohort, 0.61 Hugo11 cohort). Tumors with high RI neoepitope load and tumors unresponsive to checkpoint inhibitors, with only 38% overlap, shared common transcriptional programs consistent with cell cycle and DNA damage repair activity (Supplementary Fig. 7 and Supplementary Table 5).
Here we demonstrate that tumor-specific RI neoepitopes can be identified computationally in both patient- and cell-line-derived samples and a subset can be validated as presented in complex with MHC I. These data support the hypothesis that aberrant splicing results in intron retention, which generates abnormal transcripts that are translated into immunogenic peptides, loaded on MHC I and presented to the immune system, underscoring their relevance in patients receiving immunotherapy. Further studies will be necessary to clinically validate the immunogenicity of specific RI neoepitopes in patients, including identification of T cells specific to predicted RI neoepitopes.
Furthermore, we found that RI neoepitope load was not associated with checkpoint inhibitor response and discovered that samples from patients with high RI neoepitope load are transcriptionally similar to those whose tumors did not respond to immunotherapy: both patient groups have enrichment of cell cycle and DNA damage repair–related gene sets. Intron retention has been shown to regulate the cell cycle in both nonmalignant16 and malignant cells17. These findings warrant further investigation and experimental validation, given the emerging synergistic relationship between cell cycle inhibition and immune checkpoint blockade therapies18,19,20.
Identification of a wider array of tumor neoepitopes, including those derived from somatic mutation, aberrant gene expression and splicing dysregulation, will contribute to a more complete understanding of the tumor immune landscape. Additional work dissecting the relationship between the prediction, processing and presentation, and ultimate immunogenicity of neoepitopes derived from different sources will be required to ensure clinical relevance of this approach. It has been shown that melanoma in particular may feature certain shared epitopes across patients that are derived from incomplete splicing processes, which may render these cancers more susceptible to RI-derived neoepitopes21,22. Similar approaches across different tissues will provide further clarity on the role of RI neoepitopes in tumor immunity across cancer contexts. Currently, our findings are limited by the availability of clinically annotated cohorts with high-quality RNA sequencing and matched normal tissue. Incorporation of matched normal tissue will improve exclusion of RIs that represent normal gene expression and may help increase precision of our filtering approach. Prediction of patient-specific RI neoepitopes has the potential to contribute to the development of personalized cancer vaccines.
Methods
Clinical cohorts.
Analysis was conducted on published cohorts of melanoma patients treated with immune checkpoint inhibitors. The Hugo et al. cohort included samples from 27 melanoma patients (26 before treatment, 1 on treatment) treated with the PD-1 inhibitor pembrolizumab11. Patient outcomes were classified as responding to therapy (R) (n = 14) or not responding to therapy (NR) (n = 13), as described in the original publication. These samples were sequenced from fresh-frozen tissue using a standard, poly(A)-selecting protocol. The Snyder et al. cohort included post-treatment samples for 21 melanoma patients treated with ipilimumab (anti-CTLA-4 therapy)12,23. Outcomes were classified as receiving long-term clinical benefit (LB) (n = 8) or not receiving clinical benefit (NB) (n = 13), as described in the original publication. RNA sequencing of the Snyder cohort was performed on fresh-frozen tissue using a standard, poly(A)-selecting protocol.
RI neoepitope pipeline.
Raw RNA-seq FASTQ files were pseudoaligned to an augmented hg19 (GENCODE Release 19, GRCh37.p13)24 transcriptome index containing both exonic and intronic transcript sequences, and transcript expression was quantified via kallisto25. The KMA algorithm26, implemented as a suite of Python scripts within an R package, was used to identify the genomic loci of expressed intron retention events with limited false positives. Using these RI loci, the UCSC Table Browser27 database was queried via public MySQL server to obtain the nucleotide sequences corresponding to the intronic regions and fragments of the previous exonic sequences, as well as the open reading frame orientation at the start of the intron. RI peptide sequences of 9 or 10 amino acids, with at least 1 intronic amino acid, were generated by translating open reading frames into intronic sequences until hitting an in-frame stop codon. These peptides, along with sample HLA class I alleles identified via the POLYSOLVER algorithm28, were assessed for putative peptide–MHC I binding affinity via NetMHCpan v3.129. A threshold of rank < 0.5% was used to identify putative RI neoepitopes.
Several filters were applied at various steps throughout the pipeline to eliminate likely false positive RIs and RI neoepitopes. After expression quantification, RIs expressed at a level ≤1 transcript per million, likely artifactual, were eliminated from the analysis. Additional expression-based filters were applied within the KMA algorithm: RIs that did not reach a level of at least 5 unique counts in at least 25% of samples in a cohort and whose neighboring exons did not reach a level of at least 1 transcript per million in at least 25% of samples in a cohort were eliminated as false positives26. Owing to the absence of matched normal RNA-seq data for our melanoma clinical cohorts, a 'panel of normals' approach was taken in an attempt to filter out introns commonly retained in normal skin tissue, which would not produce immunogenic peptides as a result of likely host immune tolerance. RIs were identified in six normal skin samples (three individuals, two samples per individual: subject ERS326932 with samples ERR315339 and ERR315376, subject ERS326943 with samples ERR315372 and ERR315460, and subject ERS327007 with samples ERR315401 and ERR315464) from the Human Protein Atlas. RNA-seq paired-end FASTQ files for each sample were downloaded from the following open-access link: https://www.ebi.ac.uk/arrayexpress/experiments/E-MTAB-1733/samples/. All normal sample retention profiles were highly concordant, both within and across individuals (Supplementary Fig. 8a). The final filter set of 7,050 normal RIs was obtained by intersecting the sets of RIs shared by each unique combination of one sample per individual—eight groups total (Supplementary Fig. 8b and Supplementary Table 6). These RIs were eliminated from downstream tumor sample analyses. In addition, RI peptides with amino acid sequences present in the normal proteome, derived from the UniProt human reference proteome version 2017_03, downloaded on 5 July 2017, were filtered because of likely host immune tolerance30. Finally, a set of RIs that were flagged due to abnormally high expression values and discovered upon manual review via Integrative Genomics Viewer31 to be erroneously annotated in either the reference transcriptome or the Table Browser database were eliminated from the analysis (Supplementary Fig. 9a–d and Supplementary Table 6).
Clinical cohort somatic neoepitope analysis.
Putative somatic neoepitopes were identified in silico for each sample as described in Van Allen et al. 201514. Briefly, BAM files from each cohort underwent sequencing quality control to ensure concordance between tumor and matched normal sequences and adequate depth of sequencing coverage. Single nucleotide variants were called using MuTect32 and insertions and deletions were called using Strelka33. Annotation of identified variants was done using Oncotator (http://www.broadinstitute.org/cancer/cga/oncotator). Sequences of 9- or 10-amino acid peptides with at least one mutant amino acid were generated. These peptides, along with HLA class I alleles called with POLYSOLVER were analyzed using NetMHCpan v3.0 to identify HLA–peptide binding interactions28,29. For each patient, all peptides with predicted binding rank ≤2.0% for at least one patient HLA Class I allele were called somatic neoepitopes.
Cell line analyses.
Raw RNA-seq data from published34 cell lines CA46, DOHH-2, HL-60, THP-1, MeWo and SK-MEL-5 were obtained from the Cancer Cell Line Encyclopedia35 via the NCI Genomic Data Commons and run through our computational pipeline as previously described, with minor adaptations as follows. HLA class I alleles were used for each cell line as enumerated in publication. A threshold of predicted binding rank ≤ 2.0% for at least one HLA class I allele was used to distinguish cell line RI neoepitopes. All pipeline filters applied to patient data described above were implemented on the cell line data except that RI neoepitopes expected to be retained in normal tissue were not filtered because these experiments were focused on presentation of RI neoepitopes rather than immune system stimulation once presented.
Mass spectrometric data from Ritz et al.34, as well as previously unpublished data for cell lines MeWo, DOHH-2 and SK-MEL-5, were searched against a database consisting of 93,250 sequences of the human reference proteome downloaded from UniProt on 7 July 2017 concatenated with putative retained intron sequences (TPM > 1), or concatenated with 133,811 intron sequences with TPM < 1 (not retained) as negative control. Fragment mass spectra were searched with SEQUEST and filtered to a 1% false discovery rate with Percolator to identify high confidence events.
Gene set enrichment analysis.
Gene expression was quantified in patient samples using kallisto25. Gene set enrichment analysis (GSEA) was run to compare both patients in the top quartile vs. bottom quartile of RI load and patients whose tumors responded to immunotherapy vs. those whose did not. Initially, 50 Hallmark gene sets were tested36. GSEA analyses of the Founders gene sets underlying the Hallmark gene sets that were significantly enriched in both of the above comparisons were subsequently performed. All statistical values reported are Benjamini–Hochberg false discovery rate q values corrected for multiple hypothesis testing.
Statistical analyses.
Assessment of difference in means or medians for a continuous variable between two clinical response groups (i.e., clinical benefit vs. no clinical benefit) was performed using the two-sided nonparametric Mann–Whitney U test for non-normally-distributed variables (for example, RI neoepitope burden). All statistical analyses were conducted in the R statistical software environment (v.3.3.1).
Life Sciences Reporting Summary.
Further information on research design is available in the Nature Research Reporting Summary linked to this article.
Code availability.
Pipeline code is publicly accessible on GitHub at https://github.com/vanallenlab/retained-intron-neoantigen-pipeline and as Supplementary Software.
Data availability.
Raw RNA-seq data for the Snyder et al. 2014 patient cohort are available on dbGaP under accession code phs001038.v1.p1 and for the Hugo et al. 201611 cohort on the Sequence Read Archive under accession code SRP070710.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Accession codes
References
Ott, P.A. et al. Nature 547, 217–221 (2017).
Sahin, U. et al. Nature 547, 222–226 (2017).
Carreno, B.M. et al. Science 348, 803–808 (2015).
Hunder, N.N. et al. N. Engl. J. Med. 358, 2698–2703 (2008).
Robbins, P.F. et al. Clin. Cancer Res. 21, 1019–1027 (2015).
Dvinge, H. & Bradley, R.K. Genome Med. 7, 45 (2015).
Jung, H. et al. Nat. Genet. 47, 1242–1248 (2015).
Apcher, S. et al. Proc. Natl. Acad. Sci. USA 108, 11572–11577 (2011).
Rock, K.L., Farfán-Arribas, D.J. & Shen, L. J. Immunol. 184, 9–15 (2010).
Pearson, H. et al. J. Clin. Invest. 126, 4690–4701 (2016).
Hugo, W. et al. Cell 165, 35–44 (2016).
Snyder, A. et al. N. Engl. J. Med. 371, 2189–2199 (2014).
Li, S. et al. Nat. Biotechnol. 32, 888–895 (2014).
Van Allen, E.M. et al. Science 350, 207–211 (2015).
Rooney, M.S., Shukla, S.A., Wu, C.J., Getz, G. & Hacohen, N. Cell 160, 48–61 (2015).
Middleton, R. et al. Genome Biol. 18, 51 (2017).
Dominguez, D. et al. Elife 5, e10288 (2016).
Deng, J. et al. Cancer Discov. 8, 216–233 (2018).
Schaer, D.A. et al. Cell Rep. 22, 2978–2994 (2018).
Goel, S. et al. Nature 548, 471–475 (2017).
Lupetti, R. et al. J. Exp. Med. 188, 1005–1016 (1998).
Andersen, R.S. et al. Oncoimmunology 2, e25374 (2013).
Nathanson, T. et al. Cancer Immunol. Res. 5, 84–91 (2017).
Harrow, J. et al. Genome Res. 22, 1760–1774 (2012).
Bray, N.L., Pimentel, H., Melsted, P. & Pachter, L. Nat. Biotechnol. 34, 525–527 (2016).
Pimentel, H. et al. Nucleic Acids Res. 44, 838–851 (2016).
Karolchik, D. et al. Nucleic Acids Res. 32, D493–D496 (2004).
Shukla, S.A. et al. Nat. Biotechnol. 33, 1152–1158 (2015).
Nielsen, M. & Andreatta, M. Genome Med. 8, 33 (2016).
The UniProt Consortium. Nucleic Acids Res. 45, D158–D169 (2017).
Robinson, J.T. et al. Nat. Biotechnol. 29, 24–26 (2011).
Cibulskis, K. et al. Nat. Biotechnol. 31, 213–219 (2013).
Saunders, C.T. et al. Bioinformatics 28, 1811–1817 (2012).
Ritz, D. et al. Proteomics 16, 1570–1580 (2016).
Barretina, J. et al. Nature 483, 603–607 (2012).
Subramanian, A. et al. Proc. Natl. Acad. Sci. USA 102, 15545–15550 (2005).
Acknowledgements
We are grateful to D. Neri for fruitful discussions, D. Ritz for the purification of HLA peptides from cell lines, and M. Ghandi for assistance in coordinating access to cell line transcriptome data. This work was supported by the BroadNext10, NIH K08 CA188615, NIH R01 CA227388 and a Prostate Cancer Foundation–V Foundation Challenge Award.
Author information
Authors and Affiliations
Contributions
Conception and design: A.C.S., C.A.M., E.M.V.A. Development of methodology: C.A.M., A.C.S., H.P., M.X.H., T.F., D.M., K.-K.W., E.M.V.A. Analysis and interpretation of data (for example, pipeline development, statistical analysis, computational analysis): C.A.M., A.C.S., D.A. Writing, review and/or revision of the manuscript: C.A.M., A.C.S., H.P., M.X.H., D.M., D.A., T.F., K.-K.W., E.M.V.A. Study supervision: E.M.V.A.
Corresponding author
Ethics declarations
Competing interests
E.M.V.A. holds consulting roles with Tango Therapeutics, Invitae and Genome Medical and receives research support from Bristol-Myers Squibb and Novartis. T.F. is an employee of Philochem AG.
Integrated supplementary information
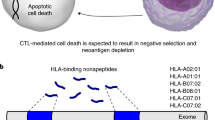
Supplementary Figure 1 Neoepitope presentation pathway illustrations.
Somatic DNA mutations (1) are transcribed (2), spliced (3) and missense mutations are translated (4) and undergo processing into 9-10mer peptides (5), which are presented on the cell surface through the MHC I pathway (6). RI neoepitopes are produced from intact DNA (1), transcribed (2), and undergo defective splicing resulting in intron retention (3). RI transcripts are translated resulting in abnormal peptides and early termination (4). Abnormal proteins are degraded through the NMD pathway, processed into 9-10mer peptides (5), and presented on the cell surface through the MHC-I pathway (6).
Supplementary Figure 2 Retained intron neoepitope load is not associated with somatic neoepitope load in patient cohorts.
Scatterplots illustrate correlation between somatic neoepitope and RI neoepitope loads, with cohort indicated by color (n = 48 patient samples). Two outliers, Hugo_Mel_PD1_Pt8 and Hugo_Mel_PD1_Pt32, indicated on upper plot with asterisks and excluded from lower plot.
Supplementary Figure 3 Mass spectra show RI neoepitopes bound to MHC class I molecules in human cell lines.
Corresponding mass spectrometry plots for RI neoepitopes identified experimentally in complex with MHC-I for each of the cell lines shown in Fig. 2B. Experiments were repeated four times with independent measurements for cell line SK-MEL-5. Neoepitope shown had five peptide-to-spectrum matches (PSMs) and was identified in all four replicates within 1% false discovery rate (FDR). Experiments were repeated four times with independent measurements for CA46. Neoepitope shown had two PSMs and was identified in two replicates within 1% FDR. Experiments were repeated three times with independent measurements for DOHH-2. Neoepitope shown had one PSM and was identified in one replicate within 1% FDR. Experiments were repeated four times with independent measurements for HL-60. Neoepitope shown had one PSM and was identified in one replicate within 1% FDR. Experiments were repeated three times with independent measurements for THP-1. Neoepitope shown had five PSMs and was identified in all three replicates within 1% FDR.
Supplementary Figure 4 RI neoepitope load is not significantly associated with clinical benefit from immunotherapy.
Association of RI load, neoepitope-yielding RI load, and RI neoepitope load with clinical benefit from immunotherapy in Hugo (n = 14 clinical benefit, n = 13 no clinical benefit) and Snyder (n = 8 clinical benefit, n = 13 no clinical benefit) patient cohorts. Boxplots show the median, first, and third quartiles, whiskers extend to 1.5 × the interquartile range, and outlying points are plotted individually. Two-sided Mann-Whitney U p-values > 0.05 for all.
Supplementary Figure 5 Correlation between RI neoepitope load and markers of immune cytolytic activity.
Scatterplots illustrate expression, measured in transcripts per million (TPM), of immune cytolytic activity markers CD8A (top), GZMA (middle), and PRF1 (bottom) vs. RI neoepitope load for both patient cohorts (n = 48 patient samples). Linear trendline and error margins (grey shaded regions) shown, as well as Pearson's correlation coefficients (denoted as rho) and accompanying Pearson's correlation p-values, are denoted on plots.
Supplementary Figure 6 Association between RI neoepitope load and patient clinical characteristics.
Top: Age vs. RI neoepitope load for Snyder cohort (n = 21 patient samples) and Hugo cohort (n = 27 patient samples). Linear trendline and error margins (grey shaded regions) shown, as well as Pearson's correlation coefficients (denoted as rho) and accompanying Pearson's correlation p-values, are denoted on plots. Center left: Disease status vs. RI neoepitope load for both cohorts (n = 48 patient samples). Two-sided Mann-Whitney U p-values shown. Center right: Prior MAP kinase inhibitor therapy vs. RI neoepitope load for Hugo cohort (n = 27 patient samples) (Data not available for Snyder cohort). Two-sided Mann-Whitney U p-values shown. Bottom left: Sex vs. RI neoepitope load for both cohorts (n = 48 patient samples). Two-sided Mann-Whitney U p-values shown. Bottom right: Time of biopsy vs. RI neoepitope load for Snyder cohort (n = 21 patient samples). Two-sided Mann-Whitney U p-values shown. All boxplots show the median, first, and third quartiles, whiskers extend to 1.5 × the interquartile range, and outlying points are plotted individually.
Supplementary Figure 7 Patients with high RI neoepitope loads and immunotherapy nonresponders show enrichment of similar transcriptional programs.
Gene Set Enrichment Analysis (GSEA) was performed comparing top (n = 12) vs. bottom (n = 11) quartile RI neoepitope load patients and immunotherapy nonresponders (n = 10) vs. responders (n = 13). Only half of the top quartile RI neoepitope load patients were overlapping as nonresponders to immunotherapy. Enrichment of cell cycle- and DNA repair-related gene sets was seen in both high RI neoepitope load patients and immunotherapy nonresponders. Representative GSEA enrichment plots from the G2M checkpoint and Downregulation of TLX targets gene sets are shown for both the top vs. bottom quartile RI neoepitope load patients and immunotherapy nonresponders vs. responders comparisons. FDR q-values are indicated on plots.
Supplementary Figure 8 Human Protein Atlas samples were used to create a ‘panel of normals’ for filtering.
A ‘panel of normals’ was created using six Human Protein Atlas (HPA) skin samples (two samples each from three distinct individuals) in order to filter intron retention events likely to occur in normal tissue which would not produce RI neoantigens due to immune tolerance. A, Histogram illustrating the number of unique retained introns shared across samples. The majority of introns are retained by all six normal samples. B, UpSet visualization of set intersections of unique retained introns in each unique grouping of one sample per individual (8 total groupings). The set of 7,050 retained introns shared by all 8 groups of normal samples was denoted the final normal retained intron set and filtered from the RI neoepitope analysis of tumors.
Supplementary Figure 9 Illustrative examples of false positive retained intron events detected upon manual review.
False positive retained intron events were discovered upon manual review of retained introns expressed at aberrantly high levels relative to all intronic expression (> 50 TPM in multiple samples). Likely artifactual introns were filtered from final analysis. IGV screenshots are shown illustrating representative examples. A, Read depth in intron is much higher and more uniform than in neighboring annotated exon; likely a result of transcript annotation error. B, Annotated intron-exon boundary is inconsistent with exon-intron boundary supported by manual review of raw sequencing reads and results in RI neoantigen predicted after an in-frame stop codon. C, Intron expression profile matches surrounding exons and sharply contrasts with other introns in similar region; this intron is likely included in the canonical form of the transcript but not reflected in the annotation. D, Exonic expression of one flanking exon is negligible and does not match with expression profile of other flanking exon, and read depth is low throughout most of the region; first exonic region may be mis-annotated.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Table 4, Supplementary Table Legends and Supplementary Code (PDF 1552 kb)
Supplementary Table 1: Clinical and molecular summary features from Hugo (n = 27) and Snyder (n = 21) patient cohorts.
Clinical characteristics included for each patient: cohort, immunotherapy response status, type of immunotherapy. These characteristics were obtained directly from original publications for each cohort. Molecular characteristics included for each patient: total retained intron (RI) load, neoepitope-yielding RI load, RI neoepitope load, mean number of RI neoepitopes yielded by each RI, somatic neoepitope load. (XLSX 49 kb)
Supplementary Table 2: All RI neoepitopes predicted for each patient in Hugo (n = 27) and Snyder (n = 21) cohorts.
Table contains one patient neoepitope (unique peptide, HLA allele combination) per row. Fields included: Pos (position in original retained intron peptide sequence), Peptide, Intron_ID (genomic coordinates of RI yielding neoepitope), Allele (HLA Class I allele), 1-log50k (NetMHCpan prediction score), nM (NetMHCpan predicted binding affinity, measured in nM), Rank (NetMHCpan rank of predicted affinity compared to a set of random natural peptides), TPM (neoepitope expression level, measured in transcripts per million), SampleID, Gene, Strand (positive or negative genomic strand). (TXT 8709 kb)
Supplementary Table 3: Cancer cell line RI neoepitopes that were both predicted computationally and discovered experimentally bound to MHC Class I molecules via mass spectrometry.
Table contains one cell line neoepitope (unique peptide, HLA allele combination) per row. Rows colored by cell line. Fields included: Cell line, Peptide, Intron ID (genomic coordinates of RI yielding neoepitope), Gene, Strand (positive or negative genomic strand), Allele (HLA Class I allele), 1-log50k (NetMHCpan prediction score), nM (NetMHCpan predicted binding affinity, measured in nM), rank (NetMHCpan rank of predicted affinity compared to a set of random natural peptides), Expression (neoepitope expression level, measured in transcripts per million). (XLSX 74 kb)
Supplementary Table 5: Gene set enrichment analysis results for Hallmark and corresponding Founders gene sets comparing both top quartile vs. bottom quartile RI neoepitope load patients and immunotherapy responders vs. nonresponders.
File contains raw Gene Set Enrichment Analysis (GSEA) results, with four tabs corresponding to Tables S4A-D. A: Hallmark gene sets, top quartile vs. bottom quartile RI neoepitope load. B: Hallmark gene sets, immunotherapy responders vs. nonresponders. C: Founders gene sets, top quartile vs. bottom quartile RI neoepitope load. D: Founders gene sets, immunotherapy responders vs. nonresponders. Founders results reported for all significantly enriched Hallmark gene sets. (XLSX 202 kb)
Supplementary Table 6: Retained introns filtered from RI neoepitope analysis due to either (a) presence in normal skin tissue yielding likely immune tolerance or (b) determination of false-positive nature upon manual review.
File contains two tabs corresponding to Tables S5A-B. A: Introns retained in Human Protein Atlas (HPA) normal skin tissue that were filtered from RI neoepitope analysis of patient tumors due to likely host immune competence (n = 7,050). B: Introns filtered from analysis of patient tumors after manual review (n = 63). (XLSX 175 kb)
Rights and permissions
About this article
Cite this article
Smart, A., Margolis, C., Pimentel, H. et al. Intron retention is a source of neoepitopes in cancer. Nat Biotechnol 36, 1056–1058 (2018). https://doi.org/10.1038/nbt.4239
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4239
This article is cited by
-
Computational immunogenomic approaches to predict response to cancer immunotherapies
Nature Reviews Clinical Oncology (2024)
-
BamQuery: a proteogenomic tool to explore the immunopeptidome and prioritize actionable tumor antigens
Genome Biology (2023)
-
Comprehensive analysis of neoantigens derived from structural variation across whole genomes from 2528 tumors
Genome Biology (2023)
-
Noncoding translation mitigation
Nature (2023)
-
CamoTSS: analysis of alternative transcription start sites for cellular phenotypes and regulatory patterns from 5' scRNA-seq data
Nature Communications (2023)