Abstract
No existing method to characterize transcription factor (TF) binding to DNA allows genome-wide measurement of all TF-binding activity in cells. Here we present a massively parallel protein activity assay, active TF identification (ATI), that measures the DNA-binding activity of all TFs in cell or tissue extracts. ATI is based on electrophoretic separation of protein-bound DNA sequences from a highly complex DNA library and subsequent mass-spectrometric identification of the DNA-bound proteins. We applied ATI to four mouse tissues and mouse embryonic stem cells and found that, in a given tissue or cell type, a small set of TFs, which bound to only ∼10 distinct motifs, displayed strong DNA-binding activity. Some of these TFs were found in all cell types, whereas others were specific TFs known to determine cell fate in the analyzed tissue or cell type. We also show that a small number of TFs determined the accessible chromatin landscape of a cell, suggesting that gene regulatory logic may be simpler than previously appreciated.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
References
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Wang, J. et al. A protein interaction network for pluripotency of embryonic stem cells. Nature 444, 364–368 (2006).
Takahashi, K. & Yamanaka, S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell 126, 663–676 (2006).
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Feng, B. et al. Reprogramming of fibroblasts into induced pluripotent stem cells with orphan nuclear receptor ESRRB. Nat. Cell Biol. 11, 197–203 (2009).
Vaquerizas, J.M., Kummerfeld, S.K., Teichmann, S.A. & Luscombe, N.M. A census of human transcription factors: function, expression and evolution. Nat. Rev. Genet. 10, 252–263 (2009).
Uhlén, M. et al. Proteomics. Tissue-based map of the human proteome. Science 347, 1260419 (2015).
Jolma, A. et al. DNA-binding specificities of human transcription factors. Cell 152, 327–339 (2013).
ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012).
Garber, M. et al. A high-throughput chromatin immunoprecipitation approach reveals principles of dynamic gene regulation in mammals. Mol. Cell 47, 810–822 (2012).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Nitta, K.R. et al. Conservation of transcription factor binding specificities across 600 million years of bilateria evolution. eLife 4 (2015).
Lim, L.S. et al. ZIC3 is required for maintenance of pluripotency in embryonic stem cells. Mol. Biol. Cell 18, 1348–1358 (2007).
Loh, Y.H. et al. The OCT4 and NANOG transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 38, 431–440 (2006).
Ivanova, N. et al. Dissecting self-renewal in stem cells with RNA interference. Nature 442, 533–538 (2006).
Jiang, J. et al. A core KLF circuitry regulates self-renewal of embryonic stem cells. Nat. Cell Biol. 10, 353–360 (2008).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor OCT4. Cell 95, 379–391 (1998).
Jolma, A. et al. DNA-dependent formation of transcription factor pairs alters their binding specificity. Nature 527, 384–388 (2015).
Tun, T. et al. Recognition sequence of a highly conserved DNA-binding protein RBP-Jκ. Nucleic Acids Res. 22, 965–971 (1994).
Wei, G.H. et al. Genome-wide analysis of ETS-family DNA-binding in vitro and in vivo. EMBO J. 29, 2147–2160 (2010).
Wikström, L. et al. Abnormal heart rate and body temperature in mice lacking thyroid hormone receptor–α1. EMBO J. 17, 455–461 (1998).
Adams, B. et al. Pax5 encodes the transcription factor BSAP and is expressed in B lymphocytes, the developing CNS and adult testis. Genes Dev. 6, 1589–1607 (1992).
Urbánek, P., Wang, Z.Q., Fetka, I., Wagner, E.F. & Busslinger, M. Complete block of early B cell differentiation and altered patterning of the posterior midbrain in mice lacking PAX5 (BSAP). Cell 79, 901–912 (1994).
Mach, B., Steimle, V., Martinez-Soria, E. & Reith, W. Regulation of MHC class II genes: lessons from a disease. Annu. Rev. Immunol. 14, 301–331 (1996).
Poirier, R. et al. Distinct functions of Egr gene family members in cognitive processes. Front. Neurosci. 2, 47–55 (2008).
Nakakura, E.K. et al. Mammalian Scratch: a neural-specific Snail family transcriptional repressor. Proc. Natl. Acad. Sci. USA 98, 4010–4015 (2001).
Sugitani, Y. et al. BRN-1 and BRN-2 share crucial roles in the production and positioning of mouse neocortical neurons. Genes Dev. 16, 1760–1765 (2002).
Wang, N.D. et al. Impaired energy homeostasis in C/EBP-α-knockout mice. Science 269, 1108–1112 (1995).
Frain, M. et al. The liver-specific transcription factor LF-B1 contains a highly diverged homeobox DNA-binding domain. Cell 59, 145–157 (1989).
Pontoglio, M. et al. Hepatocyte nuclear factor 1 inactivation results in hepatic dysfunction, phenylketonuria and renal Fanconi syndrome. Cell 84, 575–585 (1996).
Fonjallaz, P., Ossipow, V., Wanner, G. & Schibler, U. The two PAR leucine zipper proteins TEF and DBP display similar circadian and tissue-specific expression but have different target promoter preferences. EMBO J. 15, 351–362 (1996).
Du, Y. et al. Human hepatocytes with drug metabolic function induced from fibroblasts by lineage reprogramming. Cell Stem Cell 14, 394–403 (2014).
Huang, P. et al. Direct reprogramming of human fibroblasts to functional and expandable hepatocytes. Cell Stem Cell 14, 370–384 (2014).
Weirauch, M.T. et al. Determination and inference of eukaryotic transcription factor sequence specificity. Cell 158, 1431–1443 (2014).
Gupta, S., Stamatoyannopoulos, J.A., Bailey, T.L. & Noble, W.S. Quantifying similarity between motifs. Genome Biol. 8, R24 (2007).
Branca, R.M. et al. HiRIEF LC-MS enables deep proteome coverage and unbiased proteogenomics. Nat. Methods 11, 59–62 (2014).
Gabig, T.G., Mantel, P.L., Rosli, R. & Crean, C.D. Requiem: a novel zinc-finger gene essential for apoptosis in myeloid cells. J. Biol. Chem. 269, 29515–29519 (1994).
Gaarenstroom, T. & Hill, C.S. TGF-β signaling to chromatin: how SMADs regulate transcription during self-renewal and differentiation. Semin. Cell Dev. Biol. 32, 107–118 (2014).
Zhang, J. et al. Retinoic acid induces embryonic stem cell differentiation by altering both encoding RNA and microRNA expression. PLoS One 10, e0132566 (2015).
Kokkinopoulos, I. et al. Cardiomyocyte differentiation from mouse embryonic stem cells using a simple and defined protocol. Dev. Dyn. 245, 157–165 (2016).
Lambert, S.A. et al. The human transcription factors. Cell 172, 650–665 (2018).
Yin, Y. et al. Impact of cytosine methylation on DNA-binding specificities of human transcription factors. Science 356, eaaj2239 (2017).
de Boer, C.G. et al. A unified model for yeast transcript definition. Genome Res. 24, 154–166 (2014).
Yue, F. et al. A comparative encyclopedia of DNA elements in the mouse genome. Nature 515, 355–364 (2014).
Bailey, S.D. et al. ZNF143 provides sequence specificity to secure chromatin interactions at gene promoters. Nat. Commun. 2, 6186 (2015).
Noble, W.S., Kuehn, S., Thurman, R., Yu, M. & Stamatoyannopoulos, J. Predicting the in vivo signature of human gene regulatory sequences. Bioinformatics 21 (Suppl. 1), i338–i343 (2005).
Lee, D., Karchin, R. & Beer, M.A. Discriminative prediction of mammalian enhancers from DNA sequence. Genome Res. 21, 2167–2180 (2011).
Thurman, R.E. et al. The accessible chromatin landscape of the human genome. Nature 489, 75–82 (2012).
Yan, J. et al. Transcription factor binding in human cells occurs in dense clusters formed around cohesin anchor sites. Cell 154, 801–813 (2013).
Audic, S. & Claverie, J.M. The significance of digital gene expression profiles. Genome Res. 7, 986–995 (1997).
Kaplan, N. et al. The DNA-encoded nucleosome organization of a eukaryotic genome. Nature 458, 362–366 (2009).
Chronis, C. et al. Cooperative binding of transcription factors orchestrates reprogramming. Cell 168, 442–459.e20 (2017).
Fryer, C.J. & Archer, T.K. Chromatin remodeling by the glucocorticoid receptor requires the BRG1 complex. Nature 393, 88–91 (1998).
Li, Z. et al. FOXA2 and H2A.Z mediate nucleosome depletion during embryonic stem cell differentiation. Cell 151, 1608–1616 (2012).
Stampfel, G. et al. Transcriptional regulators form diverse groups with context-dependent regulatory functions. Nature 528, 147–151, 10.1038/nature15545 (2015).
Zaret, K.S. & Mango, S.E. Pioneer transcription factors, chromatin dynamics and cell fate control. Curr. Opin. Genet. Dev. 37, 76–81 (2016).
Sherwood, R.I. et al. Discovery of directional and nondirectional pioneer transcription factors by modeling DNase profile magnitude and shape. Nat. Biotechnol. 32, 171–178 (2014).
Cirillo, L.A. et al. Opening of compacted chromatin by early developmental transcription factors HNF3 (FOXA) and GATA-4. Mol. Cell 9, 279–289 (2002).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
Morris, S.A. et al. Dissecting engineered cell types and enhancing cell fate conversion via CellNet. Cell 158, 889–902 (2014).
Simicevic, J. et al. Absolute quantification of transcription factors during cellular differentiation using multiplexed targeted proteomics. Nat. Methods 10, 570–576 (2013).
Schmitges, F.W. et al. Multiparameter functional diversity of human C2H2 zinc finger proteins. Genome Res. 26, 1742–1752 (2016).
Imbeault, M., Helleboid, P.Y. & Trono, D. KRAB zinc-finger proteins contribute to the evolution of gene regulatory networks. Nature 543, 550–554 (2017).
Zhang, Y. et al. Model-based analysis of ChIP-seq (MACS). Genome Biol. 9, R137 (2008).
Neph, S. et al. BEDOPS: high-performance genomic feature operations. Bioinformatics 28, 1919–1920 (2012).
Jolma, A. et al. Multiplexed massively parallel SELEX for characterization of human transcription factor–binding specificities. Genome Res. 20, 861–873 (2010).
Korhonen, J., Martinmäki, P., Pizzi, C., Rastas, P. & Ukkonen, E. MOODS: fast search for position-weight-matrix matches in DNA sequences. Bioinformatics 25, 3181–3182 (2009).
Pizzi, C., Rastas, P. & Ukkonen, E. Finding significant matches of position-weight matrices in linear time. IEEE/ACM Trans. Comput. Biol. Bioinform. 8, 69–79 (2011).
Hubner, N.C., Nguyen, L.N., Hornig, N.C. & Stunnenberg, H.G. A quantitative proteomics tool to identify DNA–protein interactions in primary cells or blood. J. Proteome Res. 14, 1315–1329 (2015).
Acknowledgements
We thank J. Yan, E. Kaasinen, B. Schmierer and Y. Yin for critical review of the manuscript, and S. Augsten, L. Hu and P. Pandey for technical assistance. This work was supported by the Center for Innovative Medicine at the Karolinska Institutet (2015–2017; J.T.), the Knut and Alice Wallenberg Foundation (KAW 2013.0088; J.T.), the Göran Gustafsson Foundation (2011–2013; J.T.) and the Swedish Research Council (Vetenskapsrådet; Rådsprofessorprogrammet D0815201; J.T.).
Author information
Authors and Affiliations
Contributions
I.S. collected mouse tissue samples; B.W. extracted proteins, performed ATI experiments and analyzed the data; F. Zhong and F. Zhu performed the DHS analysis; B.S. performed the iHep reprogramming experiment; L.M.O. and J.L. performed the MS experiments and data analysis; A.J., T.K. and M.T. helped to supervise the project or related experiments; and B.W. and J.T. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Analysis of motifs detected in ATI assay using extract from feeder free mouse ES cells.
a, The motifs detected in ATI assay by using the mouse ES cells extract are compared with reference motifs detected by using bacterial expressed pure proteins in HT-SELEX9. The reference binding motif of TF RBPJ (asterisk) is from T. Tun et al.20; the binding motif of TF NFY (asterisk) is from the HOCOMOCO database71. The TF families or specific TFs are proposed based on the comparison of the motifs.
b, The pie charts indicate the percentage of reads containing different numbers of seed matches to the strong motifs found in mouse ES cells. Top: matches in the original input DNA pool (“Cycle 0”). Bottom: matches in the ATI-enriched DNA pool (“Cycle 4”).
71. Kulakovskiy, I. V. et al. HOCOMOCO: a comprehensive collection of human transcription factor binding sites models. Nucleic Acids Res 41, D195-202, doi:10.1093/nar/gks1089 (2013).
Supplementary Figure 2 Motif analysis of ATI data using nuclear extract from different types of mouse tissues and cell lines.
a, “Common” and “Shared” motifs that are found in different mouse cell and tissue types are compared with corresponding motifs detected by using bacterially expressed pure proteins using HT-SELEX9. There is one exception that is not in the SELEX database and corresponds to motif of TF RBPJ; the reference motif of RBPJ (asterisk) is from T. Tun et al.20, 1994.
b, Comparison of binding activities for all five “common” TFs detected in all tested samples based on the absolute molecular counts12 of each motif in the sequencing data. The areas of circles indicate the activities of the five common motifs in the indicated tissues. Data from the last cycle (cycle 4) are used as signals, and data from the previous cycle (cycle 3) are used as background to determine enrichment in one single ATI cycle. For each motif, the activities are normalized by setting its highest activity in any of the tissues as 1.
c, The motifs detected in ATI assay by using different mouse cell and tissue samples are compared with the similar motifs detected by using bacterial expressed pure proteins in HT-SELEX9. The binding motif of TF NFY (asterisk) is from the HOCOMOCO database71. For TFs having the unique binding motifs, the results are validated by the mRNA expression of those TFs; for TFs sharing the same binding motifs, the specific members are proposed based on the mRNA expression levels45,72 and functional data from previous studies.
71. Kulakovskiy, I. V. et al. HOCOMOCO: a comprehensive collection of human transcription factor binding sites models. Nucleic Acids Res 41, D195-202, doi:10.1093/nar/gks1089 (2013).
72. Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat Methods 5, 621-628, doi:10.1038/nmeth.1226 (2008).
Supplementary Figure 3 Reprogramming of induced human hepatocytes confirms the results of ATI assay.
a, Reprogramming timeline for direct conversion of human fibroblasts to induced hepatocytes (iHep).
b, Bright field images of iHep colonies from human fibroblasts after lentiviral transduction of TF combinations previously reported in Morris et al.61 (Set_a; FOXA1, HNF4A, KLF5), Du et al.33 (Set_b; HNF4A, HNF1A, HNF6/ONECUT1, ATF5, PROX1, CEBPA), Huang et al.34 (Set_c; FOXA3, HNF4A, HNF1A) and factors identified by ATI in mouse liver (Set_ATI; HNF1A, HNF1B, DBP, MAFG, CEBPA, CEBPB, HNF4A, HNF6, ESRRA). Experiment was repeated twice independently and similar result was obtained.
c, Expression levels of the liver-specific marker gene ALBUMIN in iHep cells normalized to GAPDH levels by qRT-PCR using previously reported TF cocktails and ATI-identified TF combinations. Bars indicate the means of two independent duplicate samples.
Supplementary Figure 4 ATI assay from multiple species reveals deep conservation of TF activity.
ATI analysis of four different species indicates that the assay can identify TF activities from a wide variety of organisms. The names of the TF that bind to motifs that are similar to those identified using ATI are shown above the sequence logos. Histograms on top show background corrected absolute molecular counts12 (y-axis, Motif counts) of all discovered motifs at enriched ATI cycle; for each sample the highest count is normalized to 100%. Counts more than 10% of the maximum are indicated by red bars; the relative activities of them are shown on the right corner of the corresponding sequence logos. “Unknown motifs” indicate that the motifs could not be assigned to a known TF based on the literature; there are much more unknown motifs detected in other species than in mammals (Figure 2a), suggesting that the binding specificity landscape of these species is less well characterized.
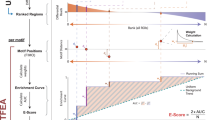
Supplementary Figure 5 Enrichment of ATI 10-mers in DNase I hypersensitive sites from different mouse tissues.
a, ATI enriched 10-mers from mouse tissues are also enriched in DNase I hypersensitive sites from the corresponding tissues. The dot plots show matches to enriched ATI 10-mers in DNase I hypersensitive sites from the indicated mouse tissues. In each dot plot, each row indicates one DHS region from the relative mouse tissues that is flanked with its genomic sequences. Red dots indicate the boundaries of the DHS regions, blue dots indicate positions of top 2000 ATI-enriched 10-mers out of all 410 (~ 1 million) 10-mers. The graph on top shows the average of scores for each 10-mer at each position across the rows.
b, Prediction of DHS regions by using the 10-mer data from the ATI assay. DHSs are sorted by position of the prediction call (yellow line). Black horizontal lines separate accurate DHS calls (middle) from calls more than 500 bp off the known DHS center that is located at the x-axis position 0 in all cases. The fraction of predictions within ± 500 bp of the center and the corresponding p-value (based on winflat) for null model where position calls are randomly distributed are also indicated. The optimization of scoring is indicated in the “Methods”. Numbers of DHSs for different tissues are as follows, heat: 4096, spleen: 4668, brain: 4865, liver: 4473.
Supplementary Figure 6 Comparison of subsequences enriched in ATI and in DHS regions from mouse ES cells.
The enrichment of all 10-mer sequences in the ATI data (y-axis) and DHS data (x-axis) from ES cells is shown. X-axis indicates the log2 fold change of 10-mer counts in DHS regions compared with non-DHS regions; y-axis indicates fold change of 10-mer counts in ATI enriched DNA pool (Cycle 4) compared with original pool (Cycle 0). Coloring of the dots indicates 10-mers that are similar to the motifs shown on the right; black dots indicate the 10-mers that are not similar to any motifs. One 10-mer sequence (“CGGCGGCGGC”) is shown as an example of repetitive CG rich sequences which displays high enrichment in DHS regions but no enrichment in ATI. The reason why such CG rich sequences enrich in gene regulatory elements may be due to the fact that methylated C is prone to mutation, and the low CpG methylation rate of regulatory elements protects these sequences from this mutational process73,74.
73. Coulondre, C., Miller, J. H., Farabaugh, P. J. & Gilbert, W. Molecular-Basis of Base Substitution Hotspots in Escherichia-Coli. Nature 274, 775-780, doi:DOI 10.1038/274775a0 (1978).
74. Hodgkinson, A. & Eyre-Walker, A. Variation in the mutation rate across mammalian genomes. Nature Reviews Genetics 12, 756-766, doi:10.1038/nrg3098 (2011).
Supplementary Figure 7 ATI assay by using genomic DNA in mouse ES cells.
a, De novo motif mining of genomic fragments bound by nuclear extract from mouse ES cells. Top twelve motifs are shown with the corresponding E-values. Motifs 4, 5, 8 and 10 were similar to motifs detected in the ATI assay.
b, Overlap between the 25,261 DHS regions from mES cells (DHSs, blue) and the peaks called from genomic fragments bound by nuclear extract (ATI peaks, red) or the peaks called from genomic fragments not bound by nuclear extract (Control peaks, red). The peak analyzed is considered overlapping with DHSs if not less than 100% (top), 50% (middle) or 1bp length (bottom) of the peak overlaps with the DHS regions. The numbers inside the circles indicate numbers of DHSs or peaks. The right panel shows three specific loci exemplifying the non-overlapped DHS regions without ATI peaks (top), DHS and ATI overlapped regions (middle) and non-overlapped ATI peaks (bottom). Relatively low overlap between ATI peaks and DHS regions indicate that the open chromatin in cells doesn’t result exclusively from the action of TFs with strong binding activity, or that binding motifs of strongly bound TFs would be sufficient to open closed chromatin states characterized by presence of HP1, histone H1 or repressive chromatin modifications75.
Supplementary Figure 8 Analysis of features of different categories of DHS fragments.
a-b, Comparison of features between DHS regions that are easy to predict (tritile_1), intermediate (tritile_2) and hard to predict (tritile_3) using DHS (a) or ATI (b) 10-mers. The DHS fragments (n= 15533) are 1 kb non-overlapping “DHS” fragments used for the final prediction in the Precision-recall analysis. Top: prediction score (sum of the scores of all 10-mers inside the window). Middle: distance from TSS. Bottom: percentage of CpG dinucleotides. Note that hard to predict DHSs using ATI and DHS 10-mer data tend to be farther from a TSS and have a low CpG content. For all six boxplots, the top and bottom of the box represent 25th and 75th percentile, and the centerline is the mean; the upper whisker is located at the smaller of the maximum value and 75th percentile + 1.5 IQR, and the lower whisker is located at the larger of the minimum value and 25th percentile – 1.5 IQR; dots represent the outliers, which are less than 25th percentile – 1.5 IQR or more than 75th percentile + 1.5 IQR.
c, The correlation between the ATI and DHS total prediction scores for all the 1 kb DHS fragments used for the final prediction in the Precision-recall analysis (n = 15533). Each dot represents one fragment and the color indicates the percentage of CpG dinucleotides within the fragment.
d, De novo motif mining of four different types of DHS regions from mouse ES cells. The different types of DHS regions are generated from intersection of different categories of DHS fragments (see “Methods” for details). The “known motifs” indicate the motifs can be assigned to the known motifs based on current knowledge. All known motifs with E-value less than 0.01, and top five unknown/repetitive motifs are shown.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 (PDF 1838 kb)
Supplementary Tables
Supplementary Tables 1–3, 5–10 (XLSX 4553 kb)
Supplementary Table 4
Comparison of the motif analysis result and MS identification result in ATI from nuclear extract of mouse ES cells. (PDF 3135 kb)
Rights and permissions
About this article
Cite this article
Wei, B., Jolma, A., Sahu, B. et al. A protein activity assay to measure global transcription factor activity reveals determinants of chromatin accessibility. Nat Biotechnol 36, 521–529 (2018). https://doi.org/10.1038/nbt.4138
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.4138
This article is cited by
-
Hold out the genome: a roadmap to solving the cis-regulatory code
Nature (2024)
-
A universal system for boosting gene expression in eukaryotic cell-lines
Nature Communications (2024)
-
ExplaiNN: interpretable and transparent neural networks for genomics
Genome Biology (2023)
-
PIM1 promotes hepatic conversion by suppressing reprogramming-induced ferroptosis and cell cycle arrest
Nature Communications (2022)
-
Sequence determinants of human gene regulatory elements
Nature Genetics (2022)