Abstract
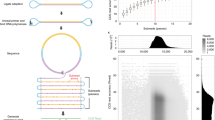
Eliminating errors in next-generation DNA sequencing has proved challenging. Here we present error-correction code (ECC) sequencing, a method to greatly improve sequencing accuracy by combining fluorogenic sequencing-by-synthesis (SBS) with an information theory–based error-correction algorithm. ECC embeds redundancy in sequencing reads by creating three orthogonal degenerate sequences, generated by alternate dual-base reactions. This is similar to encoding and decoding strategies that have proved effective in detecting and correcting errors in information communication and storage. We show that, when combined with a fluorogenic SBS chemistry with raw accuracy of 98.1%, ECC sequencing provides single-end, error-free sequences up to 200 bp. ECC approaches should enable accurate identification of extremely rare genomic variations in various applications in biology and medicine.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Shendure, J., Mitra, R.D., Varma, C. & Church, G.M. Advanced sequencing technologies: methods and goals. Nat. Rev. Genet. 5, 335–344 (2004).
Koboldt, D.C., Steinberg, K.M., Larson, D.E., Wilson, R.K. & Mardis, E.R. The next-generation sequencing revolution and its impact on genomics. Cell 155, 27–38 (2013).
Drmanac, R. The advent of personal genome sequencing. Genet. Med. 13, 188–190 (2011).
Mardis, E.R. & Wilson, R.K. Cancer genome sequencing: a review. Hum. Mol. Genet. 18, R2, R163–R168 (2009).
Schrijver, I. et al. Opportunities and challenges associated with clinical diagnostic genome sequencing: a report of the Association for Molecular Pathology. J. Mol. Diagn. 14, 525–540 (2012).
Goodwin, S., McPherson, J.D. & McCombie, W.R. Coming of age: ten years of next-generation sequencing technologies. Nat. Rev. Genet. 17, 333–351 (2016).
Mardis, E.R. A decade's perspective on DNA sequencing technology. Nature 470, 198–203 (2011).
Mardis, E.R. Next-generation sequencing platforms. Annu. Rev. Anal. Chem. (Palo Alto, Calif.) 6, 287–303 (2013).
Metzker, M.L. Sequencing technologies - the next generation. Nat. Rev. Genet. 11, 31–46 (2010).
Shendure, J. & Ji, H. Next-generation DNA sequencing. Nat. Biotechnol. 26, 1135–1145 (2008).
Fuller, C.W. et al. The challenges of sequencing by synthesis. Nat. Biotechnol. 27, 1013–1023 (2009).
Eid, J. et al. Real-time DNA sequencing from single polymerase molecules. Science 323, 133–138 (2009).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Braslavsky, I., Hebert, B., Kartalov, E. & Quake, S.R. Sequence information can be obtained from single DNA molecules. Proc. Natl. Acad. Sci. USA 100, 3960–3964 (2003).
Pushkarev, D., Neff, N.F. & Quake, S.R. Single-molecule sequencing of an individual human genome. Nat. Biotechnol. 27, 847–850 (2009).
Gao, Y. et al. Single molecule targeted sequencing for cancer gene mutation detection. Sci. Rep. 6, 26110 (2016).
Ju, J. et al. Four-color DNA sequencing by synthesis using cleavable fluorescent nucleotide reversible terminators. Proc. Natl. Acad. Sci. USA 103, 19635–19640 (2006).
Guo, J., Yu, L., Turro, N.J. & Ju, J. An integrated system for DNA sequencing by synthesis using novel nucleotide analogues. Acc. Chem. Res. 43, 551–563 (2010).
Stupi, B.P. et al. Stereochemistry of benzylic carbon substitution coupled with ring modification of 2-nitrobenzyl groups as key determinants for fast-cleaving reversible terminators. Angew. Chem. Int. Ed. 51, 1724–1727 (2012).
Margulies, M. et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature 437, 376–380 (2005).
Rothberg, J.M. et al. An integrated semiconductor device enabling non-optical genome sequencing. Nature 475, 348–352 (2011).
Sims, P.A., Greenleaf, W.J., Duan, H. & Xie, X.S. Fluorogenic DNA sequencing in PDMS microreactors. Nat. Methods 8, 575–580 (2011).
Chen, Z. et al. Fluorogenic sequencing using halogen-fluorescein-labeled nucleotides. ChemBioChem 16, 1153–1157 (2015).
Wu, W. et al. Termination of DNA synthesis by N6-alkylated, not 3′-O-alkylated, photocleavable 2′-deoxyadenosine triphosphates. Nucleic Acids Res. 35, 6339–6349 (2007).
Rothberg, J.M. & Leamon, J.H. The development and impact of 454 sequencing. Nat. Biotechnol. 26, 1117–1124 (2008).
Forgetta, V. et al. Sequencing of the Dutch elm disease fungus genome using the Roche/454 GS-FLX Titanium System in a comparison of multiple genomics core facilities. J. Biomol. Tech. 24, 39–49 (2013).
Loman, N.J. et al. Performance comparison of benchtop high-throughput sequencing platforms. Nat. Biotechnol. 30, 434–439 (2012).
Liu, L. et al. Comparison of next-generation sequencing systems. J. Biomed. Biotechnol. 2012, 251364 (2012).
Urano, Y. et al. Evolution of fluorescein as a platform for finely tunable fluorescence probes. J. Am. Chem. Soc. 127, 4888–4894 (2005).
Sood, A. et al. Terminal phosphate-labeled nucleotides with improved substrate properties for homogeneous nucleic acid assays. J. Am. Chem. Soc. 127, 2394–2395 (2005).
Rumble, S.M. et al. SHRiMP: accurate mapping of short color-space reads. PLoS Comput. Biol. 5, e1000386 (2009).
Kinde, I., Wu, J., Papadopoulos, N., Kinzler, K.W. & Vogelstein, B. Detection and quantification of rare mutations with massively parallel sequencing. Proc. Natl. Acad. Sci. USA 108, 9530–9535 (2011).
Hoang, M.L. et al. Genome-wide quantification of rare somatic mutations in normal human tissues using massively parallel sequencing. Proc. Natl. Acad. Sci. USA 113, 9846–9851 (2016).
Schmitt, M.W. et al. Detection of ultra-rare mutations by next-generation sequencing. Proc. Natl. Acad. Sci. USA 109, 14508–14513 (2012).
Paten, B., Novak, A. & Haussler, D. Mapping to a reference genome structure. Preprint available at https://arxiv.org/abs/1404.5010v1 (2014).
Acknowledgements
The authors thank Y. Men, Z. Yu, H. Qiu, C. Zheng, Y. Fu, X. Zhang, T. Chen, L. Wu, S. Zhang, X. Jiang, J. Bu, P.A. Sims, L.L. Tao, and H. Ge for experimental assistance and discussion. This work was supported by the Ministry of Science and Technology of China (863 Program 2012AA02A101), National Natural Science Foundation of China (21327808 and 21525521), Beijing Municipal Commission of Science and Technology (Z111100059111002), and Beijing Advanced Innovation Center for Genomics.
Author information
Authors and Affiliations
Contributions
Y.H. and X.S.X. conceived the project. Y.H. and Z.C. designed the experiment. Z.C., W.Z., S.Q., L.K., and H.D. performed the experiments. All authors analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
Patent applications based on this work have been filed by Peking University, and licensed to Cygnus, of which all the authors are consultants, Z.C. and H.D. are shareholders.
Supplementary information
Supplementary Note
Supplementary Information (PDF 13087 kb)
Rights and permissions
About this article
Cite this article
Chen, Z., Zhou, W., Qiao, S. et al. Highly accurate fluorogenic DNA sequencing with information theory–based error correction. Nat Biotechnol 35, 1170–1178 (2017). https://doi.org/10.1038/nbt.3982
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3982
This article is cited by
-
Reconstruction algorithms for DNA-storage systems
Scientific Reports (2024)
-
Nucleic Acids Analysis
Science China Chemistry (2021)
-
Benchmarking of computational error-correction methods for next-generation sequencing data
Genome Biology (2020)
-
Data storage in DNA with fewer synthesis cycles using composite DNA letters
Nature Biotechnology (2019)
-
Sequencing DNA, no mistake
Nature Methods (2018)