Abstract
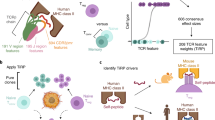
Identification of the peptides recognized by individual T cells is important for understanding and treating immune-related diseases. Current cytometry-based approaches are limited to the simultaneous screening of 10–100 distinct T-cell specificities in one sample. Here we use peptide–major histocompatibility complex (MHC) multimers labeled with individual DNA barcodes to screen >1,000 peptide specificities in a single sample, and detect low-frequency CD8 T cells specific for virus- or cancer-restricted antigens. When analyzing T-cell recognition of shared melanoma antigens before and after adoptive cell therapy in melanoma patients, we observe a greater number of melanoma-specific T-cell populations compared with cytometry-based approaches. Furthermore, we detect neoepitope-specific T cells in tumor-infiltrating lymphocytes and peripheral blood from patients with non-small cell lung cancer. Barcode-labeled pMHC multimers enable the combination of functional T-cell analysis with large-scale epitope recognition profiling for the characterization of T-cell recognition in various diseases, including in small clinical samples.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
McCutcheon, M. et al. A sensitive ELISPOT assay to detect low-frequency human T lymphocytes. J. Immunol. Methods 210, 149–166 (1997).
Altman, J.D. et al. Phenotypic analysis of antigen-specific T lymphocytes. Science 274, 94–96 (1996).
Chattopadhyay, P.K. & Roederer, M. Cytometry: today's technology and tomorrow's horizons. Methods 57, 251–258 (2012).
Bendall, S.C. et al. Single-cell mass cytometry of differential immune and drug responses across a human hematopoietic continuum. Science 332, 687–696 (2011).
Hadrup, S.R. et al. Parallel detection of antigen-specific T-cell responses by multidimensional encoding of MHC multimers. Nat. Methods 6, 520–526 (2009).
Andersen, R.S. et al. Parallel detection of antigen-specific T cell responses by combinatorial encoding of MHC multimers. Nat. Protoc. 7, 891–902 (2012).
Newell, E.W., Klein, L.O., Yu, W. & Davis, M.M. Simultaneous detection of many T-cell specificities using combinatorial tetramer staining. Nat. Methods 6, 497–499 (2009).
Newell, E.W. et al. Combinatorial tetramer staining and mass cytometry analysis facilitate T-cell epitope mapping and characterization. Nat. Biotechnol. 31, 623–629 (2013).
Rammensee, H.G., Falk, K. & Rötzschke, O. Peptides naturally presented by MHC class I molecules. Annu. Rev. Immunol. 11, 213–244 (1993).
Stevanović, S. & Schild, H. Quantitative aspects of T cell activation--peptide generation and editing by MHC class I molecules. Semin. Immunol. 11, 375–384 (1999).
Robins, H.S. et al. Comprehensive assessment of T-cell receptor beta-chain diversity in alphabeta T cells. Blood 114, 4099–4107 (2009).
Davis, M.M. & Bjorkman, P.J. T-cell antigen receptor genes and T-cell recognition. Nature 334, 395–402 (1988).
Wooldridge, L. et al. A single autoimmune T cell receptor recognizes more than a million different peptides. J. Biol. Chem. 287, 1168–1177 (2012).
Xu, Q., Schlabach, M.R., Hannon, G.J. & Elledge, S.J. Design of 240,000 orthogonal 25mer DNA barcode probes. Proc. Natl. Acad. Sci. USA 106, 2289–2294 (2009).
Kivioja, T. et al. Counting absolute numbers of molecules using unique molecular identifiers. Nat. Methods 9, 72–74 (2011).
Toebes, M. et al. Design and use of conditional MHC class I ligands. Nat. Med. 12, 246–251 (2006).
Rodenko, B. et al. Generation of peptide-MHC class I complexes through UV-mediated ligand exchange. Nat. Protoc. 1, 1120–1132 (2006).
Andersen, R.S. et al. Dissection of T-cell antigen specificity in human melanoma. Cancer Res. 72, 1642–1650 (2012).
Lyngaa, R. et al. T-cell responses to oncogenic merkel cell polyomavirus proteins distinguish patients with merkel cell carcinoma from healthy donors. Clin. Cancer Res. 20, 1768–1778 (2014).
Frøsig, T.M. et al. Broadening the repertoire of melanoma-associated T-cell epitopes. Cancer Immunol. Immunother. 64, 609–620 (2015).
van Buuren, M.M. et al. HLA micropolymorphisms strongly affect peptide-MHC multimer-based monitoring of antigen-specific CD8+ T cell responses. J. Immunol. 192, 641–648 (2014).
Frøsig, T.M. et al. Design and validation of conditional ligands for HLA-B*08:01, HLA-B*15:01, HLA-B*35:01, and HLA-B*44:05. Cytometry A 87, 967–975 (2015).
Valmori, D. et al. Enhanced generation of specific tumor-reactive CTL in vitro by selected Melan-A/MART-1 immunodominant peptide analogues. J. Immunol. 160, 1750–1758 (1998).
Derré, L. et al. A novel population of human melanoma-specific CD8 T cells recognizes Melan-AMART-1 immunodominant nonapeptide but not the corresponding decapeptide. J. Immunol. 179, 7635–7645 (2007).
Wieckowski, S. et al. Fine structural variations of alphabetaTCRs selected by vaccination with natural versus altered self-antigen in melanoma patients. J. Immunol. 183, 5397–5406 (2009).
Speiser, D.E. et al. Unmodified self antigen triggers human CD8 T cells with stronger tumor reactivity than altered antigen. Proc. Natl. Acad. Sci. USA 105, 3849–3854 (2008).
Andersen, R. et al. Long-lasting complete responses in patients with metastatic melanoma after adoptive cell therapy with tumor-infiltrating lymphocytes and an attenuated IL2 regimen. Clin. Cancer Res. 22, 3734–3745 (2016).
Snyder, A. et al. Genetic basis for clinical response to CTLA-4 blockade in melanoma. N. Engl. J. Med. 371, 2189–2199 (2014).
Rizvi, N.A. et al. Cancer immunology. Mutational landscape determines sensitivity to PD-1 blockade in non-small cell lung cancer. Science 348, 124–128 (2015).
McGranahan, N. et al. Clonal neoantigens elicit T cell immunoreactivity and sensitivity to immune checkpoint blockade. Science 351, 1463–1469 (2016).
Aleksic, M. et al. Different affinity windows for virus and cancer-specific T-cell receptors: implications for therapeutic strategies. Eur. J. Immunol. 42, 3174–3179 (2012).
Dolton, G. et al. Comparison of peptide-major histocompatibility complex tetramers and dextramers for the identification of antigen-specific T cells. Clin. Exp. Immunol. 177, 47–63 (2014).
Massilamany, C. et al. Direct staining with major histocompatibility complex class II dextramers permits detection of antigen-specific, autoreactive CD4 T cells in situ. PLoS One 9, e87519 (2014).
Han, A., Glanville, J., Hansmann, L. & Davis, M.M. Linking T-cell receptor sequence to functional phenotype at the single-cell level. Nat. Biotechnol. 32, 684–692 (2014).
Dössinger, G. et al. MHC multimer-guided and cell culture-independent isolation of functional T cell receptors from single cells facilitates TCR identification for immunotherapy. PLoS One 8, e61384 (2013).
Klein, A.M. et al. Droplet barcoding for single-cell transcriptomics applied to embryonic stem cells. Cell 161, 1187–1201 (2015).
Donia, M. et al. Characterization and comparison of 'standard' and 'young' tumour-infiltrating lymphocytes for adoptive cell therapy at a Danish translational research institution. Scand. J. Immunol. 75, 157–167 (2012).
Donia, M. et al. Aberrant expression of MHC Class II in melanoma attracts inflammatory tumor-specific CD4+ T- cells, which dampen CD8+ T-cell antitumor reactivity. Cancer Res. 75, 3747–3759 (2015).
Gerlinger, M. et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N. Engl. J. Med. 366, 883–892 (2012).
Gerlinger, M. et al. Genomic architecture and evolution of clear cell renal cell carcinomas defined by multiregion sequencing. Nat. Genet. 46, 225–233 (2014).
Szolek, A. et al. OptiType: precision HLA typing from next-generation sequencing data. Bioinformatics 30, 3310–3316 (2014).
Bakker, A.H. et al. Conditional MHC class I ligands and peptide exchange technology for the human MHC gene products HLA-A1, -A3, -A11, and -B7. Proc. Natl. Acad. Sci. USA 105, 3825–3830 (2008).
Hadrup, S.R. et al. High-throughput T-cell epitope discovery through MHC peptide exchange. Methods Mol. Biol. 524, 383–405 (2009).
Chang, C.X.L. et al. Conditional ligands for Asian HLA variants facilitate the definition of CD8+ T-cell responses in acute and chronic viral diseases. Eur. J. Immunol. 43, 1109–1120 (2013).
Garboczi, D.N., Hung, D.T. & Wiley, D.C. HLA-A2-peptide complexes: refolding and crystallization of molecules expressed in Escherichia coli and complexed with single antigenic peptides. Proc. Natl. Acad. Sci. USA 89, 3429–3433 (1992).
Nielsen, M. et al. NetMHCpan, a method for quantitative predictions of peptide binding to any HLA-A and -B locus protein of known sequence. PLoS One 2, e796 (2007).
Rammensee, H., Bachmann, J., Emmerich, N.P., Bachor, O.A. & Stevanović, S. SYFPEITHI: database for MHC ligands and peptide motifs. Immunogenetics 50, 213–219 (1999).
Langmead, B. & Salzberg, S.L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Robinson, M.D., McCarthy, D.J. & Smyth, G.K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Robinson, M.D. & Oshlack, A. A scaling normalization method for differential expression analysis of RNA-seq data. Genome Biol. 11, R25 (2010).
Acknowledgements
We would like to thank U.K. Hansen, A. Burkal and T. Seremet for technical assistance; T. Schumacher, Netherlands Cancer Institute, for scientific discussions and sharing of MHC expression plasmids; and Dr. Altman, NIH Tetramer Core Facility, for sharing expression plasmids HLA-B*1501 and HLA-B*3501. The work was funded by The Danish Cancer Society (ID:R72-A4531-13-S2), The Lundbeck Foundation Fellowship (ID: R190-2014-4178), The Danish Research Council (FSS-ID: 1331-00283 and DFF-ID:4004-00422), Familien Erichsens Mindefond, Cancer Research UK (FC001169), the UK Medical Research Council (FC001169 ), the Wellcome Trust (FC001169), the UK Medical Research Council (MR/FC001169/1) and the Novo Nordisk Foundation (ID: 16584).
Author information
Authors and Affiliations
Contributions
A.K.B. conceived the concept, designed and performed experiments, analyzed data, made figures and wrote the manuscript; A.M.M. designed the in silicon analysis, analyzed data, and made figures; R.L., S.K.S., and S.R. produced MHC monomers, designed and performed experiments, analyzed data and revised the manuscript; L.S. performed experiments, M.D. and I.M.S. provided patients material and generated tumor cell lines, discussed data; P.t.S. provided administrative support, flow facility and production of MHC monomers; A.J.S.F., S.A.Q. provided patients material and peptides from NSCLC; N.M.G., R.R., C.S. identified tumor mutagenome and predicted NSCLC-associated neoepitopes; Z.S. and A.C.E., designed the in silico analysis, and guided data analyses; S.N.J. designed DNA barcodes, conceived the concept, guided data analyses and revised the manuscript; S.R.H. conceived the concept, designed experiments, analyzed data and wrote the manuscript.
Ethics declarations
Competing interests
The technology is patented (WO2015185067 and WO2015188839) by the authors (S.R.H., A.K.B., and S.N.J.), the Capital Region of Copenhagen, University Hospital Herlev and Immudex. The technology is licensed to Immudex.
Integrated supplementary information
Supplementary Figure 1 Binding capacity of DNA-barcoded MHC multimers and recovery of antigen specificity
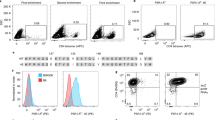
(a, b) Fluorescent-based determination of the binding capacity of DNA-barcoded MHC multimers (+barcode) compared to fluorescent labeled MHC multimers (-barcode). DNA-barcoded MHC multimer reagents were assembled with two identical DNA barcodes attached to each multimerization backbone and subsequent co-attachment of pMHC molecules. Non-barcoded multimers were generated similarly without prior attachment of DNA barcodes (i.e. ‘-barcode’ and ‘+barcode’ were both assembled on a dextramer backbone). Reagents assembled with HLA-A*0201 CMV pp65 NLV were applied for staining of (a) healthy donor PBMCs, and reagents assembled with HLA-A*0201 hTERT p988 (YLQVNSLQTV) were applied for staining of (b) expanded TILs from a melanoma patient. The binding capacity is evaluated in terms of the stain index of the multimer fluorescent intensity of T cells stained with non-barcoded or DNA-barcoded MHC multimers respectively, along with the frequencies of the given multimer positive cell population of CD8 T cells, (a) 0.9-1.2% and (b) 3.8%-4.3%. Bar plots show mean stain index values of three stainings ± SD. (c) Fluorescent-based analysis of antigen specific T cells stained with relevant virus pMHC multimers and excess of irrelevant pMHC multimers. An equimolar (1:1) mixture of individually barcode labeled HLA-B*0702 CMV pp65TPR-multimers and HLA-A*0201 HIV PolILK-multimers, or a mixture with 998 additional fluorescent labeled pMHC multimers, were used for staining of healthy donor PBMCs. The 998 additional MHC multimers comprised equal amounts of irrelevant-peptide HLA-A*0201 and HLA-B*0702 multimers, i.e. multimers carrying MHC molecules refolded with UV-sensitive ligand (Online Methods). Using either reagent mixture, the concentrations of each equimolar pMHC were 23 nM in the final staining volume, i.e. for staining with the 1:1:998 equimolar reagents, the volume of the MHC multimer pool were reduced 50x. Percentages of the multimer positive population of CD8 T cells are given in dot plots. (d) The multimer positive populations from (c) were sorted by FACS and DNA barcodes associated with the sorted cell population were subjected to qPCR with fluorescent reporter probes targeting each individual DNA barcode. The experiments were performed with reagent mixtures with DNA barcodes inverted between the CMV and HIV pMHC multimers (indicated in orange and purple respectively). Cross threshold (Ct) values of DNA barcodes recovered from qPCR with approximately 200 and 600 cells in separate reactions (derived from staining with 1:1 and 1:1:998 reagent mixtures respectively) are shown in bar plots (mean±range of duplicate samples). Hashtag indicate that a given barcode was not detected.
Supplementary Figure 2 Dynamic range and detection limit using a panel of >1000 DNA-barcoded pMHC multimers
(a) Heatmap representing the pMHC multimer analysis of duplicates of seven samples with various proportions of HLA-B*0702 CMV TPR-specific T cells: 5%, 1%, 0.2%, 0.04%, 0.008%, 0.0016% and 0.00032% of CD8 T cells. The ‘5%’ sample corresponds to 100% BC260 PBMCs. Each sample was screened with a panel of 1031 pMHC multimers, all carrying individual DNA barcodes. The heatmap shows changes in read proportions compared to background levels, as in Figure 2b. Each column represents the reads associated with a given sample. Epitopes are grouped based on their antigen origin. Within each antigen group, rows are sorted based on Log2FC, highest to lowest, compared to baseline samples. Orange-red coloring in the heatmap represents a statistically significant number of DNA barcode reads, FDR < 0.1%, defined as antigen-specific T cell responses. The gray scaling indicates non-significant number of barcode reads, i.e. any antigen-responsive T cells associated with such barcode would be present in numbers too low to discern from background. Duplicate samples are grouped side-by-side, indicated with a. and b. respectively. (b) Magnification of the panel of virus-derived peptides. Rows representing antigen specificities are sorted based on Log2FC and duplicate samples a grouped as in (a). The first row represents the target ’5%’ titrated specificity, B*0702 CMVTPR. The rows below are two T cell responses present in the HLA-B*0702 negative PBMC sample (BC262), followed by at least two lower-frequent responses that are present in the ’5%’ donor (BC260). Dark gray scaling, i.e. barcode reads that that are non-significant but with Log2FC>1, may represent T cell responses just below the detection limit. All pMHC multimers are ‘dextramers’.
Supplementary Figure 3 Comparing the dynamic range and detection limit using panels of 110 and 1031 DNA-barcoded pMHC multimers
(a) Heatmap representing the pMHC multimer analysis of duplicates of seven samples with various proportions of HLA-B*0702 CMVTPR-specific T cells: 5%, 1%, 0.2%, 0.04%, 0.008%, 0.0016% and 0.00032% of CD8 T cells. The ‘5%’ sample corresponds to 100% BC260 PBMCs. Each sample was screened with a panel of 110 pMHC multimers, all carrying individual DNA barcodes. The heatmap is organized as in Figure 2b, each column represents one donor. Duplicate samples are grouped side-by-side, indicated with a. and b. respectively. (b) Magnification of the panel of virus-derived peptides. Rows representing antigen specificities are sorted based on Log2FC and duplicate samples a grouped as in (a). The first row represents the target ’5%’ titrated specificity, B*0702 CMVTPR. The rows below are T cell responses present in the HLA-B*0702 negative PBMC sample (BC262) or lower-frequent responses present in the ‘5%’ donor (BC260). Dark gray scaling, i.e. barcode reads that are non-significant but with Log2FC>1, may represent T cell responses just below the detection limit. (c) Correlations between the frequency of HLA-B*0702 CMVTPR-specific T cells determined by analyzing the same samples using combinatorial fluorescent labeled pMHC multimers or a panel of 110 DNA barcode labeled pMHC multimers. (d, e) Correlation between results obtained from screening the same samples of 2x106 PBMCs with the panel of 1031 or 110 DNA-barcoded pMHC multimers represented as (d) the estimated frequency of HLA-B*0702 CMVTPR-specific T cells or (e) the number of clonally reduced barcode reads associated with this pMHC multimer. Error bars represent range of duplicates (SD). The accumulated number of non-reduced read counts that mapped to any of the DNA barcodes among the 14 samples screened were 2.7x106 and 1.4 x106 reads for the 110 and 1031 pMHC multimer library respectively. All pMHC multimers are ‘dextramers’.
Supplementary Figure 4 T cell reactivity in healthy donor samples analyzed by a panel of 110 DNA-barcoded pMHC multimers
(a) Analysis of PBMCs (1-2x106) from six different healthy donors using 110 pMHC multimers each carrying individual barcodes. The heatmap is organized as in figure 2b, each column represents one donor. Epitopes are grouped based on their antigen origin. Significant responses are shown in orange-red colors. Significance was defined as FDR<5% since the number of reads within a given sample were compared with only one baseline sample (opposed to three baseline samples in other experiments). (b) Magnification of the panel of virus-derived peptides (26 epitopes). Rows representing antigen specificities are grouped according to HLA-type and sorted within each group based on Log2FC. HLA types of donors can be seen in Supplementary Table 6. (c) Correlations between the frequencies of antigen-specific T cells determined by analyzing the same samples with combinatorially encoded fluorescently labeled pMHC multimers or with 110 DNA-barcoded pMHC multimers. Each dot represents one specificity. T cell populations with FDR<5% in DNA-barcode MHC multimer analysis or ≥10 events and >0.002% of CD8 T cells in combinatorial encoding analysis were plotted. All specificities included in the plot were tested using both a combinatorial encoding analysis and DNA-barcoded MHC multimers. Dots plotted on the axes are nonsignificant for one of the methods. (d, e) Correlation between results obtained from screening the same samples with the panel of 1031 or 110 DNA-barcoded pMHC multimers represented as (d) the estimated frequency of antigen-specific T cells or (e) the number of clonally reduced barcode reads associated with the given pMHC multimers. Each dot represents one specificity. Only T cell populations that fulfilled the significance criteria for DNA barcode assessment (FDR<0.1% and 5% for the 1031 and 110 library, respectively) in at least one of the analyses were plotted. Dots plotted on the axes are nonsignificant for one of the library screenings. The accumulated number of non-reduced read counts that mapped to any of the DNA barcodes among the six screened samples were 2.6x105 and 4.3x105 reads for the 110 and 1031 pMHC multimer library respectively.
Supplementary Figure 5 T cell reactivity assessed independently from fluorescent-based separation of MHC multimer binding T cells
(a) PBMCs (2x106) from one healthy donor was stained with varying amounts of DNA barcoded-pMHC multimers, i.e. a titration from 23 nM to 0.0037 nM in respect to each pMHC as indicated above each dot plot. This corresponds to 100%-0.016% of the amount used elsewhere in this study. Samples were stained in duplicates and either the full CD8 population (all cells in the dot plots) or only the MHC multimer positive populations (cells indicated in black) were sorted. (b) Bar plot representing the distribution of DNA barcodes in the isolated cells from (a). Left side is based on the full CD8 population, right side is based on the MHC multimer positive population. Each bar represents the -Log10(p) value in respect to the pMHC associated DNA barcode. Dotted line at y=3 (-Log10(0.001)) represent the threshold of FDR < 0.1%. The donor BC035 is HLA-A*0101 and B*0801 positive and A*0201 negative (see full HLA-type in Supplementary Table 6). BC035 was previously found to carry antigen-specific T cells restricted to HLA-A*0101, CMV pp50VTE (2.2% of CD8 T cells) and CMV pp65YSE (0.6% of CD8 T cells). HLA- A*0201 restricted epitopes functions as negative controls. Irrespectively of sorting the full CD8 or only the multimer positive population the same responses are detected after sequencing of the DNA barcodes. When the MHC multimer binding T cells could not be separated from the full CD8 T cell population based on their fluorescent intensity, i.e. when applying the lowest amount of MHC multimer reagent, the DNA barcodes associated with positive control reagents were still recovered after sorting the full CD8 population indicating that T cell reactivity can be assessed independent on fluorescent separation of the MHC multimer binding T cells.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Supplementary Tables 1, 4 and 6 (PDF 1606 kb)
Supplementary Table 2
Oligo B's (XLSX 11 kb)
Supplementary Table 3
Forward primers with Sample-identification barcodes (SampleID) and Ion Torrent adaptor (A-Key) and reverse primer with Ion Torrent adaptor (P1-Key) (XLSX 11 kb)
Supplementary Table 5
1031 barcode-labeled MHC multimer panel (XLSX 68 kb)
Supplementary Table 7
110 barcode-labeled MHC multimer panel (XLSX 13 kb)
Supplementary Table 8
175 barcode-labeled MHC multimer panel (XLSX 18 kb)
Supplementary Table 9
328 barcode-labeled MHC multimer panel (XLSX 29 kb)
Rights and permissions
About this article
Cite this article
Bentzen, A., Marquard, A., Lyngaa, R. et al. Large-scale detection of antigen-specific T cells using peptide-MHC-I multimers labeled with DNA barcodes. Nat Biotechnol 34, 1037–1045 (2016). https://doi.org/10.1038/nbt.3662
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.3662
This article is cited by
-
Antibody-mediated delivery of viral epitopes to redirect EBV-specific CD8+ T-cell immunity towards cancer cells
Cancer Gene Therapy (2024)
-
Herpes Virus Infections in Kidney Transplant Patients (HINT) – a prospective observational cohort study
BMC Infectious Diseases (2023)
-
Identification of patient-specific CD4+ and CD8+ T cell neoantigens through HLA-unbiased genetic screens
Nature Biotechnology (2023)
-
Rapid TCR:Epitope Ranker (RAPTER): a primary human T cell reactivity screening assay pairing epitope and TCR at single cell resolution
Scientific Reports (2023)
-
Human circulating and tissue-resident memory CD8+ T cells
Nature Immunology (2023)