Abstract
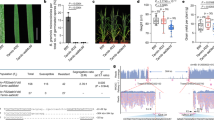
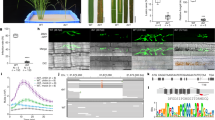
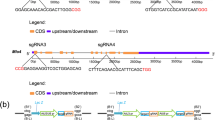
Sequence-specific nucleases have been applied to engineer targeted modifications in polyploid genomes1, but simultaneous modification of multiple homoeoalleles has not been reported. Here we use transcription activator–like effector nuclease (TALEN)2,3 and clustered, regularly interspaced, short palindromic repeats (CRISPR)-Cas9 (refs. 4,5) technologies in hexaploid bread wheat to introduce targeted mutations in the three homoeoalleles that encode MILDEW-RESISTANCE LOCUS (MLO) proteins6. Genetic redundancy has prevented evaluation of whether mutation of all three MLO alleles in bread wheat might confer resistance to powdery mildew, a trait not found in natural populations7. We show that TALEN-induced mutation of all three TaMLO homoeologs in the same plant confers heritable broad-spectrum resistance to powdery mildew. We further use CRISPR-Cas9 technology to generate transgenic wheat plants that carry mutations in the TaMLO-A1 allele. We also demonstrate the feasibility of engineering targeted DNA insertion in bread wheat through nonhomologous end joining of the double-strand breaks caused by TALENs. Our findings provide a methodological framework to improve polyploid crops.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Kim, H. & Kim, J.S. A guide to genome engineering with programmable nucleases. Nat. Rev. Genet. 15, 321–334 (2014).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Moscou, M.J. & Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501 (2009).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Elliott, C. et al. Functional conservation of wheat and rice Mlo orthologs in defense modulation to the powdery mildew fungus. Mol. Plant Microbe Interact. 15, 1069–1077 (2002).
Várallyay, É., Giczey, G. & Burgyán, J. Virus-induced gene silencing of Mlo genes induces powdery mildew resistance in Triticum aestivum. Arch. Virol. 157, 1345–1350 (2012).
Slade, A.J. et al. A reverse genetic, nontransgenic approach to wheat crop improvement by TILLING. Nat. Biotechnol. 23, 75–81 (2005).
Dvorřák, J. in Genetics and Genomics of the Triticeae Vol. 7. (eds. Muehlbauer, G.J. & Feuillet, C.) 685–711 (Springer US, 2009).
Bibikova, M., Beumer, K., Trautman, J.K. & Carroll, D. Enhancing gene targeting with designed zinc finger nucleases. Science 300, 764 (2003).
Symington, L.S. & Gautier, J. Double-strand break end resection and repair pathway choice. Annu. Rev. Genet. 45, 247–271 (2011).
Voytas, D.F. & Gao, C. Precision genome engineering and agriculture: opportunities and regulatory challenges. PLoS Biol. 12, e1001877 (2014).
Zhang, F. et al. High frequency targeted mutagenesis in Arabidopsis thaliana using zinc-finger nucleases. Proc. Natl. Acad. Sci. USA 107, 12028–12033 (2010).
Feng, Z. et al. Multigeneration analysis reveals the inheritance, specificity, and patterns of CRISPR/Cas-induced gene modifications in Arabidopsis. Proc. Natl. Acad. Sci. USA 111, 4632–4637 (2014).
Christian, M., Qi, Y., Zhang, Y. & Voytas, D.F. Targeted mutagenesis of Arabidopsis thaliana using engineered TAL effector nucleases. G3 (Bethesda) 3, 1697–1705 (2013).
Li, T. et al. High-efficiency TALEN-based gene editing produces disease-resistant rice. Nat. Biotechnol. 30, 390–392 (2012).
Shan, Q. et al. Targeted genome modification of crop plants using a CRISPR-Cas system. Nat. Biotechnol. 31, 686–688 (2013).
Büschges, R. et al. The barley Mlo gene: A novel control element of plant pathogen resistance. Cell 88, 695–705 (1997).
Piffanelli, P. et al. A barley cultivation-associated polymorphism conveys resistance to powdery mildew. Nature 430, 887–891 (2004).
Consonni, C. et al. Conserved requirement for a plant host cell protein in powdery mildew pathogenesis. Nat. Genet. 38, 716–720 (2006).
Bai, Y. et al. Naturally occurring broad-spectrum powdery mildew resistance in a central american tomato accession is caused by loss of mlo function. Mol. Plant Microbe Interact. 21, 30–39 (2008).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Shan, Q. et al. Rapid and efficient gene modification in rice and brachypodium using TALENs. Mol. Plant 6, 1365–1368 (2013).
Christensen, A. & Quail, P. Ubiquitin promoter-based vectors for high-level expression of selectable and/or screenable marker genes in monocotyledonous plants. Transgenic Res. 5, 213–218 (1996).
Feng, Z. et al. Efficient genome editing in plants using a CRISPR/Cas system. Cell Res. 23, 1229–1232 (2013).
Xie, K. & Yang, Y. RNA-guided genome editing in plants using a CRISPR-Cas system. Mol. Plant 6, 1975–1983 (2013).
McDonald, B.A. & Linde, C. Pathogen population genetics, evolutionary potential, and durable resistance. Annu. Re. Phytopathol. 40, 349–379 (2002).
Zhang, Y. et al. Transcription activator-like effector nucleases enable efficient plant genome engineering. Plant Physiol. 161, 20–27 (2013).
Guerineau, F., Lucy, A. & Mullineaux, P. Effect of two consensus sequences preceding the translation initiator codon on gene expression in plant protoplasts. Plant Mol. Biol. 18, 815–818 (1992).
Rasco-Gaunt, S. et al. Procedures allowing the transformation of a range of European elite wheat (Triticum aestivum L.) varieties via particle bombardment. J. Exp. Bot. 52, 865–874 (2001).
Hein, I. et al. Virus-induced gene silencing-based functional characterization of genes associated with powdery mildew resistance in barley. Plant Physiol. 138, 2155–2164 (2005).
Acknowledgements
We thank D. Wang for critical reading of the manuscript and providing the E22 and B13 strains, Q. Shen for providing E09 strain, and T. Li for technical support in flow cytometry (all three are at the Institute of Genetics and Developmental Biology, Chinese Academy of Sciences). This work was supported by grants from the Strategic Priority Research Program of the Chinese Academy of Sciences (XDB11030500) and the National Basic Research Program of China (973 Program, 2011CB100702) to J.-L.Q. and a grant to C.G. from the National Natural Science Foundation of China (31271795).
Author information
Authors and Affiliations
Contributions
J.-L.Q., C.G. and Y.W. designed the experiments; Y.W., X.C., Q.S., Y.Z. and J.L. performed the experiments; and J.-L.Q., C.G. and Y.W. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors have filed a patent application (Chinese patent application number 201410027631.2) based on the results reported in this paper.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7 and Supplementary Tables 1–6 (PDF 17919 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Cheng, X., Shan, Q. et al. Simultaneous editing of three homoeoalleles in hexaploid bread wheat confers heritable resistance to powdery mildew. Nat Biotechnol 32, 947–951 (2014). https://doi.org/10.1038/nbt.2969
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2969
This article is cited by
-
Precise integration of large DNA sequences in plant genomes using PrimeRoot editors
Nature Biotechnology (2024)
-
Targeted gene knockout via CRISPR/Cas9: precise genome editing in eggplant (Solanum melongena) through phytoene desaturase gene disruption
Journal of Crop Science and Biotechnology (2024)
-
Bioinformatic and literature assessment of toxicity and allergenicity of a CRISPR-Cas9 engineered gene drive to control Anopheles gambiae the mosquito vector of human malaria
Malaria Journal (2023)
-
Plant immunity research in China
Phytopathology Research (2023)
-
Functional and evolutionary study of MLO gene family in the regulation of Sclerotinia stem rot resistance in Brassica napus L.
Biotechnology for Biofuels and Bioproducts (2023)