Abstract
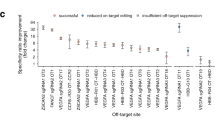
Genome editing by Cas9, which cleaves double-stranded DNA at a sequence programmed by a short single-guide RNA (sgRNA), can result in off-target DNA modification that may be detrimental in some applications. To improve DNA cleavage specificity, we generated fusions of catalytically inactive Cas9 and FokI nuclease (fCas9). DNA cleavage by fCas9 requires association of two fCas9 monomers that simultaneously bind target sites ∼15 or 25 base pairs apart. In human cells, fCas9 modified target DNA sites with >140-fold higher specificity than wild-type Cas9 and with an efficiency similar to that of paired Cas9 'nickases', recently engineered variants that cleave only one DNA strand per monomer. The specificity of fCas9 was at least fourfold higher than that of paired nickases at loci with highly similar off-target sites. Target sites that conform to the substrate requirements of fCas9 occur on average every 34 bp in the human genome, suggesting the versatility of this approach for highly specific genome-wide editing.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014).
Perez, E.E. et al. Establishment of HIV-1 resistance in CD4+ T cells by genome editing using zinc-finger nucleases. Nat. Biotechnol. 26, 808–816 (2008).
Mali, P. et al. RNA-guided human genome engineering via Cas9. Science 339, 823–826 (2013).
Fu, Y., Sander, J.D., Reyon, D., Cascio, V.M. & Joung, J.K. Improving CRISPR-Cas nuclease specificity using truncated guide RNAs. Nat. Biotechnol. 32, 279–284 (2014).
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012).
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013).
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013).
Pattanayak, V. et al. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 31, 839–843 (2013).
Fu, Y. et al. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 31, 822–826 (2013).
Hsu, P.D. et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 31, 827–832 (2013).
Cradick, T.J., Fine, E.J., Antico, C.J. & Bao, G. CRISPR/Cas9 systems targeting β-globin and CCR5 genes have substantial off-target activity. Nucleic Acids Res. 41, 9584–9592 (2013).
Cho, S.W. et al. Analysis of off-target effects of CRISPR/Cas-derived RNA-guided endonucleases and nickases. Genome Res. 24, 132–141 (2014).
Mali, P. et al. CAS9 transcriptional activators for target specificity screening and paired nickases for cooperative genome engineering. Nat. Biotechnol. 31, 833–838 (2013).
Ran, F.A. et al. Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154, 1380–1389 (2013).
Gasiunas, G., Barrangou, R., Horvath, P. & Siksnys, V. Cas9-crRNA ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. USA 109, E2579–E2586 (2012).
Ramirez, C.L. et al. Engineered zinc finger nickases induce homology-directed repair with reduced mutagenic effects. Nucleic Acids Res. 40, 5560–5568 (2012).
Wang, J. et al. Targeted gene addition to a predetermined site in the human genome using a ZFN-based nicking enzyme. Genome Res. 22, 1316–1326 (2012).
Vanamee, É.S., Santagata, S. & Aggarwal, A.K. FokI requires two specific DNA sites for cleavage. J. Mol. Biol. 309, 69–78 (2001).
Pattanayak, V., Ramirez, C.L., Joung, J.K. & Liu, D.R. Revealing off-target cleavage specificities of zinc-finger nucleases by in vitro selection. Nat. Methods 8, 765–770 (2011).
Guilinger, J.P. et al. Broad specificity profiling of TALENs results in engineered nucleases with improved DNA-cleavage specificity. Nat. Methods 11, 429–435 (2014).
Nishimasu, H. et al. Crystal structure of Cas9 in complex with guide RNA and target DNA. Cell 156, 935–949 (2014).
Jinek, M. et al. Structures of Cas9 endonucleases reveal RNA-mediated conformational activation. Science 31, 6176 (2014).
Schellenberger, V. et al. A recombinant polypeptide extends the in vivo half-life of peptides and proteins in a tunable manner. Nat. Biotechnol. 27, 1186–1190 (2009).
Qi, L.S. et al. Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152, 1173–1183 (2013).
Esvelt, K.M. et al. Orthogonal Cas9 proteins for RNA-guided gene regulation and editing. Nat. Methods 10, 1116–1121 (2013).
Kim, Y., Kweon, J. & Kim, J.-S. TALENs and ZFNs are associated with different mutation signatures. Nat. Methods 10, 185–185 (2013).
Wu, X. et al. Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat. Biotechnol. doi:10.1038/nbt.2889 (20 April 2014).
Doyon, Y. et al. Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures. Nat. Methods 8, 74–79 (2011).
Shcherbakova, D.M. & Verkhusha, V.V. Near-infrared fluorescent proteins for multicolor in vivo imaging. Nat. Methods 10, 751–754 (2013).
Schneider, C.A., Rasband, W.S. & Eliceiri, K.W. NIH Image to ImageJ: 25 years of image analysis. Nat. Methods 9, 671–675 (2012).
Guschin, D.Y. et al. in Engineering Zinc Finger Proteins (eds. Mackay, J.P. & Segal, D.J.) 247–256 (Humana Press, 2010).
Sander, J.D. et al. In silico abstraction of zinc finger nuclease cleavage profiles reveals an expanded landscape of off-target sites. Nucleic Acids Res. 41, e181 (2013).
Acknowledgements
J.P.G., D.B.T. and D.R.L. were supported by Defense Advanced Research Projects Agency HR0011-11-2-0003 and N66001-12-C-4207, US National Institutes of Health NIGMS R01 GM095501 (D.R.L.) and the Howard Hughes Medical Institute (HHMI). D.R.L. was supported as a HHMI Investigator. We thank R. McDonald for technical assistance and V. Pattanayak for helpful comments.
Author information
Authors and Affiliations
Contributions
J.P.G. and D.B.T. performed the experiments, designed the research, analyzed the data and wrote the manuscript. D.R.L. designed the research, analyzed the data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The co-authors have filed a provisional patent application related to this work. D.R.L. is a consultant for Editas Medicine, a company that applies genome editing technologies for human therapeutic applications.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–11, Supplementary Results, Supplementary Notes and Supplementary Tables 1–5 (PDF 1127 kb)
Rights and permissions
About this article
Cite this article
Guilinger, J., Thompson, D. & Liu, D. Fusion of catalytically inactive Cas9 to FokI nuclease improves the specificity of genome modification. Nat Biotechnol 32, 577–582 (2014). https://doi.org/10.1038/nbt.2909
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2909
This article is cited by
-
Undetectable off-target effects induced by FokI catalytic domain in mouse embryos
Genome Biology (2024)
-
Site-specific transgene integration in chimeric antigen receptor (CAR) T cell therapies
Biomarker Research (2023)
-
Treatment of monogenic and digenic dominant genetic hearing loss by CRISPR-Cas9 ribonucleoprotein delivery in vivo
Nature Communications (2023)
-
A cleavage rule for selection of increased-fidelity SpCas9 variants with high efficiency and no detectable off-targets
Nature Communications (2023)
-
Multi-faceted CRISPR/Cas technological innovation aspects in the framework of 3P medicine
EPMA Journal (2023)