Abstract
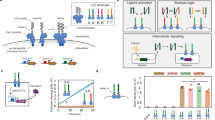
The design of synthetic biology–inspired control devices enabling entire mammalian cells to receive, process and transfer metabolic information and so communicate with each other via synthetic multichannel networks may provide new insight into the organization of multicellular organisms and future clinical interventions1. Here we describe communication networks that orchestrate behavior in individual mammalian cells in response to cell-to-cell metabolic signals. We engineered sender, processor and receiver cells that interact with each other in ways that resemble natural intercellular communication networks such as multistep information processing cascades, feed-forward–based signaling loops, and two-way communication. The engineered two-way communication devices mimicking natural control systems in the development of vertebrate extremities2 and vasculature3,4,5 was used to program temporal permeability in vascular endothelial cell layers. These synthetic multicellular communication systems may inspire future therapies or tissue engineering strategies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weber, W., Daoud-El Baba, M. & Fussenegger, M. Synthetic ecosystems based on airborne inter- and intrakingdom communication. Proc. Natl. Acad. Sci. USA 104, 10435–10440 (2007).
Freeman, M. Feedback control of intercellular signalling in development. Nature 408, 313–319 (2000).
Thurston, G. et al. Leakage-resistant blood vessels in mice transgenically overexpressing angiopoietin-1. Science 286, 2511–2514 (1999).
Thurston, G. Complementary actions of VEGF and angiopoietin-1 on blood vessel growth and leakage. J. Anat. 200, 575–580 (2002).
Lee, S.W., Kim, W.J., Jun, H.O., Choi, Y.K. & Kim, K.W. Angiopoietin-1 reduces vascular endothelial growth factor-induced brain endothelial permeability via upregulation of ZO-2. Int. J. Mol. Med. 23, 279–284 (2009).
Guido, N.J. et al. A bottom-up approach to gene regulation. Nature 439, 856–860 (2006).
Gardner, T.S., Cantor, C.R. & Collins, J.J. Construction of a genetic toggle switch in Escherichia coli. Nature 403, 339–342 (2000).
Kramer, B.P. et al. An engineered epigenetic transgene switch in mammalian cells. Nat. Biotechnol. 22, 867–870 (2004).
Auslander, S., Auslander, D., Muller, M., Wieland, M. & Fussenegger, M. Programmable single-cell mammalian biocomputers. Nature 487, 123–127 (2012).
Danino, T., Mondragon-Palomino, O., Tsimring, L. & Hasty, J. A synchronized quorum of genetic clocks. Nature 463, 326–330 (2010).
Weber, W. et al. A synthetic time-delay circuit in mammalian cells and mice. Proc. Natl. Acad. Sci. USA 104, 2643–2648 (2007).
Friedland, A.E. et al. Synthetic gene networks that count. Science 324, 1199–1202 (2009).
Tigges, M., Marquez-Lago, T.T., Stelling, J. & Fussenegger, M. A tunable synthetic mammalian oscillator. Nature 457, 309–312 (2009).
Weber, W. et al. A synthetic mammalian gene circuit reveals antituberculosis compounds. Proc. Natl. Acad. Sci. USA 105, 9994–9998 (2008).
Weber, W. & Fussenegger, M. Emerging biomedical applications of synthetic biology. Nat. Rev. Genet. 13, 21–35 (2012).
Kemmer, C. et al. Self-sufficient control of urate homeostasis in mice by a synthetic circuit. Nat. Biotechnol. 28, 355–360 (2010).
Ye, H., Daoud-El Baba, M., Peng, R.W. & Fussenegger, M. A synthetic optogenetic transcription device enhances blood-glucose homeostasis in mice. Science 332, 1565–1568 (2011).
Miki, T. et al. ATP-sensitive K+ channels in the hypothalamus are essential for the maintenance of glucose homeostasis. Nat. Neurosci. 4, 507–512 (2001).
Yarilina, A., Park-Min, K.H., Antoniv, T., Hu, X. & Ivashkiv, L.B. TNF activates an IRF1-dependent autocrine loop leading to sustained expression of chemokines and STAT1-dependent type I interferon-response genes. Nat. Immunol. 9, 378–387 (2008).
Tamsir, A., Tabor, J.J. & Voigt, C.A. Robust multicellular computing using genetically encoded NOR gates and chemical 'wires'. Nature 469, 212–215 (2011).
Regot, S. et al. Distributed biological computation with multicellular engineered networks. Nature 469, 207–211 (2011).
Chen, M.T. & Weiss, R. Artificial cell-cell communication in yeast Saccharomyces cerevisiae using signaling elements from Arabidopsis thaliana. Nat. Biotechnol. 23, 1551–1555 (2005).
Shou, W., Ram, S. & Vilar, J.M. Synthetic cooperation in engineered yeast populations. Proc. Natl. Acad. Sci. USA 104, 1877–1882 (2007).
Basu, S., Gerchman, Y., Collins, C.H., Arnold, F.H. & Weiss, R. A synthetic multicellular system for programmed pattern formation. Nature 434, 1130–1134 (2005).
Basu, S., Mehreja, R., Thiberge, S., Chen, M.T. & Weiss, R. Spatiotemporal control of gene expression with pulse-generating networks. Proc. Natl. Acad. Sci. USA 101, 6355–6360 (2004).
Hartman, S.C. & Mulligan, R.C. Two dominant-acting selectable markers for gene transfer studies in mammalian cells. Proc. Natl. Acad. Sci. USA 85, 8047–8051 (1988).
Akers, J.C. & Tan, M. Molecular mechanism of tryptophan-dependent transcriptional regulation in Chlamydia trachomatis. J. Bacteriol. 188, 4236–4243 (2006).
Weber, W. et al. Gas-inducible transgene expression in mammalian cells and mice. Nat. Biotechnol. 22, 1440–1444 (2004).
Schlatter, S., Rimann, M., Kelm, J. & Fussenegger, M. SAMY, a novel mammalian reporter gene derived from Bacillus stearothermophilus alpha-amylase. Gene 282, 19–31 (2002).
Lee, S.W. et al. SSeCKS regulates angiogenesis and tight junction formation in blood-brain barrier. Nat. Med. 9, 900–906 (2003).
Acknowledgements
We thank M. Tan (pMT1182), M. Ehrbar (pVEGF165pcDNA3), G.Y. Koh (pAng-1) and K. Frei (EA.hy926 cells) for providing reagents and D. Grasso for critical comments on the manuscript. This work was supported by the Swiss National Science Foundation (grant no. 31003A0-126022) and in part by the EC Framework 7 (Persist). Work of W.W. is supported by the Excellence Initiative of the German Federal and State Governments (EXC 294).
Author information
Authors and Affiliations
Contributions
W.B., M.L., W.W., J.S. and M.F. designed the project, analyzed results and wrote the manuscript. W.B. and M.D.E.-B. performed the experimental work. M.L. and J.S. designed and analyzed the mathematical model and performed the simulations.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Model, Supplementary Figures 1–19, Supplementary Table 1 and Supplementary Assembly (PDF 1058 kb)
Rights and permissions
About this article
Cite this article
Bacchus, W., Lang, M., El-Baba, M. et al. Synthetic two-way communication between mammalian cells. Nat Biotechnol 30, 991–996 (2012). https://doi.org/10.1038/nbt.2351
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2351
This article is cited by
-
Combinatorial protein dimerization enables precise multi-input synthetic computations
Nature Chemical Biology (2023)
-
Microbial methionine transporters and biotechnological applications
Applied Microbiology and Biotechnology (2021)
-
Functional and morphological adaptation in DNA protocells via signal processing prompted by artificial metalloenzymes
Nature Nanotechnology (2020)
-
Intercellular communication between artificial cells by allosteric amplification of a molecular signal
Nature Communications (2020)
-
De novo design of an intercellular signaling toolbox for multi-channel cell–cell communication and biological computation
Nature Communications (2020)