Abstract
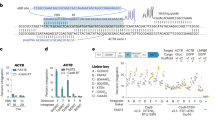
Engineered transcription activator–like effector nucleases (TALENs) have shown promise as facile and broadly applicable genome editing tools. However, no publicly available high-throughput method for constructing TALENs has been published, and large-scale assessments of the success rate and targeting range of the technology remain lacking. Here we describe the fast ligation-based automatable solid-phase high-throughput (FLASH) system, a rapid and cost-effective method for large-scale assembly of TALENs. We tested 48 FLASH-assembled TALEN pairs in a human cell–based EGFP reporter system and found that all 48 possessed efficient gene-modification activities. We also used FLASH to assemble TALENs for 96 endogenous human genes implicated in cancer and/or epigenetic regulation and found that 84 pairs were able to efficiently introduce targeted alterations. Our results establish the robustness of TALEN technology and demonstrate that FLASH facilitates high-throughput genome editing at a scale not currently possible with other genome modification technologies.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bogdanove, A.J. & Voytas, D.F. TAL effectors: customizable proteins for DNA targeting. Science 333, 1843–1846 (2011).
Bogdanove, A.J., Schornack, S. & Lahaye, T. TAL effectors: finding plant genes for disease and defense. Curr. Opin. Plant Biol. 13, 394–401 (2010).
Scholze, H. & Boch, J. TAL effectors are remote controls for gene activation. Curr. Opin. Microbiol. 47–53 (2011).
Boch, J. et al. Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326, 1509–1512 (2009).
Moscou, M.J. & Bogdanove, A.J. A simple cipher governs DNA recognition by TAL effectors. Science 326, 1501 (2009).
Miller, J.C. et al. A TALE nuclease architecture for efficient genome editing. Nat. Biotechnol. 29, 143–148 (2011).
Morbitzer, R., Romer, P., Boch, J. & Lahaye, T. Regulation of selected genome loci using de novo-engineered transcription activator-like effector (TALE)-type transcription factors. Proc. Natl. Acad. Sci. USA 107, 21617–21622 (2010).
Morbitzer, R., Elsaesser, J., Hausner, J. & Lahaye, T. Assembly of custom TALE-type DNA binding domains by modular cloning. Nucleic Acids Res. 39, 5790–5799 (2011).
Zhang, F. et al. Efficient construction of sequence-specific TAL effectors for modulating mammalian transcription. Nat. Biotechnol. 29, 149–153 (2011).
Geissler, R. et al. Transcriptional activators of human genes with programmable DNA-specificity. PLoS ONE 6, e19509 (2011).
Weber, E., Gruetzner, R., Werner, S., Engler, C. & Marillonnet, S. Assembly of designer TAL effectors by Golden Gate cloning. PLoS ONE 6, e19722 (2011).
Christian, M. et al. Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186, 757–761 (2010).
Li, T. et al. TAL nucleases (TALNs): hybrid proteins composed of TAL effectors and FokI DNA-cleavage domain. Nucleic Acids Res. 39, 359–372 (2011).
Mahfouz, M.M. et al. De novo-engineered transcription activator-like effector (TALE) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc. Natl. Acad. Sci. USA 108, 2623–2628 (2011).
Mussolino, C. et al. A novel TALE nuclease scaffold enables high genome editing activity in combination with low toxicity. Nucleic Acids Res. 9283–9293 (2011).
Li, T. et al. Modularly assembled designer TAL effector nucleases for targeted gene knockout and gene replacement in eukaryotes. Nucleic Acids Res. 39, 6315–6325 (2011).
Cermak, T. et al. Efficient design and assembly of custom TALEN and other TAL effector-based constructs for DNA targeting. Nucleic Acids Res. 39, e82 (2011).
Wood, A.J. et al. Targeted genome editing across species using ZFNs and TALENs. Science 333, 307 (2011).
Hockemeyer, D. et al. Genetic engineering of human pluripotent cells using TALE nucleases. Nat. Biotechnol. 29, 731–734 (2011).
Tesson, L. et al. Knockout rats generated by embryo microinjection of TALENs. Nat. Biotechnol. 29, 695–696 (2011).
Sander, J.D. et al. Targeted gene disruption in somatic zebrafish cells using engineered TALENs. Nat. Biotechnol. 29, 697–698 (2011).
Huang, P. et al. Heritable gene targeting in zebrafish using customized TALENs. Nat. Biotechnol. 29, 699–700 (2011).
Maeder, M.L. et al. Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 31, 294–301 (2008).
Vogelstein, B. & Kinzler, K.W. Cancer genes and the pathways they control. Nat. Med. 10, 789–799 (2004).
Kim, H.J., Lee, H.J., Kim, H., Cho, S.W. & Kim, J.S. Targeted genome editing in human cells with zinc finger nucleases constructed via modular assembly. Genome Res. 19, 1279–1288 (2009).
Pattanayak, V., Ramirez, C.L., Joung, J.K. & Liu, D.R. Revealing off-target cleavage specificities of zinc-finger nucleases by in vitro selection. Nat. Methods 8, 765–770 (2011).
Gabriel, R. et al. An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nat. Biotechnol. 29, 816–823 (2011).
Yusa, K. et al. Targeted gene correction of α1-antitrypsin deficiency in induced pluripotent stem cells. Nature 478, 391–394 (2011).
Miller, J.C. et al. An improved zinc-finger nuclease architecture for highly specific genome editing. Nat. Biotechnol. 25, 778–785 (2007).
Doyon, Y. et al. Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures. Nat. Methods 8, 74–79 (2011).
Szczepek, M. et al. Structure-based redesign of the dimerization interface reduces the toxicity of zinc-finger nucleases. Nat. Biotechnol. 25, 786–793 (2007).
Guschin, D.Y. et al. A rapid and general assay for monitoring endogenous gene modification. Methods Mol. Biol. 649, 247–256 (2010).
Acknowledgements
We thank T. Cathomen for providing polyclonal U2OS-EGFP reporter cells and a T7EI protocol, Y. Fu for deriving the clonal U2OS-EGFP cell line, D. Dobbs for support and encouragement, and M. Maeder, J. Angstman and C. Ramirez for helpful comments. This work was supported by a US National Institutes of Health (NIH) Director's Pioneer Award DP1 OD006862 (J.K.J.), NIH P50HG005550 (J.K.J.), the Jim and Ann Orr MGH Research Scholar Award (J.K.J.), NIH T32 CA009216 (J.D.S.) and National Science Foundation DBI-0923827 (D.R.).
Author information
Authors and Affiliations
Contributions
J.D.S. and J.K.J. conceived of the FLASH method. D.R., S.Q.T., C.K., J.A.F. and J.D.S. performed experiments. D.R., S.Q.T., C.K., J.A.F., J.D.S. and J.K.J. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
J.D.S. and J.K.J. are inventors on a patent application describing the FLASH method. J.K.J. is a member of the Scientific Advisory Board of Transposagen Biopharmaceuticals, Inc.
Supplementary information
Supplementary Text and Figures
Supplementary Methods, Supplementary Discussion, Supplementary Tables and Supplementary Figures 1–7 (PDF 6093 kb)
Supplementary Table 1
DNA sequences encoding individual TALE repeats used to construct the archive of preassembled units required for FLASH assembly (XLSX 9 kb)
Supplementary Table 2
Archive of 376 plasmids encoding pre-assembled TALE repeat units required to practice FLASH assembly (XLSX 48 kb)
Supplementary Table 3
EGFP reporter gene sequences targeted by 48 pairs of FLASH TALENs (XLSX 15 kb)
Supplementary Table 4
Endogenous human gene sequences targeted by 96 pairs of FLASH TALENs (XLSX 17 kb)
Supplementary Table 5
PCR primers and conditions used to amplify TALEN-targeted sequences of 96 endogenous human genes (XLSX 16 kb)
Rights and permissions
About this article
Cite this article
Reyon, D., Tsai, S., Khayter, C. et al. FLASH assembly of TALENs for high-throughput genome editing. Nat Biotechnol 30, 460–465 (2012). https://doi.org/10.1038/nbt.2170
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2170
This article is cited by
-
Oral Administration of the Antimicrobial Peptide Mastoparan X Alleviates Enterohemorrhagic Escherichia coli–Induced Intestinal Inflammation and Regulates the Gut Microbiota
Probiotics and Antimicrobial Proteins (2024)
-
Correlation of gut microorganisms and non-volatile flavor substances provides new insights for breeding Scylla paramamosain
Journal of Oceanology and Limnology (2024)
-
Correlation study on microbial communities and volatile flavor compounds in cigar tobacco leaves of diverse origins
Applied Microbiology and Biotechnology (2024)
-
Integrated Microbiome and Metabolomic Analysis Reveal Responses of Rhizosphere Bacterial Communities and Root exudate Composition to Drought and Genotype in Rice (Oryza sativa L.)
Rice (2023)
-
Impact of influenza virus infection on lung microbiome in adults with severe pneumonia
Annals of Clinical Microbiology and Antimicrobials (2023)