Abstract
RNA editing is a post-transcriptional event that recodes hereditary information. Here we describe a comprehensive profile of the RNA editome of a male Han Chinese individual based on analysis of ∼767 million sequencing reads from poly(A)+, poly(A)− and small RNA samples. We developed a computational pipeline that carefully controls for false positives while calling RNA editing events from genome and whole-transcriptome data of the same individual. We identified 22,688 RNA editing events in noncoding genes and introns, untranslated regions and coding sequences of protein-coding genes. Most changes (∼93%) converted A to I(G), consistent with known editing mechanisms based on adenosine deaminase acting on RNA (ADAR). We also found evidence of other types of nucleotide changes; however, these were validated at lower rates. We found 44 editing sites in microRNAs (miRNAs), suggesting a potential link between RNA editing and miRNA-mediated regulation. Our approach facilitates large-scale studies to profile and compare editomes across a wide range of samples.
Similar content being viewed by others
Main
RNA editing is an integral step in generating the diversity and plasticity of cellular RNA signatures. Most editing events convert A to I(G) (adenosine to inosine, which is translated as guanosine), and are catalyzed by the double-stranded RNA-specific ADAR family of proteins. On the basis of its overabundance in repetitive Alu elements and the brain transcriptome1,2,3, RNA editing has been viewed as a key determinant in primate evolution and the development of higher brain functions4. Many outstanding questions on the extent and consequences of RNA editing in humans remain unanswered, despite extensive documentation of edited sites through bioinformatics approaches5,6,7,8,9 and the reported roles of editing in altering genetic messages and other post-transcriptional events such as RNA splicing and miRNA regulation2,10,11,12. Global and unequivocal identification of RNA editing targets represents a critical first step in further understanding this post-transcriptional modification. This calls for complete information on whole-genome and transcriptome sequences from the same individual, so as to exclude polymorphisms and mutations among populations, as well as experimental approaches with the necessary high-throughput sequencing and base resolution13,14. Whole-transcriptome deep-sequencing technologies (e.g., RNA-Seq)15,16,17, with their capacity to simultaneously assay the entire transcriptome, represent an excellent choice of tool in this regard. Recent studies reporting the use of target-specific RNA-Seq4,18, the combination of DNA capture and parallel sequencing19, and mRNA-Seq20,21 to find human RNA editing sites attest to the notion that this strategy is advantageous in addressing many outstanding questions of the editing phenomenon and its implications on the transcriptome.
In this study, we used RNA-Seq to identify post-transcriptional editing events. Our unbiased and in-depth approach revealed many editing sites in transcripts corresponding to coding, noncoding and small RNA genes. Some editing events are not of the well-known A→I(G) type. This study, to our knowledge, represents the most extensive analysis of RNA editing in a single individual (22,688 sites). These findings hint at a widespread occurrence of transcript variation at the single-nucleotide level in the human transcriptome and underscore the importance of complete delineation of these polymorphisms to advance our understanding of human development and normal and pathophysiological conditions.
Results
Whole-transcriptome deep sequencing
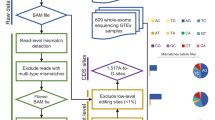
To exhaustively analyze a human RNA editome, we first obtained whole-transcriptome data from a lymphoblastoid cell line of a male Han Chinese individual (YH), whose genome sequence was reported previously22. We fractionated total RNA into poly(A)+ RNA, poly(A)− RNA and small RNA species and constructed whole-transcriptome sequencing libraries (Fig. 1a). Because poly(A)− transcripts have been observed to be enriched for noncoding RNA23 and are less well annotated in databases, we performed strand-specific sequencing on the poly(A)− RNA libraries (Online Methods). We generated 75 bp and 100 bp RNA-Seq reads for the poly(A)+ libraries and 90 bp reads for poly(A)− libraries. In total, 767.58 million reads could be uniquely aligned, representing ∼67.32 Gbp of expressed sequences (Supplementary Table 1). The aligned reads provided substantial coverage (≥90%) for the vast majority of the identified mRNA transcripts (Supplementary Fig. 1a and Supplementary Table 2); and they provided considerable sequencing depth (Supplementary Fig. 1b) and coverage (Table 1 and Supplementary Table 3). Such deep sampling of the total expressed sequences in human cells provides an ideal data set for definitive and quantitative characterization of the RNA editome.
(a) Schematic depiction of the experimental design of the study. Total RNA was isolated from the lymphoblastoid cell line (LCL) derived from a male Han Chinese individual (YH) and further processed into three different libraries for high-throughput whole-transcriptome sequencing. (b) Overview of algorithm for calling RNA editing sites or RNA-centric SNVs. The pipeline takes raw sequencing reads as input, filters them on the basis of several stringent criteria and outputs the inferred variants that are to be analyzed further. (c) Accuracy and sensitivity of the pipeline for each given filter stage. As successive filters were applied to simulated reads (harboring A→G variants at known positions categorized in DARNED; see Methods), the performance of the approach was evaluated. Accuracy is defined as the false discovery rate (FDR; dotted lines). Sensitivity (SN; gray bars) equals positive calling rate of the simulated editing sites. Notably, the pipeline yielded candidates at a high sensitivity while significantly eliminating the false positives. (d) Validation of inferred editing sites from RNA-Seq by Sanger sequencing. Sequencing chromatogram traces from two exemplary gene loci, CLEC2D and PLEKHA9, are shown. The editing positions (located in the intron of CLEC2D and coding sequence of PLEKHA9) are highlighted by yellow shading. Note the clustering of editing sites in the CLEC2D transcript. Top trace is genomic DNA (gDNA), bottom trace cDNA.
Data analysis pipeline for identification of editing sites
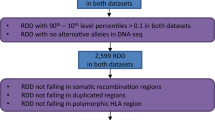
We devised a bioinformatics analysis scheme that implements multiple filters with stringent thresholds to identify sites in RNA transcripts that were different from the corresponding genome sequence (Fig. 1b). In contrast to previous large-scale in silico studies, our global assessment of base changes was not biased toward A→G changes. We performed initial variant calling using a strategy based on algorithms implemented in the SOAPsnp software package, which was originally designed to identify genomic single-nucleotide polymorphisms (SNPs). We applied several adjustments and additional filters in the SNP-calling algorithm to facilitate the detection of bona fide editing or base substitution events in the RNA-Seq reads. Briefly, we first optimized parameters, such as sequence quality score, distance of a potential single-nucleotide variant (SNV) to the end of the supporting read and number of SNV-supporting reads, on the basis of their effects on pipeline performance on simulated data sets (Supplementary Table 4 and Supplementary Fig. 2). Second, to distinguish RNA editing from allele-specific expression and duplication polymorphisms, we only kept sites whose corresponding genome sequences were homozygous in genotype and nonvariant in copy number. Third, to address the problem of intrinsic mapping errors, of which paralogs represent a major source24, we removed SNVs whose supporting reads were misaligned owing to mapping bias inherent to the mapping algorithm; we call these SNVs the mapping errors set (MES). We also used a strand bias filter to remove potential strand-specific errors in sequences generated by the Illumina platform. Additionally, as another means to reduce false positives stemming from paralogous sequences, we used BLAT to search for and subsequently filter SNVs found in such highly similar regions. Remaining sites were filtered against known SNPs to eliminate germline variants, and then we eliminated SNVs for which more than two types of nucleotide sequence were found as these are likely false positives ('multiple-mismatches (nonbinary) type,' Online Methods). Finally, taking into consideration the intrinsic mapping error of the high-throughput sequence data, we excluded polymorphic sites with an extreme degree of variation (100%).
To evaluate our analytic workflow, we simulated two sets of 75-bp paired-end reads from chromosome 1 of the NCBI human RefSeq. These reads contained either random substitutions (first set) or A→G substitution at positions found in the DARNED database9 (second set), which contains known A→G sites in the human transcriptome reported by previous studies. We found that our method achieved a high level of performance in both accuracy and sensitivity. Applying the method to the first simulation set identified >80% of the simulated sites at ∼99% specificity (at 50× depth of simulated sequencing; Supplementary Fig. 2a,b and Supplementary Table 4), and applying it to the second simulation set identified ∼70% of the simulated sites at ∼90% specificity (Fig. 1c and Supplementary Table 5). Our approach identified a sizable portion (∼71%) of the edits found in clusters (≥3 sites in 100 bp), which is a distinct hallmark of A→I(G) editing. This demonstrated the applicability of our pipeline in calling known RNA editing sites, as ∼82% of the DARNED sites in the simulated data were in clusters. Moreover, the simulation results showed that the successive application of more filters progressively improved the specificity of identifying editing sites (Fig. 1c), underscoring the importance of controlling for multiple sources of error in the comparative search of sequence polymorphisms.
We next applied the pipeline to our poly(A)+ and poly(A)− RNA-Seq data. From the poly(A)+ RNA-Seq reads, we identified 16,905 RNA editing sites. To ascertain the editing type for these sites, which requires the strand orientation to be annotated, we cross-referenced data against known coding gene models in RefSeq. This step revealed that ∼30% of the identified sites were located in sequences that were either unannotated in the database (5,381) or corresponded to overlapping transcript units on both strands (57). The remaining 11,467 sites were unambiguously mapped to known gene models, and thus were selected for further analysis (Supplementary Table 6). To identify editing sites in the intergenic regions of the transcriptome, we applied our pipeline on the strand-specific sequencing reads from the poly(A)− RNA library. Overall, 11,221 RNA editing sites were inferred from this approach (Supplementary Table 6 and Supplementary Discussion). In total, we identified 22,688 editing sites with editing type information. Among these sites, 21,113 were of the A→G type. Of the non-A→G sites, 1,146 were transitions and 429 were transversions. This data set and the corresponding RNA-Seq data are available through an online browser database (http://yheditome.genomics.cn/mgb2/gbrowse).
To experimentally validate our calls, we verified a subset of the inferred sites by two replicates of PCR amplification and Sanger sequencing of both DNA and RNA from the same batch of cells from the YH cell line. With regard to previously undescribed editing in coding sequence, we validated candidate genes such as SON-1, PLEKHA9 and UTP14C. An example of a validated editing target, CLEC2D, which exhibited confirmed extensive transcript alteration at sites in the intron, is shown in Figure 1d (left). Notably, CLEC2D mRNA undergoes sequence substitution at multiple sites within a short span of sequence, representing an example of a multiply edited gene. We identified a total of 208 genes of this type, each with more than ten editing sites (Supplementary Table 6). To verify the existence of editing sites with a low degree of variation (<20%), we carried out TA cloning for six of these A→G amplicons, followed by Sanger sequencing. We detected edited sequences in the individually sequenced clones from all of these sites; notably, sites with ∼5% of editing were all confirmed. Overall, of the 74 A→G targets chosen from the data set (distributed in 21 distinct PCR amplicons), 69 sites (or 19 regions) showed clear signals for sequence variation (Supplementary Tables 7 and 8). These validation results yielded a false-discovery rate of 6.74% (5/74), or ∼9.52% (2/21) at the amplicon level, for A→G sites (Supplementary Table 8 and Supplementary Fig. 3), the predominant type of editing identified in our analysis.
Our extensive characterization of the human RNA editome also uncovered an intriguing but mostly overlooked characteristic of the transcriptome, the existence of SNVs that are of the non-A→G substitution type (Fig. 2a and Supplementary Table 6). These nucleotide variants constitute a small but nonnegligible fraction of our data set. Among the 11 possible base substitutions, three types (T→C, G→A and C→T) show sizable representation and together account for 2/3 of these noncanonical events (Fig. 2a). However, validation by Sanger sequencing demonstrated that only a subset of them might represent bona fide nucleotide variants in the RNAs (Fig. 1d, Supplementary Tables 7,8 and Supplementary Fig. 3), as the false-discovery rate for the non-A→G type was ∼49%. Although we also verified about a dozen candidate sites of the transversion type (Fig. 1d, Supplementary Table 8 and Supplementary Fig. 3), given the low percentage of the non-A→G editing in our findings and the lack of sufficient data validation and a clear mechanism, the significance of these non-ADAR target sites remains unresolved.
(a,b) Distribution of editing sites in the poly(A)+ RNAs (left) and poly(A)− RNAs (right) transcriptome by editing type (a) or incidence per unit length (Mbp) (b) of the indicated structure classes. 'Unknown' denotes editing sites located in regions with conflicting annotations in the database. (c–e) Distribution of RNA editing levels in poly(A)+ RNAs (c), the protein-coding (CDS) region of mRNA (d) or poly(A)− RNAs (e). (f) Two examples of noncoding RNA genes with multiple edits (Jpx/NR_024582, top; Malat1/NR_002819, bottom) that, to our knowledge, have not been reported to show evidence of RNA editing. RNA editing sites identified from YH RNA-Seq data are highlighted by red boxes. Green boxes denote editing sites from DARNED. (g,h) Frequency of nucleotides in the flanking sequences (10 bp both upstream and downstream) of the editing loci in poly(A)+ RNA (g) andin poly(A)− RNA (h). The editing loci are denoted as nucleotide position 0. (i) The conservation of editing sites in poly(A)+ RNA (left) and poly(A)− RNA (right). Total sites as well as the non-Alu component of the data set were analyzed independently, as indicated. Shown are fractions of all, A→G or non-A→G sites that are located in the most evolutionarily conserved regions (score ≥200 in the UCSC conservation table). Statistical significance of the A→G versus non-A→G comparisons was calculated by the Fisher's exact test.
Characterization of the inferred editing sites
We next sought to take advantage of the large size of our data set to analyze sequence and structural attributes of the identified RNA editing sites. Consistent with previous findings, >90% of the sites were A→G substitutions (Fig. 2a). Among sites in poly(A)+ RNA, most were located in intronic RNAs (Table 1). However, for both A→G and non-A→G variants, the 3′-untranslated region (UTR) of transcripts contained the greatest number of variants per Mbp (Fig. 2b). Conversely, coding regions contained considerably fewer edited sites, with only 80 found in total (Supplementary Table 9). Of all of the base alterations located in coding regions, 50% led to changes in the encoded amino acids. We observed that in transcripts from both coding and noncoding regions sites were edited to varying degrees—that is, not all transcripts from a region contained the editing event (Fig. 2c,d).
Further analysis of the sites in poly(A)− RNAs showed that they shared similar sequence attributes with those in the poly(A)+ RNAs (Table 1), particularly in terms of editing type distribution (∼95% of the total editing sites were of the A→G type; Fig. 2a, right) and degree of editing (Fig. 2e). Figure 2f shows two examples of previously unreported, multiply edited, long noncoding RNA transcripts, Jpx25 (41 sites) and Malat1 (refs. 26,27) (31 sites). A complete summary of the sequence and structural attributes of the editing sites identified in poly(A)+ and poly(A)− RNA species is shown in Supplementary Table 10.
Known attributes of candidate ADAR target sites
To further evaluate whether the A→G sites we identified are potential ADAR substrates, we analyzed several sequence and structural features that have been shown to underlie the occurrence of RNA editing. Consistent with the previous observations5,7,8, our set of A→G sites was significantly enriched in sequence regions that either overlap with the repetitive Alu elements or likely form RNA double-stranded structure, as compared with the other 11 types of variants (Table 2). For the A→G group, we confirmed that sites in both poly(A)+ (Fig. 2g) and poly(A)− (Fig. 2h) RNAs were flanked by sequences that conform to established signatures of A→I(G) editing, an under- or overrepresentation of G nucleotides at the upstream or downstream positions, respectively, of the variant. Furthermore, A→G editing sites showed such sequence signatures irrespective of their association with Alu elements (Supplementary Fig. 4a,b). Notably, for some other types of substitution with a considerable number of incidences, we found overrepresentation of particular nucleotides at the immediately flanking positions (overrepresentation of A or G at, respectively, the upstream or downstream positions of the C→T variation, and overrepresentation of G immediately downstream of T→C editing; Supplementary Fig. 4c,d). Such observed cis preferences support the notion that these non-A→G variants may in fact represent nonrandom base-altering events, likely triggered by enzymatic activities.
Conversely, the extent of A→G sites clustering for our data set (30.89% of sites patterned in clusters of ≥3 sites in 100 bp) is considerably lower than what is found in the DARNED database (85.02%), but is more in line with that of another deep-sequencing data set of breast cancer that generated both DNA and RNA sequence information28 (22.36%) (Supplementary Discussion). Despite the relative underrepresentation of clustered sites in our data set, we found a large number of transcripts with widespread editing (208 genes with ≥10 sites each). Notably, some of these genes are known to have functions that are generally important for cell physiology, such as pathways in p53 signaling and cancer, as well as those specific to the lymphoid cells and immune response (Supplementary Table 11).
Based on the notion that A→I(G) editing and Alu-associated editing are a primate-specific phenomenon with a high prevalence in humans4,29, we next characterized the degree of flanking sequence conservation. Overall, A→G sites displayed a significantly lower degree of flanking sequence conservation than did non-A→G sites (P = 8.16 × 10−101 for the poly(A)+ data and P = 1.12 × 10−82 for poly(A)− data; Fisher's exact test). With regard to the A→G editing sites, although the results above confirmed the extensive association of A→G editing with the Alu sequences, only 0.64% of the Alu-associated A→G edits (67 out of 10,469) were situated in sequences evolutionarily conserved among the primates (based on conserved elements defined by the UCSC PhastCons program; Fig. 2i, left, and Supplementary Table 12). This is a significant underrepresentation in comparison to the non-Alu–associated A→G sites (P = 8.10 × 10−55 for poly(A)+ data and P = 8.70 × 10−13 for poly(A)− data; Fisher's exact test). In contrast, there was a higher prevalence of conserved flanking sequences (>10%) for the non-A→G edits that were not associated with Alu repeats (Fig. 2i, left). Similar observations were made for poly(A)− RNAs (Fig. 2i, right, and Supplementary Table 12). Taken together with the above results, the A→G editing sites identified in this study exhibited sequence and structural features that are largely consistent with those of known RNA editing sites. These results further demonstrate the validity and reliability of our strategy for calling editing sites.
Analysis of a low-coverage RNA-Seq data set
To demonstrate the broader applicability of our pipeline, and also as an independent assessment of data presented in a recent study20 that identified RNA-DNA differences from multiple samples sequenced at lower coverage , we randomly chose data sets generated from two individuals (GM12144 and GM12044; http://www.coriell.org/) and screened for editing sites using our workflow. Such analysis was feasible because these sequence data were obtained from the same tissue source (immortalized B cells) and sequencing platforms as our sequencing data. Previous analyses20 identified ∼1,200 exonic editing sites in each individual. In contrast, our method detected hundreds of candidate sites in total (110 in GM12144 and 328 in GM12044) and only 65 exonic sites in each (Supplementary Table 13), suggesting that the approach used previously20 to analyze these data likely overestimated the number of candidate editing sites or sites with RNA-DNA differences. Most of the editing sites we found were of the A→G type and showed sequence and structural features similar to the sites we identified in YH (Supplementary Table 13). We found 52 sites in GM12144 and 91 in GM12044 that were also identified in YH. Taken together, these findings illustrate the applicability of our pipeline to other data sets as well as the importance of stringent filters to accurately call editing sites (Supplementary Discussion).
Comparison of the YH editome with other data sets
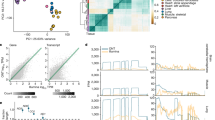
We compared sites called in this study with entries in the DARNED database9 and editing sites found in a breast cancer data set28. This comparison revealed that 1,449 sites identified by our approach corresponded to annotated entities in the DARNED database. However, the vast majority of editing sites in the two sets are not the same, with the overlapping sites representing 6.39% of our calls and 2.73% of the DARNED data set. (Fig. 3a, top). This observation is intriguing insofar as our identified sites essentially display known features of RNA editing. Similar findings were also made in the comparison to the breast cancer data set28 (356 common sites, representing 1.54% and 10.24%, respectively, of our findings and the breast cancer data set; Fig. 3a, bottom), as well as between DARNED and the breast cancer data set (504 common sites, representing ∼1% of total sites from both sets combined; Supplementary Fig. 5).
(a) Extent of overlap in editing sites between data sets in terms of nucleotide position ('site') and corresponding gene ('gene'). The YH data were compared with those of DARNED and the breast cancer RNA-Seq study. Proportions of sites and genes that are unique or common between data sets are shown. (b,c) Examples of genes with multiple editing sites. Distribution of sites (nucleotide positions indicated on the right) in an mRNA gene (ARPC2) (b) and a noncoding gene (SLC35E3) (c) is shown.
Although the marked disparity in editing patterns may be explained by variability in tissues and acquisition methods from which sequences were derived, further analysis revealed that, as compared to the editing sites, greater proportions of the edited gene transcripts, irrespective of the editing positions, are shared among the data sets (Fig. 3a and Supplementary Fig. 5). Notably, most of these genes contain clustered editing sites that are patterned differently between data sets (Fig. 3b,c, and Supplementary Fig. 6), thus supporting the possibility that ADAR targets transcripts in a context-dependent manner.
RNA editing and miRNA-mediated regulation
Previous reports have documented the functional connection between RNA editing and miRNA-mediated post-transcriptional gene silencing30. Therefore, to characterize the potential effects of RNA editing on miRNAs, we first identified 2,474 editing sites in 3′-UTRs. Next, to determine whether an editing site is located in an miRNA target region or not, we extracted a 13-bp sequence centered on the site (6 bp on either side) to search if any 7-bp substring is completely complementary to a known miRNA seed in miRBase19. To simplify our analysis, we focused only on those regions that perfectly matched seed sequence. We then identified mRNA base substitutions that match or disrupt a known miRNA seed. Based on our analysis, 398 of the 1,905 editing sites (20.89%) in the 3′-UTR resided in, and thus may alter, candidate miRNA target sites (Fig. 4a). Moreover, 21.57% of these editing sites may contribute to the creation of new miRNA binding sites in the 3′-UTR. Although these nucleotide changes may affect miRNA targeting, the extent to which the candidate editing event affects the expression of a transcript remains unresolved.
(a) Distribution of editing sites relative to miRNA target sites in the 3′-UTR and possible consequence of RNA editing. In total, 1,905 possible miRNA target sites were predicted in the 3′-UTRs. RNA editing may disrupt or switch miRNA recognition if editing occurs at nucleotides in the miRNA seed region (“altered”). Editing on non-miRNA target sites (“no match”) may generate new miRNA targets (“new targets”). (b) Distribution of RNA edits identified from miRNAs. (c,d) Examples of miRNA species that harbor RNA edits. The most abundant perfect-match and single-mismatch reads from the hsa-mir-200b (c) and hsa-mir-548o (d) loci support A→I editing in the seed region and G→A editing, respectively.
Lastly, we profiled nucleotide sequence variants in miRNAs identified from the small RNA-Seq data (Supplementary Tables 3 and 14). A total of 44 editing sites distributed among 30 different mature miRNA sequences were called by our algorithm (Online Methods and Supplementary Table 15). A→G variants represented the most abundant type, but they were not considerably overrepresented, as we had observed for poly(A)+ and poly(A)− RNAs. Other types of editing were also detected to various degrees (Fig. 4b and Supplementary Tables 15,16). Of the 11 putative A→G editing sites, 10 (Supplementary Table 16) had not been found by previous studies31,32,33,34. Figure 4c,d illustrates two examples of miRNAs with inferred editing sites in their mature sequences and the distribution of corresponding reference and variant sequence reads. Further sequence characterization uncovered a slight positional enrichment of edits at the nucleotide 20 of the mature sequence (Supplementary Fig. 7). Moreover, for a subset of the miRNAs (11 sites), nucleotide variants fall within the seed regions (Supplementary Table 15), and thus may alter the target specificity during regulation. Taken together, these observations support the notion that RNA editing is linked to miRNA-mediated regulation of gene expression.
Discussion
We report evidence of extensive RNA editing in a human cell line, which underscores the need for robust methods to detect these events. We developed a pipeline for identifying RNA editing events by screening RNA-DNA differences in the same individual through successive quality control filters. The pipeline performed well on simulated data in terms of both sensitivity and specificity (Fig. 1c and Supplementary Tables 4,5).
False positives are a critical issue in the analysis of SNVs from RNA-Seq data, as evidenced by the simulation results (Fig. 1c) and our experience with an earlier version of our method that did not include strict quality control filters (data not shown). This problem, which is also evident in genomic SNP detection24,35 and the recent reevaluation of the large-scale RNA-Seq study36, may be due to several factors. Some of the false positives are certainly a result of inaccuracy in read alignment, of which paralogous, highly similar sequences, and splice junctions represent major sources. In addition, mapping reliability can be further compromised in the presence of RNA transcript sequence variation. From a technical perspective, paired-end reads with greater length (75, 90 or 100 bp in our study) presumably should increase the fidelity of read alignment37. To address the inconsistency in read mapping, we implemented two independent filters in our calling method (the MES and BLAT steps) that remove false-positive results that were identified in a naive analysis of simulated reads. Finally, biases in calling editing sites may also stem from insufficient coverage and accuracy of the genome sequence, which becomes problematic in ascertaining potential edits at positions that correspond to genomic polymorphisms. However, the 36-fold average coverage of our genome-resequencing data, combined with the YH genome variants filter that addresses zygosity and copy number variation of the genome at the editing sites, reduces the likelihood of such errors.
Our method and results affirm that quality control filters are necessary to accurately identify RNA editing sites. This study also indicates the need to archive RNA edits for the development of more thorough statistical models that incorporate prior knowledge of sequence variation and the sequencing technologies used. Notably, recent computational approaches for detecting A→I(G) base changes in human mRNA databases also incorporated molecular features that underlie RNA editing, such as RNA folding characteristics or tissue-preferred distribution of editing events38,39. These filter criteria may thus be included as additional modules in our workflow to analyze more complex or functionally relevant data sets in future deep-sequencing studies.
As this manuscript was being prepared, two large-scale screens for RNA-DNA differences that used deep-sequencing approaches similar to ours were reported20,21. Several differences were notable between these studies, including the design of the site-calling pipelines, the extent of the sampled transcriptome, the number of sites identified and the distribution of editing types. Notably, these studies reported that ∼23% (ref. 20) and 62% (ref. 21) of editing sites were A→G changes, whereas the vast majority (>90%) of our candidate sites were A→G changes. Furthermore, our work complements previous and recent findings with in-depth information of editing across a broader sampling of the transcriptome, particularly the intergenic transcripts.
Amid the recent deep-sequencing studies of RNA editing, there has been substantial controversy centered on the technical drawbacks of this technology as well as related analysis algorithms and experimental design. We suggest that our overall methodology thoroughly addresses these concerns and minimizes errors when inferring editing sites from RNA-Seq data. The need for stringent criteria in identifying RNA-DNA differences is reinforced by a recent report showing that, after accounting for paralogous and genomic variant sequences, a considerable portion of the candidate sites that were identified in a previous study20 might actually represent spurious results36. We therefore also did an independent assessment of these data20 using our workflow (Supplementary Table 13). This analysis revealed that candidate editing or RNA-DNA difference sites were likely overestimated by their approach. In addition to the potential contribution of paralogous sequences and genomic variants as the error source, we also found that data quality and depth played a role in the possibly erroneous calling of variants. Almost 60% of the sites identified previously20 could be removed by our “read parameter” filter based on their location within 8 bp from the ends of 50-bp reads. Moreover, because of the low depth of individual genome sequences sampled in that report, some of the putative editing sites did not fulfill the requirements defined by our “genome variants” filter and may actually represent polymorphic sequences encoded by the genome. Notably, however, features of the editing sites called by our pipeline from the data20 are similar to features of sites called from our RNA-Seq data (Supplementary Table 13). This suggests that the discrepancies between the two studies might be attributed mainly to the different study designs rather than the underlying molecular biology.
In summary, our results support the accuracy of our multifilter modular pipeline to annotate an editome and to provide a global and quantitative catalog of nucleotide variants in a transcriptome. The next step is clearly to apply this methodology to larger-scale deep-sequencing studies involving additional physiologically relevant samples, so as to profile and compare editomes more comprehensively and accurately.
Methods
Library construction, sequencing and validation.
Experimental methods, including library construction and sequencing of the whole-transcriptome, as well as validation by Sanger sequencing (primer sequences are listed in Supplementary Table 17), are described in detail in the Supplementary Methods. The reads are available in the NCBI Sequence Read Archive, under accession number SRA043767.1 and also in reference 40. The Asian genome data were sampled from an anonymous male Han Chinese and sequenced by the Illumina Genome Analyzer22. These data are available in the EBI/NCBI41 Short Read Archive and in the YH database (http://yh.genomics.org.cn/). The editome and RNA-Seq data sets generated from this study can also be viewed through an online browser (http://yheditome.genomics.cn/mgb2/gbrowse).
Illumina reads alignment.
The paired-end reads obtained from Illumina GAIIx and HiSeq 2000 were aligned to the reference genome (NCBI Build 36.1, hg18) using the SOAP2 program41, with two reads belonging to a pair being aligned together with both in the correct orientation. Due to the potential uncertainty in read alignment across splice junctions, SOAP2 was used in this regard rather than tools that utilize gapped alignment across exon boundary, such as SOAPsplice42. To provide enhanced SNV calling, at most three mismatches for the 75-bp reads and four mismatches for the 90-bp and 100-bp reads were allowed when aligning the cDNA reads to the reference genome (the -v parameter). The alignments with the least number of mismatches were defined as 'best hits'. If there was only one single best hit for a read, then the read was taken as uniquely placed; a read with multiple equal best hits was taken as repeatedly placed. In this study, we discarded from paired-end reads repeatedly placed reads and also potential PCR duplicates, and used only those reads with unique ungapped genome alignment for consensus calling and editing site detection.
Simulated data.
Paired-end reads with fixed length of 75 bp were simulated randomly from chromosome 1 of the human RefSeq. We carried out the following process to create simulated reads that were similar to that which would be generated by GA sequencing:
-
1
Use chromosome 1 of the NCBI human RefSeq as a reference and produce an identical copy of the corresponding transcriptome sequence.
-
2
Two sets of simulated data were created. In the first set, SNVs were randomly generated in the copy sequence by MAQ43 with default options. In the other simulated set, only A→G substitution at positions that are referenced in the DARNED database9 (814 sites total for coding regions of chromosome 1) were introduced.
-
3
Reads were generated from random locations for both forward and reverse strands of the copy; 5-, 10-, 20-, and 50-fold coverage reads were generated for the random-sites data, 50-fold coverage reads for the DARNED data sets.
-
4
Both data sets were subjected to read alignment and SNV calling as described below. Results from each step of the analysis were compared with the known types and positions of simulated SNVs and used to evaluate our SNV calling approach (Fig. 1c, Supplementary Fig. 2 and Supplementary Tables 4 and 5).
RNA editing sites/RNA-centric SNVs detection.
Our bioinformatics analysis scheme implemented multiple filters with stringent thresholds to facilitate unbiased detection of bona fide editing or base substitution events in the RNA-Seq reads (Supplementary Data). RNA-centric SNVs were first identified from aligned cDNA reads using SOAPsnp44, which uses a method based on Bayes' theorem (the reverse probability model) to call consensus genotype by carefully considering the data quality, alignment and recurring experimental errors, with parameters e = 0.0001 and r = 0.00005. We further lifted a default filter in the basic filter step of the program that was designed to discard sequence reads with more than one variant within a 5-bp span. Since independent analysis of the DSN-normalized and nonnormalized libraries did not reveal much differences in the editing sites identified, SOAP2 alignment results from all poly(A)+ RNA-Seq libraries were therefore merged as a single data set for analysis.
The initially identified SNVs were then filtered by the following steps:
-
1
Basic filter. Retain SNVs that meet the following criteria: quality score of consensus genotype ≥ 20; covered depth ≥ 5; repeats (estimated copy number of the flanked sequence in genome) ≤ 1.
-
2
Read parameter filter. Parameters, such as sequencing quality score, distance of a potential SNV to the end of the supporting read, and coverage depth of the SNV, were optimized on the basis of their effects on pipeline performance on the first simulated data set (Supplementary Table 4 and Supplementary Fig. 2): distance cutoff = 15 (m, the minimal distance of a SNV site to its supporting reads' ends); quality score cutoff = 20 (q, minimal sequencing quality score of SNV-corresponding nucleotide); and supporting reads number cutoff = 2 (n, minimal number of supporting reads that meet the above two cutoff parameters).
-
3
RNA-DNA variants filter. Further, to focus on RNA-DNA variants only, sites of which DNA genotypes are the same as RNA genotypes were removed.
-
4
YH genome variants filter. To distinguish RNA editing from allele-specific expression and duplication polymorphisms, we kept SNVs remaining from step 3 only if their corresponding DNA genotypes are homozygous and diploid in copy number. The first component of the filter, the zygosity filter, was designed to assess several parameters of YH genome sequence reads that correspond to a given candidate site: depth is ≥5; consensus quality is ≥20; average quality of the first best allele ≥ 20; depth of the second best allele, if present, is <5% of the total number of reads; the second best allele should not be the variant allele in the RNA data; and average sequencing quality of the second best allele is <10. The second component was designed to exclude genomic duplication polymorphisms, which may also contribute to inaccurate calling of editing sites36. To this end, we assessed potential copy number variations (CNVs) within the YH genome using the CNVnator tool45 with bin set to 50, and removed sites that were nondiploid in nature. This analysis revealed that ∼10% of this genome exhibits variation in copy number, ∼95% of which is located in the intergenic region (data not shown).
-
5
MES filter. Next, we removed misaligned reads that arise from mapping error inherent to the mapping algorithm (MES). The MES set was generated as follows: read sequences were simulated based on all human genes (hg18 transcriptome) using MAQ without mutation (-r parameter). After alignment and SNV calling by SOAP2 and SOAPsnp, respectively, the identified SNVs were passed through the above two filters. The resultant collection of SNVs is termed MES and represents an inherently error-prone set of sites that are incorrectly called owing to the nature of mapping and/or calling algorithms. Any SNVs derived from step 2 that matched the MES were removed.
-
6
Strand filter. A strand bias filter was also installed to remove potential strand-specific errors in sequences generated by the Illumina platform46,47. For a particular SNV site, the reads carrying a reference or alternative allele that maps to the plus and minus strand in the genome were counted and evaluated using a Fisher's exact test47. Sites whose reads exhibited strand bias in distribution (P < 0.01), and whose number of supporting reads mapped to either strand is <2, were discarded.
-
7
BLAT filter. To further address the potential pitfall of paralogous sequences in site calling, we used BLAT to search for SNVs whose supporting reads are multiply aligned to the reference genome with the same mismatch tolerance used in SOAP alignment, and subsequently discarded all supporting reads with more than one hit within the mismatch tolerance. We then filtered SNVs that had <2 qualified supporting reads.
-
8
Known SNPs filter. Further, to eliminate germline variants, SNVs remaining from step 5 were cross-referenced against known SNP databases, including the 1000 Genomes Project (http://www.1000genomes.org/), the genomes of Yoruba48, Watson49, Korean50 and dbSNP (version 129).
-
9
Multiple type of mismatches filter. Discard SNV candidate sites that display more than one nonreference type (e.g., reference allele is A, nonreference alleles are G and T).
-
10
Editing degree filter. Finally, polymorphic sites with extreme degree of variation (100%) were excluded, based on our observation that >90% of sites in MES exhibited 100% variation (Supplementary Fig. 8). Degree ofediting for a particular site was calculated as the ratio of reads supporting the variant allele to the total number of reads covering the site.
Analysis of the sequence and structural features of RNA editing.
To identify sites that are potentially located within a double-stranded RNA (dsRNA) structure, or sites in 3′-UTR that are likely microRNA seed matches, we followed procedures described elsewhere19. Clustering of editing sites is defined as occurrence of at least 3 nucleotide variants within a 100-bp window. A site is considered to be located in a conserved region, if it is annotated as “most conserved” by the UCSC genome browser (http://genome.ucsc.edu/). The coding sequence is defined by the RefSeq annotation. For gene enrichment analysis, after selecting highly edited genes (≥10 variant sites per gene), we sorted this set into pathways using the DAVID pathway-classification tool (http://david.abcc.ncifcrf.gov/).
Identification of miRNA and editing.
Overview of the small RNA-Seq data is shown in Supplementary Table 14. The small RNA reads were subjected to the following filtering processes: (i) Filter out low-quality reads; (ii) trim 3′ adaptor sequence by a dynamic programming algorithm; (iii) remove adaptor contaminations formed by adaptor ligation; and (iv) retain only short trimmed reads of sizes from 18 to 30 nt. To annotate and categorize small RNAs into different classes, we performed the following analyses. We filtered out small RNA reads that might be from known noncoding RNAs by comparing them with known noncoding RNAs (we considered rRNA, tRNA, snRNA and snoRNA) deposited in the Rfam database (http://rfam.sanger.ac.uk/, release 9.1) and the NCBI Genbank (http://www.ncbi.nlm.nih.gov/genbank/). Small RNA reads assigned to exonic regions were also discarded. After removing small RNA reads belonging to the above categories, the rest were subjected to MIREAP (https://sourceforge.net/projects/mireap/), which identifies miRNA candidates according to the canonical hairpin structure and sequencing data. The identified small RNA (miRNA) reads were then aligned (using BLAST) to miRNA reference sequences (http://www.mirbase.org/, release 16) with tolerance for, at most, one mismatch. Reads that were uniquely aligned and overlapped with known miRNAs were used to identify miRNA editing sites. First, we identified reads with mismatch to hg18 genome. Reads with mismatch within 1 nt at 5′ end or 2 nt at 3′ end were discarded. Then we identified miRNA edits by the following criteria: (i) Sequencing depth of editing sites should be equal to or larger than 5; (ii) frequency of SNV occurrence ≥5% & ≤95%; and (iii) variants that were not found in previous SNP annotations (YH, 1000 genomes project, Yoruba, Watson, Korean and dbSNP version 129).
Accession code.
EBI/NCBI Short Read Archive: ERA000005 (YH genome data). RNA-Seq reads are available in the NCBI Sequence Read Archive, under accession number SRA043767.1.
Accession codes
References
Jepson, J.E. & Reenan, R.A. RNA editing in regulating gene expression in the brain. Biochim. Biophys. Acta 1779, 459–470 (2008).
Nishikura, K. Functions and regulation of RNA editing by ADAR deaminases. Annu. Rev. Biochem. 79, 321–349 (2010).
Osenberg, S. et al. Alu sequences in undifferentiated human embryonic stem cells display high levels of A-to-I RNA editing. PLoS ONE 5, e11173 (2010).
Paz-Yaacov, N. et al. Adenosine-to-inosine RNA editing shapes transcriptome diversity in primates. Proc. Natl. Acad. Sci. USA 107, 12174–12179 (2010).
Athanasiadis, A., Rich, A. & Maas, S. Widespread A-to-I RNA editing of Alu-containing mRNAs in the human transcriptome. PLoS Biol. 2, e391 (2004).
Blow, M., Futreal, P.A., Wooster, R. & Stratton, M.R. A survey of RNA editing in human brain. Genome Res. 14, 2379–2387 (2004).
Kim, D.D. et al. Widespread RNA editing of embedded alu elements in the human transcriptome. Genome Res. 14, 1719–1725 (2004).
Levanon, E.Y. et al. Systematic identification of abundant A-to-I editing sites in the human transcriptome. Nat. Biotechnol. 22, 1001–1005 (2004).
Kiran, A. & Baranov, P.V. DARNED: a DAtabase of RNa EDiting in humans. Bioinformatics 26, 1772–1776 (2010).
Hundley, H.A. & Bass, B.L. ADAR editing in double-stranded UTRs and other noncoding RNA sequences. Trends Biochem. Sci. 35, 377–383 (2010).
Farajollahi, S. & Maas, S. Molecular diversity through RNA editing: a balancing act. Trends Genet. 26, 221–230 (2010).
Pullirsch, D. & Jantsch, M.F. Proteome diversification by adenosine to inosine RNA editing. RNA Biol. 7, 205–212 (2010).
Eisenberg, E., Li, J.B. & Levanon, E.Y. Sequence based identification of RNA editing sites. RNA Biol. 7, 248–252 (2010).
Reid, J.G. et al. Mouse let-7 miRNA populations exhibit RNA editing that is constrained in the 5′-seed/cleavage/anchor regions and stabilize predicted mmu-let-7a:mRNA duplexes. Genome Res. 18, 1571–1581 (2008).
Wang, Z., Gerstein, M. & Snyder, M. RNA-Seq: a revolutionary tool for transcriptomics. Nat. Rev. Genet. 10, 57–63 (2009).
Mortazavi, A., Williams, B.A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Morozova, O., Hirst, M. & Marra, M.A. Applications of new sequencing technologies for transcriptome analysis. Annu. Rev. Genomics Hum. Genet. 10, 135–151 (2009).
Wahlstedt, H., Daniel, C., Enstero, M. & Ohman, M. Large-scale mRNA sequencing determines global regulation of RNA editing during brain development. Genome Res. 19, 978–986 (2009).
Li, J.B. et al. Genome-wide identification of human RNA editing sites by parallel DNA capturing and sequencing. Science 324, 1210–1213 (2009).
Li, M. et al. Widespread RNA and DNA sequence differences in the human transcriptome. Science 333, 53–58 (2011).
Bahn, J.H. et al. Accurate Identification of A-to-I RNA editing in human by transcriptome sequencing. Genome Res. 142–150 (2012).
Wang, J. et al. The diploid genome sequence of an Asian individual. Nature 456, 60–65 (2008).
Morin, R. et al. Profiling the HeLa S3 transcriptome using randomly primed cDNA and massively parallel short-read sequencing. Biotechniques 45, 81–94 (2008).
Cirulli, E.T. et al. Screening the human exome: a comparison of whole genome and whole transcriptome sequencing. Genome Biol. 11, R57 (2010).
Tian, D., Sun, S. & Lee, J.T. The long noncoding RNA, Jpx, is a molecular switch for X chromosome inactivation. Cell 143, 390–403 (2010).
Tripathi, V. et al. The nuclear-retained noncoding RNA MALAT1 regulates alternative splicing by modulating SR splicing factor phosphorylation. Mol. Cell 39, 925–938 (2010).
Bernard, D. et al. A long nuclear-retained non-coding RNA regulates synaptogenesis by modulating gene expression. EMBO J. 29, 3082–3093 (2010).
Shah, S.P. et al. Mutational evolution in a lobular breast tumour profiled at single nucleotide resolution. Nature 461, 809–813 (2009).
Eisenberg, E. et al. Is abundant A-to-I RNA editing primate-specific? Trends Genet. 21, 77–81 (2005).
Borchert, G.M. et al. Adenosine deamination in human transcripts generates novel microRNA binding sites. Hum. Mol. Genet. 18, 4801–4807 (2009).
Chiang, H.R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Kawahara, Y. et al. Frequency and fate of microRNA editing in human brain. Nucleic Acids Res. 36, 5270–5280 (2008).
Kawahara, Y. et al. Redirection of silencing targets by adenosine-to-inosine editing of miRNAs. Science 315, 1137–1140 (2007).
Kawahara, Y., Zinshteyn, B., Chendrimada, T.P., Shiekhattar, R. & Nishikura, K. RNA editing of the microRNA-151 precursor blocks cleavage by the Dicer-TRBP complex. EMBO Rep. 8, 763–769 (2007).
Degner, J.F. et al. Effect of read-mapping biases on detecting allele-specific expression from RNA-sequencing data. Bioinformatics 25, 3207–3212 (2009).
Schrider, D.R., Gout, J.F. & Hahn, M.W. Very few RNA and DNA sequence differences in the human transcriptome. PLoS ONE 6, e25842 (2011).
Pepke, S., Wold, B. & Mortazavi, A. Computation for ChIP-seq and RNA-seq studies. Nat. Methods 6, S22–S32 (2009).
He, T. et al. Computational detection and functional analysis of human tissue-specific A-to-I RNA editing. PLoS ONE 6, e18129 (2011).
Maas, S. et al. Genome-wide evaluation and discovery of vertebrate A-to-I RNA editing sites. Biochem. Biophys. Res. Commun. 412, 407–412 (2011).
Tian, Z. et al. Transcriptome from a lymphoblastoid cell line taken from the YH Han Chinese individual. Giga Sci. http://dx.doi.org/10.5524/100013. (2011).
Li, R. et al. SOAP2: an improved ultrafast tool for short read alignment. Bioinformatics 25, 1966–1967 (2009).
Huang, S. et al. SOAPsplice: genome-wide ab initio detection of splice junctions from RNA-Seq data. Front. Genet. 2 (2011).
Li, H., Ruan, J. & Durbin, R. Mapping short DNA sequencing reads and calling variants using mapping quality scores. Genome Res. 18, 1851–1858 (2008).
Li, R. et al. SNP detection for massively parallel whole-genome resequencing. Genome Res. 19, 1124–1132 (2009).
Abyzov, A., Urban, A.E., Snyder, M. & Gerstein, M. CNVnator: an approach to discover, genotype, and characterize typical and atypical CNVs from family and population genome sequencing. Genome Res. 21, 974–984 (2011).
Nakamura, K. et al. Sequence-specific error profile of Illumina sequencers. Nucleic Acids Res. 39, e90 (2011).
DePristo, M.A. et al. A framework for variation discovery and genotyping using next-generation DNA sequencing data. Nat. Genet. 43, 491–498 (2011).
Bentley, D.R. et al. Accurate whole human genome sequencing using reversible terminator chemistry. Nature 456, 53–59 (2008).
Wheeler, D.A. et al. The complete genome of an individual by massively parallel DNA sequencing. Nature 452, 872–876 (2008).
Kim, J.I. et al. A highly annotated whole-genome sequence of a Korean individual. Nature 460, 1011–1015 (2009).
Acknowledgements
This project was supported by a National Basic Research Program of China (973 program no. 2011CB809200, 2011CB809202 and 2011CB809203), the National Natural Science Foundation of China (30725008, 30890032 and 30811130531), the Chinese 863 program (2006AA02A302 and 2009AA022707), the Shenzhen Municipal Government of China and the Local Government of Yantian District of Shenzhen (grants JC200903190767A, JC200903190772A, ZYC200903240076A, CXB200903110066A, ZYC200903240077A and ZYC200903240080A) and the Ole Rømer grant from the Danish Natural Science Research Council.
Author information
Authors and Affiliations
Contributions
Z.P., B.C.-M.T. and J.W. conceived and designed the experiment; Z.P., Y.C., B.C.-M.T., L.K. and Y.Z. performed data analysis and informatics; Z.T., Yu L., X.H., Yan L. and L.B. carried out sample preparation and sequencing experiments; Y.C., Z.T., W.Z., X.T., J.G. and Z.D. designed and executed experimental validation; Z.P., B.C.-M.T. and J.W. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1, 4–5, 7, 10–11, 14, 16, Supplementary Discussion, Supplementary Methods and Supplementary Figures 1–8 (PDF 10917 kb)
Rights and permissions
About this article
Cite this article
Peng, Z., Cheng, Y., Tan, BM. et al. Comprehensive analysis of RNA-Seq data reveals extensive RNA editing in a human transcriptome. Nat Biotechnol 30, 253–260 (2012). https://doi.org/10.1038/nbt.2122
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.2122
This article is cited by
-
L-GIREMI uncovers RNA editing sites in long-read RNA-seq
Genome Biology (2023)
-
Changes in ADAR RNA editing patterns in CMV and ZIKV congenital infections
BMC Genomics (2023)
-
Deep transcriptome profiling reveals limited conservation of A-to-I RNA editing in Xenopus
BMC Biology (2023)
-
DeepEdit: single-molecule detection and phasing of A-to-I RNA editing events using nanopore direct RNA sequencing
Genome Biology (2023)
-
Precision RNA base editing with engineered and endogenous effectors
Nature Biotechnology (2023)