Abstract
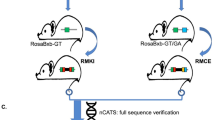
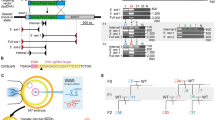
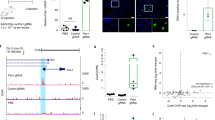
The 21–23 nucleotide, single-stranded RNAs classified as microRNAs (miRNA) perform fundamental roles in diverse cellular and developmental processes. In contrast to the situation for protein-coding genes, no public resource of miRNA mouse mutant alleles exists. Here we describe a collection of 428 miRNA targeting vectors covering 476 of the miRNA genes annotated in the miRBase registry. Using these vectors, we generated a library of highly germline-transmissible C57BL/6N mouse embryonic stem (ES) cell clones harboring targeted deletions for 392 miRNA genes. For most of these targeted clones, chimerism and germline transmission can be scored through a coat color marker. The targeted alleles have been designed to be adaptable research tools that can be efficiently altered by recombinase-mediated cassette exchange to create reporter, conditional and other allelic variants. This miRNA knockout (mirKO) resource can be searched electronically and is available from ES cell repositories for distribution to the scientific community.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Bartel, D.P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Guo, H., Ingolia, N.T., Weissman, J.S. & Bartel, D.P. Mammalian microRNAs predominantly act to decrease target mRNA levels. Nature 466, 835–840 (2010).
Wang, Y. et al. Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nat. Genet. 40, 1478–1483 (2008).
Melton, C., Judson, R.L. & Blelloch, R. Opposing microRNA families regulate self-renewal in mouse embryonic stem cells. Nature 463, 621–626 (2010).
van Rooij, E. et al. Control of stress-dependent cardiac growth and gene expression by a microRNA. Science 316, 575–579 (2007).
Zhao, Y. et al. Dysregulation of cardiogenesis, cardiac conduction, and cell cycle in mice lacking miRNA-1–2. Cell 129, 303–317 (2007).
Saba, R. & Schratt, G.M. MiRNAs in neuronal development, function and dysfunction. Brain Res. 1338, 3–13 (2010).
O'Connell, R.M., Rao, D.S., Chaudhuri, A.A. & Baltimore, D. Physiological and pathological roles for microRNAs in the immune system. Nat. Rev. Immunol. 10, 111–122 (2010).
Calin, G.A. & Croce, C.M. MicroRNA signatures in human cancers. Nat. Rev. Cancer 6, 857–866 (2006).
Miska, E.A. et al. Most Caenorhabditis elegans microRNAs are individually not essential for development or viability. PLoS Genet. 3, e215 (2007).
Austin, C.P. et al. The knockout mouse project. Nat. Genet. 36, 921–924 (2004).
Skarnes, W.C. et al. A conditional knockout resource for the genome-wide study of mouse gene function. Nature 474, 337–342 (2011).
Ringwald, M. et al. The IKMC web portal: a central point of entry to data and resources from the International Knockout Mouse Consortium. Nucleic Acids Res. 39, D849–D855 (2011).
Kanellopoulou, C. et al. Dicer-deficient mouse embryonic stem cells are defective in differentiation and centromeric silencing. Genes Dev. 19, 489–501 (2005).
Harfe, B.D., McManus, M.T., Mansfield, J.H., Hornstein, E. & Tabin, C.J. The RNase III enzyme Dicer is required for morphogenesis but not patterning of the vertebrate limb. Proc. Natl. Acad. Sci. USA 102, 10898–10903 (2005).
Cobb, B.S. et al. T cell lineage choice and differentiation in the absence of the RNase III enzyme Dicer. J. Exp. Med. 201, 1367–1373 (2005).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007).
Thai, T.H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007).
Lu, L.F. et al. Function of miR-146a in controlling Treg cell-mediated regulation of Th1 responses. Cell 142, 914–929 (2010).
Griffiths-Jones, S., Grocock, R.J., van Dongen, S., Bateman, A. & Enright, A.J. miRBase: microRNA sequences, targets and gene nomenclature. Nucleic Acids Res. 34, D140–D144 (2006).
Cai, X., Hagedorn, C.H. & Cullen, B.R. Human microRNAs are processed from capped, polyadenylated transcripts that can also function as mRNAs. RNA 10, 1957–1966 (2004).
Marson, A. et al. Connecting microRNA genes to the core transcriptional regulatory circuitry of embryonic stem cells. Cell 134, 521–533 (2008).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
Rodriguez, A., Griffiths-Jones, S., Ashurst, J.L. & Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 14, 1902–1910 (2004).
Chan, W. et al. A recombineering based approach for high-throughput conditional knockout targeting vector construction. Nucleic Acids Res. 35, e64 (2007).
Chen, Y.T. & Bradley, A. A new positive/negative selectable marker, puDeltatk, for use in embryonic stem cells. Genesis 28, 31–35 (2000).
Schlake, T. & Bode, J. Use of mutated FLP recognition target (FRT) sites for the exchange of expression cassettes at defined chromosomal loci. Biochemistry 33, 12746–12751 (1994).
Pettitt, S.J. et al. Agouti C57BL/6N embryonic stem cells for mouse genetic resources. Nat. Methods 6, 493–495 (2009).
Raymond, C.S. & Soriano, P. High-efficiency FLP and PhiC31 site-specific recombination in mammalian cells. PLoS ONE 2, e162 (2007).
Sauer, B. & McDermott, J. DNA recombination with a heterospecific Cre homolog identified from comparison of the pac-c1 regions of P1-related phages. Nucleic Acids Res. 32, 6086–6095 (2004).
Anastassiadis, K. et al. Dre recombinase, like Cre, is a highly efficient site-specific recombinase in E. coli, mammalian cells and mice. Dis. Model. Mech. 2, 508–515 (2009).
Houbaviy, H.B., Murray, M.F. & Sharp, P.A. Embryonic stem cell-specific MicroRNAs. Dev. Cell 5, 351–358 (2003).
Chiang, H.R. et al. Mammalian microRNAs: experimental evaluation of novel and previously annotated genes. Genes Dev. 24, 992–1009 (2010).
Griffiths-Jones, S. The microRNA Registry. Nucleic Acids Res. 32, D109–D111 (2004).
Liu, P., Jenkins, N.A. & Copeland, N.G. A highly efficient recombineering-based method for generating conditional knockout mutations. Genome Res. 13, 476–484 (2003).
McMahon, A.P. & Bradley, A. The Wnt-1 (int-1) proto-oncogene is required for development of a large region of the mouse brain. Cell 62, 1073–1085 (1990).
Wobus, A.M., Guan, K., Yang, H.T. & Boheler, K.R. Embryonic stem cells as a model to study cardiac, skeletal muscle, and vascular smooth muscle cell differentiation. Methods Mol. Biol. 185, 127–156 (2002).
Sauer, B. & Henderson, N. Cre-stimulated recombination at loxP-containing DNA sequences placed into the mammalian genome. Nucleic Acids Res. 17, 147–161 (1989).
Araki, K., Araki, M., Miyazaki, J. & Vassalli, P. Site-specific recombination of a transgene in fertilized eggs by transient expression of Cre recombinase. Proc. Natl. Acad. Sci. USA 92, 160–164 (1995).
Seibler, J., Schubeler, D., Fiering, S., Groudine, M. & Bode, J. DNA cassette exchange in ES cells mediated by Flp recombinase: an efficient strategy for repeated modification of tagged loci by marker-free constructs. Biochemistry 37, 6229–6234 (1998).
Buchholz, F., Angrand, P.O. & Stewart, A.F. Improved properties of FLP recombinase evolved by cycling mutagenesis. Nat. Biotechnol. 16, 657–662 (1998).
Acknowledgements
We thank Y. Mok, D. Thomas, M. Li and G. Duddy for assistance with sections of the project. For database support and high-throughput vector designs, we are indebted to V. Iyer, D. Oakley, W. Yang, D. Klose and S. Briois. We thank A. West for advice on long range PCR genotyping. We are grateful for the advice and reagents for high-throughput vector construction received from P. Liu. We thank E. Miska for helpful comments on the manuscript. We thank R. Rad for advice with Q-PCR. We thank the Sanger Institute mouse mutant generation and animal facilities teams. We thank the public distribution repositories at MMRRC University of California Davis, USA, and as part of the EUCOMMTOOLS initiative EuMMCR at the Helmholtz Zentrum München, Munich, Germany, for agreeing to support the future distribution of the mirKO resource. This work was funded by the Wellcome Trust (WT079643). The IKMC database was developed with funding from the European Union and the US National Institutes of Health.
Author information
Authors and Affiliations
Contributions
H.M.P. planned and managed the project, constructed novel plasmid reagents and performed high-throughput vector construction, ES cell targeting experiments, post-targeting modifications, collated data and wrote the paper. H.K.-Y. performed high-throughput vector construction, planned and performed the ES cell targeting experiments, ES cell genotyping and collated data. J.D.C. performed high-throughput vector construction, ES cell targeting and genotyping. F.C.L. performed ES cell targeting experiments. A.B. initiated and supervised the project and wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Table 2 and Supplementary Figures 1–4 (PDF 5058 kb)
Supplementary Table 1
Current status of individual mirKO targeting projects. (XLS 141 kb)
Rights and permissions
About this article
Cite this article
Prosser, H., Koike-Yusa, H., Cooper, J. et al. A resource of vectors and ES cells for targeted deletion of microRNAs in mice. Nat Biotechnol 29, 840–845 (2011). https://doi.org/10.1038/nbt.1929
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1929
This article is cited by
-
A mouse model that is immunologically tolerant to reporter and modifier proteins
Communications Biology (2020)
-
Sensational MicroRNAs: Neurosensory Roles of the MicroRNA-183 Family
Molecular Neurobiology (2020)
-
MiR-211 is essential for adult cone photoreceptor maintenance and visual function
Scientific Reports (2017)
-
MicroRNAs: master regulators of drug resistance, stemness, and metastasis
Journal of Molecular Medicine (2014)
-
TALEN-based knockout library for human microRNAs
Nature Structural & Molecular Biology (2013)