Abstract
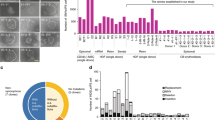
Realizing the therapeutic potential of human induced pluripotent stem (iPS) cells will require robust, precise and safe strategies for genetic modification, as cell therapies that rely on randomly integrated transgenes pose oncogenic risks. Here we describe a strategy to genetically modify human iPS cells at 'safe harbor' sites in the genome, which fulfill five criteria based on their position relative to contiguous coding genes, microRNAs and ultraconserved regions. We demonstrate that ∼10% of integrations of a lentivirally encoded β-globin transgene in β-thalassemia-patient iPS cell clones meet our safe harbor criteria and permit high-level β-globin expression upon erythroid differentiation without perturbation of neighboring gene expression. This approach, combining bioinformatics and functional analyses, should be broadly applicable to introducing therapeutic or suicide genes into patient-specific iPS cells for use in cell therapy.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
Accession codes
References
Takahashi, K. et al. Induction of pluripotent stem cells from adult human fibroblasts by defined factors. Cell 131, 861–872 (2007).
Yu, J. et al. Induced pluripotent stem cell lines derived from human somatic cells. Science 318, 1917–1920 (2007).
Park, I.H. et al. Reprogramming of human somatic cells to pluripotency with defined factors. Nature 451, 141–146 (2008).
Hanna, J. et al. Treatment of sickle cell anemia mouse model with iPS cells generated from autologous skin. Science 318, 1920–1923 (2007).
Raya, A. et al. Disease-corrected haematopoietic progenitors from Fanconi anaemia induced pluripotent stem cells. Nature 460, 53–59 (2009).
Schroder, A.R. et al. HIV-1 integration in the human genome favors active genes and local hotspots. Cell 110, 521–529 (2002).
Hacein-Bey-Abina, S. et al. LMO2-associated clonal T cell proliferation in two patients after gene therapy for SCID-X1. Science 302, 415–419 (2003).
Ott, M.G. et al. Correction of X-linked chronic granulomatous disease by gene therapy, augmented by insertional activation of MDS1–EVI1, PRDM16 or SETBP1. Nat. Med. 12, 401–409 (2006).
Howe, S.J. et al. Insertional mutagenesis combined with acquired somatic mutations causes leukemogenesis following gene therapy of SCID-X1 patients. J. Clin. Invest. 118, 3143–3150 (2008).
Cavazzana-Calvo, M. et al. Transfusion independence and HMGA2 activation after gene therapy of human beta-thalassaemia. Nature 467, 318–322 (2010).
Bejerano, G. et al. Ultraconserved elements in the human genome. Science 304, 1321–1325 (2004).
Kustikova, O. et al. Clonal dominance of hematopoietic stem cells triggered by retroviral gene marking. Science 308, 1171–1174 (2005).
May, C. et al. Therapeutic haemoglobin synthesis in beta-thalassaemic mice expressing lentivirus-encoded human beta-globin. Nature 406, 82–86 (2000).
Sadelain, M., Boulad, F., Lisowki, L., Moi, P. & Riviere, I. Stem cell engineering for the treatment of severe hemoglobinopathies. Curr. Mol. Med. 8, 690–697 (2008).
Papapetrou, E.P. et al. Stoichiometric and temporal requirements of Oct4, Sox2, Klf4, and c-Myc expression for efficient human iPSC induction and differentiation. Proc. Natl. Acad. Sci. USA 106, 12759–12764 (2009).
Chang, K.H. et al. Definitive-like erythroid cells derived from human embryonic stem cells coexpress high levels of embryonic and fetal globins with little or no adult globin. Blood 108, 1515–1523 (2006).
Qiu, C., Olivier, E.N., Velho, M. & Bouhassira, E.E. Globin switches in yolk sac-like primitive and fetal-like definitive red blood cells produced from human embryonic stem cells. Blood 111, 2400–2408 (2008).
Chang, K.H. et al. Globin phenotype of erythroid cells derived from human induced pluripotent stem cells. Blood 115, 2553–2554 (2010).
Giudice, A. & Trounson, A. Genetic modification of human embryonic stem cells for derivation of target cells. Cell Stem Cell 2, 422–433 (2008).
Hockemeyer, D. et al. Efficient targeting of expressed and silent genes in human ESCs and iPSCs using zinc-finger nucleases. Nat. Biotechnol. 27, 851–857 (2009).
Zou, J. et al. Gene targeting of a disease-related gene in human induced pluripotent stem and embryonic stem cells. Cell Stem Cell 5, 97–110 (2009).
Smith, J.R. et al. Robust, persistent transgene expression in human embryonic stem cells is achieved with AAVS1-targeted integration. Stem Cells 26, 496–504 (2008).
Irion, S. et al. Identification and targeting of the ROSA26 locus in human embryonic stem cells. Nat. Biotechnol. 25, 1477–1482 (2007).
Safaya, S., Rieder, R.F., Dowling, C.E., Kazazian, H.H. Jr. & Adams, J.G. 3rd Homozygous beta-thalassemia without anemia. Blood 73, 324–328 (1989).
Werbowetski-Ogilvie, T.E. et al. Characterization of human embryonic stem cells with features of neoplastic progression. Nat. Biotechnol. 27, 91–97 (2009).
Ji, J. et al. OP9 stroma augments survival of hematopoietic precursors and progenitors during hematopoietic differentiation from human embryonic stem cells. Stem Cells 26, 2485–2495 (2008).
Deichmann, A. et al. Vector integration is nonrandom and clustered and influences the fate of lymphopoiesis in SCID-X1 gene therapy. J. Clin. Invest. 117, 2225–2232 (2007).
Aiuti, A. et al. Multilineage hematopoietic reconstitution without clonal selection in ADA-SCID patients treated with stem cell gene therapy. J. Clin. Invest. 117, 2233–2240 (2007).
Schwarzwaelder, K. et al. Gammaretrovirus-mediated correction of SCID-X1 is associated with skewed vector integration site distribution in vivo. J. Clin. Invest. 117, 2241–2249 (2007).
Lee, G. et al. Modelling pathogenesis and treatment of familial dysautonomia using patient-specific iPSCs. Nature 461, 402–406 (2009).
Papapetrou, E.P., Kovalovsky, D., Beloeil, L., Sant'angelo, D. & Sadelain, M. Harnessing endogenous miR-181a to segregate transgenic antigen receptor expression in developing versus post-thymic T cells in murine hematopoietic chimeras. J. Clin. Invest. 119, 157–168 (2009).
Saenz, D.T. et al. Unintegrated lentivirus DNA persistence and accessibility to expression in nondividing cells: analysis with class I integrase mutants. J. Virol. 78, 2906–2920 (2004).
Watanabe, K. et al. A ROCK inhibitor permits survival of dissociated human embryonic stem cells. Nat. Biotechnol. 25, 681–686 (2007).
Papapetrou, E.P., Ziros, P.G., Micheva, I.D., Zoumbos, N.C. & Athanassiadou, A. Gene transfer into human hematopoietic progenitor cells with an episomal vector carrying an S/MAR element. Gene Ther. 13, 40–51 (2006).
Schmidt, M. et al. High-resolution insertion-site analysis by linear amplification-mediated PCR (LAM-PCR). Nat. Methods 4, 1051–1057 (2007).
Wang, G.P., Ciuffi, A., Leipzig, J., Berry, C.C. & Bushman, F.D. HIV integration site selection: analysis by massively parallel pyrosequencing reveals association with epigenetic modifications. Genome Res. 17, 1186–1194 (2007).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Papapetrou, E.P., Korkola, J.E. & Sadelain, M. A genetic strategy for single and combinatorial analysis of miRNA function in mammalian hematopoietic stem cells. Stem Cells 28, 287–296 (2009).
Huber, W., von Heydebreck, A., Sueltmann, H., Poustka, A. & Vingron, M. Parameter estimation for the calibration and variance stabilization of microarray data. Stat. Appl. Genet. Mol. Biol. 2, Article 3 (2003).
Smyth, G.K. Linear models and empirical bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, Article 3 (2004).
Venkatraman, E.S. & Olshen, A.B. A faster circular binary segmentation algorithm for the analysis of array CGH data. Bioinformatics 23, 657–663 (2007).
Acknowledgements
We thank X. Wang and N. Wu for assistance with HPLC analysis; L. Ferro, E. Reed, J. Miller, M. Leversha and M. Tomishima for technical assistance; F. Boulad, Memorial Sloan-Kettering Cancer Center New York for bone marrow specimens; and A. Athanassiadou for advice on β-thalassemia genotyping. pCMVΔR8.91N/N was kindly provided by E. Poeschla, Mayo Clinic, Rochester, Minnesota. This work was supported by the Starr Foundation (Tri-Institutional Stem Cell Initiative, Tri-SCI-018), the New York State Stem Cell Science, NYSTEM (N08T-060) and National Heart, Blood, and Lung Institute grant HL053750 (M.S.). F.D.B., S.L.R. and N.M. were supported by National Institutes of Health grants AI052845 and AI082020 (F.D.B.). G.L. was supported by a New York Stem Cell Foundation Druckenmiller fellowship.
Author information
Authors and Affiliations
Contributions
E.P.P. conceived and designed the study, designed and performed experiments, analyzed data and wrote the manuscript; G.L. performed iPS cell differentiation experiments; N.M. performed bioinformatics analyses; M.S. and C.L. analyzed microarray data; L.M.S.T. provided technical assistance; K.K. performed histological analyses of teratomas; S.L.R. generated and analyzed integration site data; P.G. provided skin biopsy samples from β-thalassemia patients; A.V. generated microarray data; I.R., F.D.B. and L.S. analyzed data; M.S. conceived and designed the study, analyzed data and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Tables 1–9 and Supplementary Figs. 1–20 (PDF 3117 kb)
Supplementary Movie 1
Beating putative cardiomyocytes derived from iPS cell line thal1.52. (MOV 20336 kb)
Rights and permissions
About this article
Cite this article
Papapetrou, E., Lee, G., Malani, N. et al. Genomic safe harbors permit high β-globin transgene expression in thalassemia induced pluripotent stem cells. Nat Biotechnol 29, 73–78 (2011). https://doi.org/10.1038/nbt.1717
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nbt.1717
This article is cited by
-
Integrating Omics and CRISPR Technology for Identification and Verification of Genomic Safe Harbor Loci in the Chicken Genome
Biological Procedures Online (2023)
-
Site-specific transgene integration in chimeric antigen receptor (CAR) T cell therapies
Biomarker Research (2023)
-
PERSIST platform provides programmable RNA regulation using CRISPR endoRNases
Nature Communications (2022)
-
HLA-independent T cell receptors for targeting tumors with low antigen density
Nature Medicine (2022)
-
Parkinson’s disease patient-specific neuronal networks carrying the LRRK2 G2019S mutation unveil early functional alterations that predate neurodegeneration
npj Parkinson's Disease (2021)