Abstract
A key limitation of the use of the CRISPR–Cas9 system for genome editing and other applications is the requirement that a protospacer adjacent motif (PAM) be present at the target site. For the most commonly used Cas9 from Streptococcus pyogenes (SpCas9), the required PAM sequence is NGG. No natural or engineered Cas9 variants that have been shown to function efficiently in mammalian cells offer a PAM less restrictive than NGG. Here we use phage-assisted continuous evolution to evolve an expanded PAM SpCas9 variant (xCas9) that can recognize a broad range of PAM sequences including NG, GAA and GAT. The PAM compatibility of xCas9 is the broadest reported, to our knowledge, among Cas9 proteins that are active in mammalian cells, and supports applications in human cells including targeted transcriptional activation, nuclease-mediated gene disruption, and cytidine and adenine base editing. Notably, despite its broadened PAM compatibility, xCas9 has much greater DNA specificity than SpCas9, with substantially lower genome-wide off-target activity at all NGG target sites tested, as well as minimal off-target activity when targeting genomic sites with non-NGG PAMs. These findings expand the DNA targeting scope of CRISPR systems and establish that there is no necessary trade-off between Cas9 editing efficiency, PAM compatibility and DNA specificity.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Doudna, J. A. & Charpentier, E. Genome editing. The new frontier of genome engineering with CRISPR–Cas9. Science 346, 1258096 (2014)
Hsu, P. D., Lander, E. S. & Zhang, F. Development and applications of CRISPR–Cas9 for genome engineering. Cell 157, 1262–1278 (2014)
Mitsunobu, H., Teramoto, J., Nishida, K. & Kondo, A. Beyond native Cas9: manipulating genomic information and function. Trends Biotechnol. 35, 983–996 (2017)
Komor, A. C., Kim, Y. B., Packer, M. S., Zuris, J. A. & Liu, D. R. Programmable editing of a target base in genomic DNA without double-stranded DNA cleavage. Nature 533, 420–424 (2016)
Gaudelli, N. M. et al. Programmable base editing of A•T to G•C in genomic DNA without DNA cleavage. Nature 551, 464–471 (2017)
Komor, A. C., Badran, A. H. & Liu, D. R. CRISPR-based technologies for the manipulation of eukaryotic genomes. Cell 168, 20–36 (2017)
Jinek, M. et al. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337, 816–821 (2012)
Findlay, G. M., Boyle, E. A., Hause, R. J., Klein, J. C. & Shendure, J. Saturation editing of genomic regions by multiplex homology-directed repair. Nature 513, 120–123 (2014)
Yang, L. et al. Optimization of scarless human stem cell genome editing. Nucleic Acids Res. 41, 9049–9061 (2013)
Ran, F. A. et al. In vivo genome editing using Staphylococcus aureus Cas9. Nature 520, 186–191 (2015)
Zetsche, B. et al. Cpf1 is a single RNA-guided endonuclease of a class 2 CRISPR–Cas system. Cell 163, 759–771 (2015)
Kim, E. et al. In vivo genome editing with a small Cas9 orthologue derived from Campylobacter jejuni. Nat. Commun. 8, 14500 (2017)
Müller, M. et al. Streptococcus thermophilus CRISPR–Cas9 systems enable specific editing of the human genome. Mol. Ther. 24, 636–644 (2016)
Lee, C. M., Cradick, T. J. & Bao, G. The Neisseria meningitidis CRISPR–Cas9 system enables specific genome editing in mammalian cells. Mol. Ther. 24, 645–654 (2016)
Kleinstiver, B. P. et al. Broadening the targeting range of Staphylococcus aureus CRISPR–Cas9 by modifying PAM recognition. Nat. Biotechnol. 33, 1293–1298 (2015)
Kleinstiver, B. P. et al. Engineered CRISPR–Cas9 nucleases with altered PAM specificities. Nature 523, 481–485 (2015)
Badran, A. H. & Liu, D. R. Development of potent in vivo mutagenesis plasmids with broad mutational spectra. Nat. Commun. 6, 8425 (2015)
Esvelt, K. M., Carlson, J. C. & Liu, D. R. A system for the continuous directed evolution of biomolecules. Nature 472, 499–503 (2011)
Badran, A. H. et al. Continuous evolution of Bacillus thuringiensis toxins overcomes insect resistance. Nature 533, 58–63 (2016)
Hubbard, B. P. et al. Continuous directed evolution of DNA-binding proteins to improve TALEN specificity. Nat. Methods 12, 939–942 (2015)
Bryson, D. I. et al. Continuous directed evolution of aminoacyl-tRNA synthetases. Nat. Chem. Biol. 13, 1253–1260 (2017)
Packer, M. S., Rees, H. A. & Liu, D. R. Phage-assisted continuous evolution of proteases with altered substrate specificity. Nat. Commun. 8, 956 (2017)
Meng, X. & Wolfe, S. A. Identifying DNA sequences recognized by a transcription factor using a bacterial one-hybrid system. Nat. Protocols 1, 30–45 (2006)
Dove, S. L., Joung, J. K. & Hochschild, A. Activation of prokaryotic transcription through arbitrary protein-protein contacts. Nature 386, 627–630 (1997)
Anders, C., Niewoehner, O., Duerst, A. & Jinek, M. Structural basis of PAM-dependent target DNA recognition by the Cas9 endonuclease. Nature 513, 569–573 (2014)
Chen, J. S. et al. Enhanced proofreading governs CRISPR–Cas9 targeting accuracy. Nature 550, 407–410 (2017)
Sternberg, S. H., LaFrance, B., Kaplan, M. & Doudna, J. A. Conformational control of DNA target cleavage by CRISPR–Cas9. Nature 527, 110–113 (2015)
Gao, L. et al. Engineered Cpf1 variants with altered PAM specificities. Nat. Biotechnol. 35, 789–792 (2017)
Tsai, S. Q. et al. GUIDE-seq enables genome-wide profiling of off-target cleavage by CRISPR–Cas nucleases. Nat. Biotechnol. 33, 187–197 (2015)
Kim, S. et al. Rescue of high-specificity Cas9 variants using sgRNAs with matched 5′ nucleotides. Genome Biology 18, 218 (2017)
Chavez, A. et al. Highly efficient Cas9-mediated transcriptional programming. Nat. Methods 12, 326–328 (2015)
Doench, J. G. et al. Rational design of highly active sgRNAs for CRISPR–Cas9-mediated gene inactivation. Nat. Biotechnol. 32, 1262–1267 (2014)
Zhang, Y. et al. Comparison of non-canonical PAMs for CRISPR/Cas9-mediated DNA cleavage in human cells. Sci. Rep. 4, 5405 (2014)
Kim, Y. B. et al. Increasing the genome-targeting scope and precision of base editing with engineered Cas9-cytidine deaminase fusions. Nat. Biotechnol. 35, 371–376 (2017)
Komor, A. C. et al. Improved base excision repair inhibition and bacteriophage Mu Gam protein yields C:G-to-T:A base editors with higher efficiency and product purity. Sci. Adv. 3, eaao4774 (2017)
Pattanayak, V., Ramirez, C. L., Joung, J. K. & Liu, D. R. Revealing off-target cleavage specificities of zinc-finger nucleases by in vitro selection. Nat. Methods 8, 765–770 (2011)
Slaymaker, I. M. et al. Rationally engineered Cas9 nucleases with improved specificity. Science 351, 84–88 (2016)
Kleinstiver, B. P. et al. High-fidelity CRISPR–Cas9 nucleases with no detectable genome-wide off-target effects. Nature 529, 490–495 (2016)
Landrum, M. J. et al. ClinVar: public archive of relationships among sequence variation and human phenotype. Nucleic Acids Res. 42, D980–D985 (2014)
Crooks, G. E., Hon, G., Chandonia, J. M. & Brenner, S. E. WebLogo: a sequence logo generator. Genome Res. 14, 1188–1190 (2004)
Acknowledgements
We thank A. Badran, B. Hubbard, J. Levy, T. Huang, G. Church, A. Chavez, K. Esvelt, S. Vora and J. Scheiman for discussions. This work was supported by DARPA HR0011-17-2-0049, US NIH RM1 HG009490, R01 EB022376 and R35 GM118062 and the HHMI. J.H.H. was supported by NDSEG and NSF graduate fellowships. S.M.M. was supported by an NSF graduate fellowship. W.T. is an HHMI Fellow of the Jane Coffin Childs Memorial Fund. L.C. was supported by the Agency for Science, Technology, and Research, Singapore.
Author information
Authors and Affiliations
Contributions
J.H.H. designed the research, performed PACE, characterized variants in bacteria, conducted human cell experiments, analysed data, performed off-target analysis and wrote the manuscript. S.M.M. performed human cell experiments, analysed data and wrote the manuscript. M.H.G. performed human cell experiments and data analysis. W.T. performed human cell experiments and cloning. L.C. and C.M.Z. assisted with cloning and PACE. N.S. optimized Cas9 PACE. X.G. assisted with off-target analysis. H.A.R. assisted with indel and base-editing analyses. Z.L. assisted with human cell experiments. D.R.L. designed and supervised the research and wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
J.H.H. and D.R.L. have filed patent applications on this work (WO2017070633). D.R.L. is a consultant and co-founder of Editas Medicine, Beam Therapeutics, and Pairwise Plants, companies that use genome editing technologies. The authors declare no other competing interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Optimization of Cas9 PACE.
Luciferase expression in E. coli was used as a proxy of gene III expression levels during efforts to link Cas9 binding to gene expression for PACE. a, b, Seven guide RNAs targeting the luciferase reporter (G1–G7′, see Supplementary Table 1), as well as a scrambled guide RNA negative control (G0) were tested without dCas9 (white bars) and with ω–dCas9 (a) or dCas9–ω (b) fusions (grey bars). c, Tests of seven different linkers between ω and dCas9. See Supplementary Table 2 for linker sequences. d, Evolution of ω–dCas9 on an NGG PAM site in PACE yielded variants (PACE1, PACE2 and PACE3) that were tested in comparison with canonical wild-type (wt) ω–dCas9, ω tethered to PACE1 dCas9 and the I12N ω mutant tethered to canonical wild-type dCas9. a–d, Data are mean and s.d. of three biologically independent samples.
Extended Data Figure 2 PAM profiling of xCas9 variants.
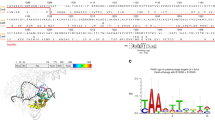
a, In separate experiments, a plasmid library containing a protospacer with all possible NNN PAM sequences and a spectinomycin-resistance gene was electroporated into E. coli along with a plasmid expressing SpCas9 or the xCas9 variant shown. PAMs that are cleaved are depleted from the library when plated on medium containing spectinomycin. HTS of the library before and after selection enables quantification of the change in library composition, resulting in a sequence logo40 for the PAM preference of SpCas9 (left) and xCas9-3.7 (right). b, c, PAM depletion scores of Cas9 variants from spectinomycin selection in E. coli, calculated as described previously36, with 1.0 representing complete cleavage of that PAM sequence. Scores for NGN, NNG, GAA, GAT and CAA are shown in b and the rest of the PAM sequences are shown in c.
Extended Data Figure 3 Transcriptional activation of reporter site PAM libraries with xCas9.
Transcriptional activation by dSpCas9–VPR and dxCas9–VPR variants (transfected as plasmids) on GFP reporter plasmids containing different PAM sites in HEK293T cells. a, Earlier generations of xCas9 variants were tested on the R1 site with the NGG, NNN or NNNNN PAM libraries. b, c, Transcriptional activators dxCas9(3.6)–VPR and dxCas9(3.7)–VPR were tested on two different protospacer reporters, R1 (a) and R2 (b), containing adjacent NGG, NNN, or NNNNN PAM libraries. a–c, Data are mean and s.d. of three biologically independent samples.
Extended Data Figure 4 Transcriptional activation with xCas9-2.0.
a–d, The transcriptional activator dxCas9(2.0)–VPR was tested on the R1 protospacer (Extended Data Fig. 3 and Supplementary Table 8) with each of the 64 possible NNN PAMs (NAN, NCN, NGN and NTN) in HEK293T cells. Data are mean and s.d. of three biologically independent samples.
Extended Data Figure 5 Transcriptional activation with xCas9-3.7 on all 64 NNN PAM sites and endogenous gene activation in human cells.
a–d, The transcriptional activator dxCas9(3.7)–VPR was tested on the R1 protospacer (Extended Data Fig. 3 and Supplementary Table 8) with each of the 64 possible NNN PAMs (NAN, NCN, NGN and NTN) in HEK293T cells. e, Endogenous gene activation was tested using both dSpCas9–VPR and dxCas9(3.7)–VPR to activate expression of the NEUROD1, ASCL1, MIAT or RHOXF2 at six total sites. RNA expression as measured by qPCR with reverse transcription (RT–qPCR) was compared to background expression levels for each gene (measured in the control with no sgRNA) and was normalized to ACTB expression. a–e, Data are mean and s.d. of three biologically independent samples. Target sites are listed in Supplementary Table 9.
Extended Data Figure 6 Transcriptional activation with xCas9-3.6 on all 64 NNN PAM sites.
a–d, The transcriptional activator dxCas9(3.6)–VPR was tested on the R1 protospacer (Extended Data Fig. 3 and Supplementary Table 8) with each of the 64 possible NNN PAMs (NAN, NCN, NGN and NTN) in HEK293T cells. Data are mean and s.d. of three biologically independent samples.
Extended Data Figure 7 Genomic DNA cleavage and base editing by evolved xCas9-3.6.
a, Genomic DNA cleavage by SpCas9 or xCas9-3.6 (transfected as plasmids), in HEK293-GFP cells containing a genomically integrated GFP gene. After 5 days, the cells were analysed for loss of GFP fluorescence by flow cytometry. Sequences for all target sites are listed in Supplementary Table 11. b, DNA cleavage of endogenous genomic DNA sites with a variety of NGG and non-NGG PAMs by SpCas9 and xCas9-3.6 in HEK293T cells. Indel rates were measured using HTS 5 days after plasmid transfection. Sequences for all target sites are listed in Supplementary Table 12. c, C•G-to-T•A base editing in HEK293T cells by SpCas9–BE3 or xCas9(3.6)–BE3 was tested at 20 sites containing NG, GAA, GAT, or CAA PAM sites. The C•G-to-T•A conversion frequency, ascertained using HTS, at the most efficiently edited base 3 days after plasmid transfection is shown. d, Of the 20 sites in c, seven contained an A in the canonical window for ABE editing5 and were tested for A•T-to-G•C base editing by SpCas9–ABE and xCas9(3.6)–ABE. The A•T-to-G•C conversion frequency, ascertained using HTS, at the most efficiently edited base 5 days after plasmid transfection is shown. a–d, Data are mean and s.d. of three biologically independent samples. Complete HTS results across the protospacer are provided in Supplementary Table 14.
Extended Data Figure 8 Negative controls lacking sgRNA for nuclease and base editing experiments.
To verify genomic DNA cleavage and base editing results, the same sites were sequenced after treatment with SpCas9 nuclease, SpCas9–BE3, or SpCas9–ABE but without any sgRNA. a, Indel rates at endogenous target sites 5 days after treatment of HEK293T cells with SpCas9. b, Target C•G-to-T•A conversion 3 days after treatment of HEK293T cells with SpCas9–BE3. c, Target A•T-to-G•C base conversion 5 days after treatment of HEK293T cells with SpCas9–ABE. a–c, Data are mean and s.d. of three biologically independent samples. Complete HTS results across the protospacer are provided in Supplementary Table 14.
Extended Data Figure 9 Cytidine base editing at 15 additional genomic sites and xCas9 base editing with the BE4 architecture.
a, Base editing by SpCas9–BE3 and xCas9(3.7)–BE3 at 15 sites within the FANCF gene in HEK293T cells. The C•G-to-T•A conversion frequency at the most efficiently edited base 3 days after plasmid transfection is shown. b, Test of xCas9-3.7 in the BE4 architecture35 on the same sites tested in Fig. 3. The C•G-to-T•A conversion frequency in HEK293T cells at the most efficiently edited base 3 days after plasmid transfection is shown. c, d, Indel frequency following treatment with BE3 or BE4 variants targeting sites with NGG PAMs (c) and non-NGG PAMs (d). e, f, Product distribution among edited DNA sequence reads (reads in which the target C is mutated) following treatment with BE3 or BE4 variants targeting sites with NGG PAMs (e) and non-NGG PAMs (f). Because SpCas9 has minimal activity on non-NGG PAM sites, only xCas9(3.7)–BE3 and xCas9(3.7)–BE4 data are compared on non-NGG PAM sites. a–f, Data are mean and s.d. of three biologically independent samples. Target sites are listed in Supplementary Table 12.
Extended Data Figure 10 Additional characterization of xCas9-3.7 and xCas9-3.6 by GUIDE-seq.
a–d, In addition to the GUIDE-seq data shown in Fig. 4, two additional sites in HEK293T cells (a, b) and two sites in U2OS cells (c, d) were analysed after treatment with SpCas9 and xCas9-3.7. e, All six GUIDE-seq sites with an NGG PAM that were tested with SpCas9 and xCas9-3.7 in HEK293T and U2OS cells were also tested with xCas9-3.6. On-target reads (indicated by an asterisk) and off-target reads for all sites are shown. Target sequences are listed in Supplementary Table 15.
Extended Data Figure 11 Validation by high-throughput sequencing of GUIDE-seq results.
a–d, The most frequent off-target sites identified using GUIDE-seq were verified by HTS of genomic DNA following treatment of HEK293T cells with SpCas9 or xCas9-3.7 (a, b), or following treatment with SpCas9 or xCas9-3.6 (c, d). New sites with non-NGG PAMs that were identified as off-target sites of the xCas9 proteins were also analysed in a–d. a–d, Data are mean and s.d. of three biologically independent samples. Target sequences are listed in Supplementary Table 16.
Supplementary information
Supplementary Information
This file contains the data for Supplementary Tables 1-17, Supplementary Sequences 1-3 and Supplementary Notes 1-3. (PDF 1003 kb)
Rights and permissions
About this article
Cite this article
Hu, J., Miller, S., Geurts, M. et al. Evolved Cas9 variants with broad PAM compatibility and high DNA specificity. Nature 556, 57–63 (2018). https://doi.org/10.1038/nature26155
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature26155
This article is cited by
-
Enhancing prime editor flexibility with coiled-coil heterodimers
Genome Biology (2024)
-
Recent advances in CRISPR-based functional genomics for the study of disease-associated genetic variants
Experimental & Molecular Medicine (2024)
-
Phage-assisted evolution of compact Cas9 variants targeting a simple NNG PAM
Nature Chemical Biology (2024)
-
Discovery and structural mechanism of DNA endonucleases guided by RAGATH-18-derived RNAs
Cell Research (2024)
-
Unraveling the mechanisms of PAMless DNA interrogation by SpRY-Cas9
Nature Communications (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.