Abstract
Histone modifications, such as the frequently occurring lysine succinylation1,2, are central to the regulation of chromatin-based processes. However, the mechanism and functional consequences of histone succinylation are unknown. Here we show that the α-ketoglutarate dehydrogenase (α-KGDH) complex is localized in the nucleus in human cell lines and binds to lysine acetyltransferase 2A (KAT2A, also known as GCN5) in the promoter regions of genes. We show that succinyl-coenzyme A (succinyl-CoA) binds to KAT2A. The crystal structure of the catalytic domain of KAT2A in complex with succinyl-CoA at 2.3 Å resolution shows that succinyl-CoA binds to a deep cleft of KAT2A with the succinyl moiety pointing towards the end of a flexible loop 3, which adopts different structural conformations in succinyl-CoA-bound and acetyl-CoA-bound forms. Site-directed mutagenesis indicates that tyrosine 645 in this loop has an important role in the selective binding of succinyl-CoA over acetyl-CoA. KAT2A acts as a succinyltransferase and succinylates histone H3 on lysine 79, with a maximum frequency around the transcription start sites of genes. Preventing the α-KGDH complex from entering the nucleus, or expression of KAT2A(Tyr645Ala), reduces gene expression and inhibits tumour cell proliferation and tumour growth. These findings reveal an important mechanism of histone modification and demonstrate that local generation of succinyl-CoA by the nuclear α-KGDH complex coupled with the succinyltransferase activity of KAT2A is instrumental in histone succinylation, tumour cell proliferation, and tumour development.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Weinert, B. T. et al. Lysine succinylation is a frequently occurring modification in prokaryotes and eukaryotes and extensively overlaps with acetylation. Cell Reports 4, 842–851 (2013)
Zhang, Z. et al. Identification of lysine succinylation as a new post-translational modification. Nat. Chem. Biol. 7, 58–63 (2011)
Yang, W. et al. Nuclear PKM2 regulates β-catenin transactivation upon EGFR activation. Nature 480, 118–122 (2011)
Jiang, Y. et al. Local generation of fumarate promotes DNA repair through inhibition of histone H3 demethylation. Nat. Cell Biol. 17, 1158–1168 (2015)
Sutendra, G. et al. A nuclear pyruvate dehydrogenase complex is important for the generation of acetyl-CoA and histone acetylation. Cell 158, 84–97 (2014)
Wang, L. & Dent, S. Y. Functions of SAGA in development and disease. Epigenomics 6, 329–339 (2014)
Grant, P. A. et al. Yeast Gcn5 functions in two multisubunit complexes to acetylate nucleosomal histones: characterization of an Ada complex and the SAGA (Spt/Ada) complex. Genes Dev. 11, 1640–1650 (1997)
Nguyen Ba, A. N., Pogoutse, A., Provart, N. & Moses, A. M. NLStradamus: a simple Hidden Markov Model for nuclear localization signal prediction. BMC Bioinformatics 10, 202 (2009)
Schuetz, A. et al. Crystal structure of a binary complex between human GCN5 histone acetyltransferase domain and acetyl coenzyme A. Proteins 68, 403–407 (2007)
Lin, Y., Fletcher, C. M., Zhou, J., Allis, C. D. & Wagner, G. Solution structure of the catalytic domain of GCN5 histone acetyltransferase bound to coenzyme A. Nature 400, 86–89 (1999)
Clements, A. et al. Structural basis for histone and phosphohistone binding by the GCN5 histone acetyltransferase. Mol. Cell 12, 461–473 (2003)
Poux, A. N. & Marmorstein, R. Molecular basis for Gcn5/PCAF histone acetyltransferase selectivity for histone and nonhistone substrates. Biochemistry 42, 14366–14374 (2003)
Poux, A. N., Cebrat, M., Kim, C. M., Cole, P. A. & Marmorstein, R. Structure of the GCN5 histone acetyltransferase bound to a bisubstrate inhibitor. Proc. Natl Acad. Sci. USA 99, 14065–14070 (2002)
Ringel, A. E. & Wolberger, C. Structural basis for acyl-group discrimination by human Gcn5L2. Acta Crystallogr. D 72, 841–848 (2016)
Dyda, F., Klein, D. C. & Hickman, A. B. GCN5-related N-acetyltransferases: a structural overview. Annu. Rev. Biophys. Biomol. Struct. 29, 81–103 (2000)
Kaczmarska, Z. et al. Structure of p300 in complex with acyl-CoA variants. Nat. Chem. Biol. 13, 21–29 (2017)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Subramanian, A. et al. Gene set enrichment analysis: a knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl Acad. Sci. USA 102, 15545–15550 (2005)
Acknowledgements
We thank L. Li at the University of Texas Health Science at Houston for technical support and D. Norwood and T. Locke for critical reading of this manuscript. This work was supported by National Institute of Neurological Disorders and Stroke grant R01 NS089754 (Z.L.), National Cancer Institute grants 2R01 CA109035 (Z.L.), R01 CA169603 (Z.L.), MD Anderson Support Grant P30CA016672, Welch Foundation grant C-1565 (Y.J.T.), and the National Institutes of Health Brain Cancer Specialized Program of Research Excellence (2P50 CA127001). Z.L. is a Ruby E. Rutherford Distinguished Professor.
Author information
Authors and Affiliations
Contributions
Z.L., Y.J.T., D.X., and Y.W. conceived and designed the study and wrote the manuscript. Y.W., R.L., Y.X., Y.Z., and J.L. performed the biochemistry experiments to identify the succinyltransferase activity of KAT2A. Y.R.G., Y.W., and Y.J.T. generated co-crystal structures of KAT2A and Co-enzyme A analogues. Z.Y. and K.L. performed bioinformatics analyses. L.T. and P.Y. performed quantification analyses of acetyl-CoA and succinyl-CoA in cells. X.L. identified the nuclear localization sequence on DLST. X.Q. performed animal tissue studies. D.H. identified KAT2A-mediated histone H3 succinylation residue. P.Y., J.-H.L., D.X., J.L., and J.H. provided technical support.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks J. Denu, C. Frezza and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 KAT2A interacts with the α-KGDH complex in the nucleus.
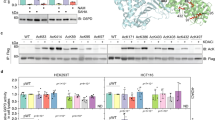
a, OGDH binds to KAT2A. KAT2A was immunoprecipitated from U251 cells using an anti-KAT2A antibody. The immunoprecipitate was analysed by mass spectrometry. A specific peptide (920-DMVGQVAIT-929) of OGDH was identified. The IonScore is 30, the q value is 0.004, and the Exp value (P) is 0.1792. One-sided t-tests were conducted. Representative results of two experiments are shown. b, Quantitative analyses of α-KGDH subunits in the nucleus. Immunoblotting analyses of α-KGDH subunits in the nucleus extraction of U251 cells were performed with the indicated antibodies displayed in Fig. 1b. The quantitative standard curves were used to determine the percentage of nuclear OGDH, DLST, and DLD. The x-axis presents the intensities of quantified proteins detected by immunoblotting analysis, and the y-axis presents the percentage of whole-cell lysate corresponding to each quantified protein. The percentages of nuclear OGDH, DLST, and DLD were calculated using the equations displayed above the standard curves. Representative images of triplicate experiments are shown. The R2 values represent the relationship between logY and X to the linear model (logY = log[A + B] × X). Y, percentage of whole cell lysate; X, intensity. c, Nuclear distribution of the α-KGDH complex. Immunofluorescent staining of U251 cells was performed with anti-OGDH, anti-DLST, or anti-DLD antibodies (green). The mitochondria and nuclei were stained with an anti-Tom20 antibody (red) and DAPI (blue), respectively. Representative images of more than 50 cells are shown. The majority of the α-KGDH complex overlapped with the mitochondrial protein Tom20, indicating primary mitochondrial localization of the α-KGDH complex. d, Quantitative analyses of the interaction between KAT2A and α-KGDH subunits. Immunoprecipitation analyses of U251 cell nuclei with anti-KAT2A or anti-OGDH antibodies were performed. KAT2A- or OGDH-associated proteins were analysed via immunoblotting analyses. Representative images of triplicate experiments are shown. e, f, KAT2A is not localized in mitochondria. e, Immunoblotting analyses of the mitochondria extraction and total cell lysate of U251 cells were performed with the indicated antibodies. Representative images of triplicate experiments are shown. f, Immunofluorescent analyses of U251 cells were performed with DAPI (blue) and anti-KAT2A (green) and anti-TOM20 (red) antibodies. Representative images of more than 50 cells are shown. g, OGDH depletion, which does not affect the interaction between DLST and DLD, inhibits the binding of DLST to KAT2A. OGDH was depleted in U251 cells. Immunoprecipitation analyses with an anti-DLST antibody were followed by immunoblotting analyses with the indicated antibodies. Representative images of triplicate experiments are shown. h, i, The NLS of DLST is required for nuclear distribution of the α-KGDH complex. DLST was depleted from U87 cells, which were reconstituted with expression of wild-type rDLST or the rDLST(R224A/K226E) NLS mutant. h, Immunofluorescent staining was performed with an anti-DLST or anti-OGDH antibody. The mitochondria and nuclei were stained with an anti-Tom20 antibody (red) and DAPI (blue), respectively. i, Immunoblotting analyses were performed with the indicated antibodies. Mitochondrial COX IV and IDH2 and nuclear PCNA were used to show lack of mitochondrial contamination of nuclear fractions and loading controls. Representative images of triplicate experiments are shown.
Extended Data Figure 2 KAT2A associates with α-KGDH and acts as a histone succinyltransferase.
a, rDLST NLS mutation does not affect the enzymatic activity of α-KGDH. Wild-type His–rDLST or His–rDLST NLS mutant expressed in 293 cells was precipitated using Ni-NTA agarose beads. The enzymatic activity of α-KGDH was measured. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from five independent experiments (n = 5). b, rDLST NLS mutant expression does not affect cellular levels of acetyl-CoA or succinyl-CoA. The levels of acetyl-CoA and succinyl-CoA in U251 cells with depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant were measured by LC–MS/MS. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from three independent experiments (n = 3). c, rDLST NLS mutant expression does not affect total cellular protein succinylation. Lysine succinylation of whole-cell lysate of U251 cells expressing wild-type V5–rDLST or V5–rDLST NLS mutant was analysed by immunoblotting analysis with an anti-succinyl-lysine antibody. The protein levels of each sample were shown by Ponceau S staining. Representative images of triplicate experiments are shown. d, Quantification of the colocalization of KAT2A, α-KGDH complex, and histone H3. Immunofluorescent analyses were performed with anti-DLST (red), anti-KAT2A (blue), and anti-histone H3 (green) antibodies. The Pearson’s R value is used to present the extent of co-localization of two different targets. The random co-localization tests were conducted by 90° clockwise rotation of KAT2A (blue channel) and 180° clockwise rotation of histone H3 (green channel). Two-sided t-tests were conducted. Statistical analysis for 79 individual cells (n = 79) was performed. *P < 0.05. e, KAT2A-mediated histone H3 succinylation is inhibited by CoA. KAT2A-mediated histone H3 succinylation was analysed by mixing purified KAT2A, histone H3, and succinyl-CoA (2 μM) with or without addition of the indicated concentrations of CoA. Representative images of triplicate experiments are shown. f, The chemical structures of acetyl-CoA and succinyl-CoA. The various acyl tail groups in acetyl-CoA and succinyl-CoA are highlighted in blue. g, Comparison of the structures of the catalytic domain of KAT2A in complex with different acyl-CoA ligands. The four structures shown from left to right are: apo structure of human KAT2A (this study), acetyl-CoA complex of human KAT2A (PDB ID 1Z4R), succinyl-CoA complex of human KAT2A (this study), and the bisubstrate analogue complex of Tetrahymena thermophila KAT2A (PDB ID 1M1D). For clarity, only residues surrounding the acyl groups are shown. The CoA substrate molecules are shown as yellow sticks. Hydrogen bonds are shown as dotted lines. h, KAT2A-mediated histone H3 succinylation is moderately affected by acetyl-CoA. KAT2A-mediated histone H3 succinylation was analysed by mixing purified KAT2A, histone H3, and succinyl-CoA (2 μM) with or without acetyl-CoA (2 μM). Immunoblotting analyses were performed with the indicated antibodies. Representative images of triplicate experiments are shown. i, j, The steady-state kinetics of KAT2A-mediated histone H3 modifications. Immunoprecipitated wild-type Flag–rKAT2A or Flag–rKAT2A(Y645A) from 293 cells was incubated with purified histone H3 in the presence of acetyl-CoA or succinyl-CoA. The steady-state kinetics of KAT2A activity was analysed by measuring CoA production. The Km values of wild-type Flag–rKAT2A for histone H3 acetylation and succinylation were 0.83 ± 0.13 μM (means ± s.d.) and 0.36 ± 0.04 μM, respectively. The Km values of Flag–rKAT2A(Y645A) for histone H3 acetylation and succinylation were 0.79 ± 0.08 μM and 1.73 ± 0.29 μM, respectively. The steady-state kinetic curves show the mean values from three independent measurements (i, n = 3). The Vmax values of wild-type Flag–rKAT2A for histone H3 acetylation and succinylation were 1.56 ± 0.07 nM s−1 (mean ± s.d.) and 1.80 ± 0.06 nM s−1, respectively. The Vmax values of Flag–rKAT2A(Y645A) for histone H3 acetylation and succinylation were 1.50 ± 0.05 nM s−1 and 0.87 ± 0.09 nM s−1, respectively (j). Two-sided t-tests were conducted. The data are presented as means ± s.d. from three independent experiments (n = 3). *P < 0.05. k, The KAT2A binding pocket for succinyl-CoA. The structurally conserved core regions of the HAT domains are shown in blue, whereas the flanking N- and C-terminal regions are shown in grey. The two enlarged panels, viewed with 90° rotation, display the van der Waals surface of KAT2A in grey around the succinyl group of succinyl-CoA. The cofactors are shown as follows: carbon in yellow, oxygen in red, nitrogen in blue, and sulfur in light brown. The alternative pockets are marked by dotted circles. The polypeptide regions surrounding the two extra pockets are highlighted in magenta and cyan, respectively. l, KAT2A, but not EP300, succinylates histone H3 in vitro. Purified histone H3 and succinyl-CoA were mixed with immunoprecipitated Flag–KAT2A or Flag–EP300 from 293 cells. Immunoblotting analyses were performed with the indicated antibodies. Representative images of triplicate experiments are shown.
Extended Data Figure 3 KAT2A coupled with the α-KGDH complex regulates H3K79 succinylation and gene expression.
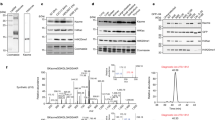
a, H3K79 is succinylated. Purified wild-type or mutated histone H3 was incubated with purified KAT2A and succinyl-CoA. Mass spectrometric analysis of a tryptic fragment of histone H3 at monoisotopic m/z 718.35706 Da (+0.21 milli-mass unit (m.m.u.)/+0.29 p.p.m.) matched with the doubly charged peptide 73-EIAQDFKTDLR-83 with K7-succinyl (100.01604 Da), suggesting that H3K79 was succinylated. The IonScore (Mascot) was 51, and the expectation value (P) was 5.5 × 10−3. b, Analyses of the specificity of the antibody against H3K79 succinylation. Synthetic histone H3 peptides with or without H3K79 succinylation were used to test the specificity of the antibody against H3K79 succinylation. Representative images of triplicate experiments are shown. MW, molecular weight. c, Identification of histone H3 K79 succinylation in U251 cells. Histones were extracted from U251 cells. Mass spectrometric analysis of a tryptic fragment of histone H3 extracts from U251 cells at monoisotopic m/z 718.35559 Da (−1.26 m.m.u./ −1.75 p.p.m.) matched with the doubly charged peptide 73-EIAQDFKTDLR-83 with K7-succinyl (100.01604 Da), suggesting that H3K79 was succinylated. The IonScore (Mascot) was 40, and the expectation value (P) was 2.5 × 10−2. d, Semi-quantitative comparison of posttranslational modifications of histone H3 with or without KAT2A depletion. Histones were extracted from U251 cells with or without KAT2A depletion. Mass spectrometric analyses identified the specific peptides of H3K14 acetylation (KSTGGKacAPR), H3K24 acetylation (KQLATKacAAR), H3K79 succinylation (EIAQDFKsuccTDLR), and H3K122 acetylation (RVTIMPKacDIQLAR). The succinylated H3K79 peptide was identified in the histone extracts from U251 cells in the absence but not presence of KAT2A depletion (Mascot IonScore, 40; expectation value, 2.5 × 10−2). The peptides of H3K14ac, H3K24ac, and H3K122ac were identified in histone extracts from U251 cells with or without KAT2A depletion (Mascot IonScore >30; expectation value < 0.05). KAT2A depletion reduced H3K14 acetylation, but not acetylation of H3K24 and H3K122. The semiquantitative results were obtained by the ratio of specific peptide iron intensity to total histone H3 iron intensity (n = 1). e, EP300 does not regulate histone H3 K79 succinylation. EP300 shRNA was expressed in U251 cells. Immunoblotting analyses were performed with the indicated antibodies. EP300 depletion reduced the acetylation of H3K18 and H3K27. Representative images of triplicate experiments are shown. f, Quantitative analyses of acetyl-CoA and succinyl-CoA levels in GBM cells. The levels of acetyl-CoA and succinyl-CoA in the nuclear extracts of U87 and U251 cells were measured by LC–MS/MS. Quantitative limits: ratio of signal intensity to noise intensity > 5. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from three independent experiments (n = 3). *P < 0.05. g, The enzymatic activity of wild-type OGDH and inactive OGDH(P459A/Y460A) was determined. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from three independent experiments (n = 3). *P < 0.05. h, KAT2A uses α-KGDH-produced succinyl-CoA for histone H3 succinylation in the presence of acetyl-CoA and absence of exogenous succinyl-CoA. Purified KAT2A, histone H3, CoA, α-ketoglutarate, NAD+, and acetyl-CoA were incubated with or without immunoprecipitated wild-type Flag–OGDH from 293 cells. Immunoblotting analyses were performed with the indicated antibodies. Representative images of triplicate experiments are shown. i, The DLST NLS mutant does not succinylate purified histone H3. Wild-type Flag–rKAT2A was co-expressed with wild-type V5–rDLST or V5–rDLST NLS mutant in 293 cells. The nuclei of 293 cells were extracted. Immunoprecipitated nuclear wild-type Flag–rKAT2A (which associated with wild-type V5–rDLST but not V5–rDLST NLS mutant) was incubated with purified histone H3, CoA, α-ketoglutarate, and NAD+. Immunoblotting analyses were performed with the indicated antibodies. Representative images of triplicate experiments are shown. j, KAT2A in the SAGA complex does not regulate H3K79 succinylation. U251 cells with or without TAF9 shRNA were analysed by immunoblotting assay with the indicated antibodies. Representative images of triplicate experiments are shown. k, Genomic distributions of ChIP–seq peaks for H3K79 succinylation, KAT2A, and α-KGDH (OGDH) in U251 cells. l, Venn diagram shows the overlap of H3K79 succinylation (red)-, KAT2A (blue)-, and α-KGDH (yellow)-occupied gene promoters (P = 0.02 (one-sided Fisher’s exact test)). m, H3K79 succinylation by KAT2A is a dynamic process. Purified KAT2A was incubated with streptavidin bead-immobilized nucleosomes and biotin-labelled DNA. The immobilized KAT2A–nucleosome complex was washed with PBS three times and incubated with succinyl-CoA in the presence or absence of purified and soluble nucleosome-containing His–histone H3, which was assembled with bacterially purified nonmodified histone components. Immunoblotting analyses were performed with the indicated antibodies. Representative images of triplicate experiments are shown. n, KAT2A depletion reduces H3K79 succinylation in the gene promoter regions. Venn diagram showing the overlap of H3K79 succinylation-occupied gene promoters in U251 cells (red) and U251 cells expressing KAT2A shRNA (green) (P = 0.02 (one-sided Fisher’s exact test)).
Extended Data Figure 4 Cell signalling pathways enriched with genes with H3K79-succinylated promoter regions.
Nodes stand for 130 GSEA-identified pathways (P < 0.005 and FDR q < 0.001). Each node represents one pathway. The node size reflects the number of genes in the pathway, whereas the node colour reflects the P value from the GSEA. Red indicates that the pathways are significantly enriched with genes with H3K79 succinylation in their promoter regions. Edges: two nodes are connected if the group of genes from the smaller node is strictly contained in the gene group from the larger node (that is, every member of the smaller pathway is also a member of the larger pathway). Highly connected node clusters reflect functional modules, which share a large portion of genes regulated by H3K79 succinylation. The 130 nodes are connected by 147 edges. Among the 130 nodes, 21 nodes, which are connected by 67 edges, are linked to the pathways of Cell Cycle and Cell Cycle Mitotic, both of which are defined in the Reactome database.
Extended Data Figure 5 Selected cell signalling pathways enriched with the genes with H3K79-succinylated promoter regions.
a, H3K79 succinylation regulates critical cellular signalling pathways. Hypergeometric tests selected H3K79 succinylation-enriched signalling pathways in U251 cells with DLST(R224A/K226E) or KAT2A(Y645A) mutant expression. y-axis: level of significance from RNA-seq data, which takes the form of (−log10P) × reg, where reg = −1, indicating that the pathway is enriched with downregulated genes based on the comparison of mutant with wild-type proteins. One-sided hypergeometric test for overrepresentation was conducted. The data are from one RNA-seq sample. b, c, Combined view of ChIP–seq and RNA-seq data revealing histone H3K79 succinylation-enriched pathways that were significantly suppressed by the expression of V5–rDLST(R224A/K226E) (b) or Flag–rKAT2A(Y645A) (c). Significantly downregulated pathways were identified via hypergeometric tests (blue nodes, P < 0.01 and FDR q < 0.05) and were enriched with downregulated genes (individual genes with fold enrichment >2 and FDR q < 0.1 in ChIP–seq data and P < 0.05 and FDR q < 0.1 in RNA-seq data). The selected genes (PRKDC, JUN, and PIK3R1) were those that were most significantly suppressed (red nodes, P < 0.005 and FDR q < 0.05 in RNA-seq; fold enrichment >4 and FDR q < 0.05 in ChIP–seq) or important hitches between these suppressed pathways. The blue node size reflects the number of significantly suppressed genes in the pathway. Edges represent more than 40% of gene members shared between the pathways. d, e, α-KGDH-coupled KAT2A regulates gene expression. mRNA expression levels of PIK3R1, JUN, and PRKDC in U87 cells with depleted endogenous KAT2A and reconstituted expression of wild-type Flag–rKAT2A or Flag–rKAT2A(Y645A) mutant (d) or with depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant (e) were determined using quantitative PCR. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from three independent experiments (n = 3). *P < 0.05. f, g, α-KGDH-coupled KAT2A regulates gene expression. mRNA expression levels of PIK3R1, JUN, and PRKDC in U251 cells with depleted endogenous KAT2A and reconstituted expression of wild-type Flag–rKAT2A or Flag–rKAT2A(Y645A) (f) or with depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant (g) were determined using quantitative PCR. Two-sided t-tests were conducted. The data are presented as the means ± s.d. from three independent experiments (n = 3). *P < 0.05. h, i, α-KGDH-coupled KAT2A regulates gene expression. Expression levels of p85α, c-Jun, and DNA-PKcs in U251 cells with depleted endogenous KAT2A and reconstituted expression of wild-type Flag–rKAT2A or Flag–rKAT2A(Y645A) (h) or with depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant (i) were determined using immunoblotting analyses with the indicated antibodies. Representative images of triplicate experiments are shown.
Extended Data Figure 6 α-KGDH-coupled KAT2A promotes tumour growth.
a, α-KGDH-coupled KAT2A promotes tumour cell proliferation. A total of 104 U251 cells with depleted endogenous KAT2A and reconstituted expression of wild-type Flag–rKAT2A or Flag–rKAT2A(Y645A) or depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant were plated. The cells were collected and counted daily for nine days. The data are presented as the means ± s.d. from three independent experiments (n = 3). b, α-KGDH-coupled KAT2A promotes tumour growth. U87 cells with depleted endogenous DLST and reconstituted expression of wild-type V5–rDLST or V5–rDLST NLS mutant were intracranially injected into athymic nude mice. Representative haematoxylin and eosin-stained coronal brain sections are shown. Tumour volumes were calculated. Two-sided t-tests were conducted. Data represent the means ± s.d. of 5 mice (n = 5). *P < 0.05. c, Immunohistochemical analyses of the brain tumour sections were performed with an anti-Ki67 antibody. d, Immunohistochemical analyses of the brain tumour sections were performed with the indicated antibodies. e, f, α-KGDH-coupled KAT2A promotes H3K79 succinylation in tumour cells. Quantification of the immunoblotting (Fig. 4d) was performed using ImageJ software (e). Immunoblotting analyses of histone extracts from tumour tissues were performed with the indicated antibodies (f). Two-sided t-tests were conducted. Quantitative data represent the means ± s.d. of 5 samples (n = 5). *P < 0.05. Representative images of triplicate experiments are shown.
Supplementary information
Supplementary Information
This file contains a Supplementary Note, Supplementary References and the Gel Source Data. (PDF 7550 kb)
Supplementary Data
This zipped file contains Supplementary Tables 1-9. (ZIP 3515 kb)
Rights and permissions
About this article
Cite this article
Wang, Y., Guo, Y., Liu, K. et al. KAT2A coupled with the α-KGDH complex acts as a histone H3 succinyltransferase. Nature 552, 273–277 (2017). https://doi.org/10.1038/nature25003
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature25003
This article is cited by
-
KAT2A-mediated succinylation modification of notch1 promotes the proliferation and differentiation of dental pulp stem cells by activating notch pathway
BMC Oral Health (2024)
-
Novel histone post-translational modifications in Alzheimer’s disease: current advances and implications
Clinical Epigenetics (2024)
-
Deciphering functional roles of protein succinylation and glutarylation using genetic code expansion
Nature Chemistry (2024)
-
Regulation of urea cycle by reversible high-stoichiometry lysine succinylation
Nature Metabolism (2024)
-
A glimpse into novel acylations and their emerging role in regulating cancer metastasis
Cellular and Molecular Life Sciences (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.