Abstract
Although individuals age and die with time, an animal species can continue indefinitely, because of its immortal germ-cell lineage1. How the germline avoids transmitting damage from one generation to the next remains a fundamental question in biology. Here we identify a lysosomal switch that enhances germline proteostasis before fertilization. We find that Caenorhabditis elegans oocytes whose maturation is arrested by the absence of sperm2 exhibit hallmarks of proteostasis collapse, including protein aggregation. Remarkably, sperm-secreted hormones re-establish oocyte proteostasis once fertilization becomes imminent. Key to this restoration is activation of the vacuolar H+-ATPase (V-ATPase), a proton pump that acidifies lysosomes3. Sperm stimulate V-ATPase activity in oocytes by signalling the degradation of GLD-1, a translational repressor4 that blocks V-ATPase synthesis. Activated lysosomes, in turn, promote a metabolic shift that mobilizes protein aggregates for degradation, and reset proteostasis by enveloping and clearing the aggregates. Lysosome acidification also occurs during Xenopus oocyte maturation; thus, a lysosomal switch that enhances oocyte proteostasis in anticipation of fertilization may be conserved in other species.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Medvedev, Z. A. On the immortality of the germ line: genetic and biochemical mechanism. A review. Mech. Ageing Dev. 17, 331–359 (1981)
Miller, M. A. et al. A sperm cytoskeletal protein that signals oocyte meiotic maturation and ovulation. Science 291, 2144–2147 (2001)
Ohkuma, S., Moriyama, Y. & Takano, T. Identification and characterization of a proton pump on lysosomes by fluorescein-isothiocyanate-dextran fluorescence. Proc. Natl Acad. Sci. USA 79, 2758–2762 (1982)
Nousch, M. & Eckmann, C. R. Translational control in the Caenorhabditis elegans germ line. Adv. Exp. Med. Biol. 757, 205–247 (2013)
Goudeau, J. & Aguilaniu, H. Carbonylated proteins are eliminated during reproduction in C. elegans. Aging Cell 9, 991–1003 (2010)
Greenstein, D. Control of oocyte meiotic maturation and fertilization. WormBook doi:10.1895/wormbook.1.53.1 (2005)
David, D. C. et al. Widespread protein aggregation as an inherent part of aging in C. elegans. PLoS Biol. 8, e1000450 (2010)
McCarter, J., Bartlett, B., Dang, T. & Schedl, T. On the control of oocyte meiotic maturation and ovulation in Caenorhabditis elegans. Dev. Biol. 205, 111–128 (1999)
Singson, A., Mercer, K. B. & L’Hernault, S. W. The C. elegans spe-9 gene encodes a sperm transmembrane protein that contains EGF-like repeats and is required for fertilization. Cell 93, 71–79 (1998)
Govindan, J. A., Cheng, H., Harris, J. E. & Greenstein, D. Gαo/i and Gαs signaling function in parallel with the MSP/Eph receptor to control meiotic diapause in C. elegans. Curr. Biol. 16, 1257–1268 (2006)
Kamath, R. S. et al. Systematic functional analysis of the Caenorhabditis elegans genome using RNAi. Nature 421, 231–237 (2003)
Hughes, A. L. & Gottschling, D. E. An early age increase in vacuolar pH limits mitochondrial function and lifespan in yeast. Nature 492, 261–265 (2012)
Yoshimori, T., Yamamoto, A., Moriyama, Y., Futai, M. & Tashiro, Y. Bafilomycin A1, a specific inhibitor of vacuolar-type H+-ATPase, inhibits acidification and protein degradation in lysosomes of cultured cells. J. Biol. Chem. 266, 17707–17712 (1991)
Zhou, Q., Li, H. & Xue, D. Elimination of paternal mitochondria through the lysosomal degradation pathway in C. elegans. Cell Res. 21, 1662–1669 (2011)
Kostich, M., Fire, A. & Fambrough, D. M. Identification and molecular-genetic characterization of a LAMP/CD68-like protein from Caenorhabditis elegans. J. Cell Sci. 113, 2595–2606 (2000)
Sato, M. & Sato, K. Degradation of paternal mitochondria by fertilization-triggered autophagy in C. elegans embryos. Science 334, 1141–1144 (2011)
Mijaljica, D., Prescott, M. & Devenish, R. J. Microautophagy in mammalian cells: revisiting a 40-year-old conundrum. Autophagy 7, 673–682 (2011)
Wright, J. E. et al. A quantitative RNA code for mRNA target selection by the germline fate determinant GLD-1. EMBO J. 30, 533–545 (2011)
Jones, A. R., Francis, R. & Schedl, T. GLD-1, a cytoplasmic protein essential for oocyte differentiation, shows stage- and sex-specific expression during Caenorhabditis elegans germline development. Dev. Biol. 180, 165–183 (1996)
Francis, R., Barton, M. K., Kimble, J. & Schedl, T. gld-1, a tumor suppressor gene required for oocyte development in Caenorhabditis elegans. Genetics 139, 579–606 (1995)
Merritt, C. & Seydoux, G. Transgenic solutions for the germline. WormBook doi:10.1895/wormbook.1.148.1 (2010)
Brand, M. D. & Nicholls, D. G. Assessing mitochondrial dysfunction in cells. Biochem. J. 435, 297–312 (2011)
Chance, B. & Williams, G. R. Respiratory enzymes in oxidative phosphorylation. I. Kinetics of oxygen utilization. J. Biol. Chem. 217, 383–393 (1955)
Tantama, M., Martínez-François, J. R., Mongeon, R. & Yellen, G. Imaging energy status in live cells with a fluorescent biosensor of the intracellular ATP-to-ADP ratio. Nat. Commun. 4, 2550 (2013)
Hung, Y. P., Albeck, J. G., Tantama, M. & Yellen, G. Imaging cytosolic NADH-NAD+ redox state with a genetically encoded fluorescent biosensor. Cell Metab. 14, 545–554 (2011)
Parry, B. R. et al. The bacterial cytoplasm has glass-like properties and is fluidized by metabolic activity. Cell 156, 183–194 (2014)
Pastore, A. & Temussi, P. Protein aggregation and misfolding: good or evil? J. Phys. Condens. Matter 24, 244101 (2012)
Unal, E., Kinde, B. & Amon, A. Gametogenesis eliminates age-induced cellular damage and resets life span in yeast. Science 332, 1554–1557 (2011)
Roux, A. E., Langhans, K., Huynh, W. & Kenyon, C. Reversible age-related phenotypes induced during larval quiescence in C. elegans. Cell Metab. 23, 1113–1126 (2016)
Conboy, I. M. et al. Rejuvenation of aged progenitor cells by exposure to a young systemic environment. Nature 433, 760–764 (2005)
Redemann, S. et al. Codon adaptation-based control of protein expression in C. elegans. Nat. Methods 8, 250–252 (2011)
Dantuma, N. P., Lindsten, K., Glas, R., Jellne, M. & Masucci, M. G. Short-lived green fluorescent proteins for quantifying ubiquitin/proteasome-dependent proteolysis in living cells. Nat. Biotechnol. 18, 538–543 (2000)
Kelly, W. G., Xu, S., Montgomery, M. K. & Fire, A. Distinct requirements for somatic and germline expression of a generally expressed Caernorhabditis elegans gene. Genetics 146, 227–238 (1997)
Mello, C. C., Kramer, J. M., Stinchcomb, D. & Ambros, V. Efficient gene transfer in C.elegans: extrachromosomal maintenance and integration of transforming sequences. EMBO J. 10, 3959–3970 (1991)
Doniach, T. & Hodgkin, J. A sex-determining gene, fem-1, required for both male and hermaphrodite development in Caenorhabditis elegans. Dev. Biol. 106, 223–235 (1984)
Yang, J. S. et al. OASIS: online application for the survival analysis of lifespan assays performed in aging research. PLoS One 6, e23525 (2011)
Acknowledgements
We thank members of the Kenyon laboratory and our colleagues at Calico for discussions and comments on the manuscript, and the Calico microscopy core, especially M. Ingaramo, for help with microscopy and sensors. We thank the E. Blackburn and P. O’Farrell laboratories for sharing equipment at UCSF. K. Sato provided the Ppie-1::gfp::lgg-1 strain. Other strains were provided by the CGC, funded by the NIH Office of Research Infrastructure Programs (P40 OD010440). Research was performed at UCSF and then Calico, and was supported at UCSF by the George and Judy Marcus Family Foundation, the Life Extension and Chuan Lyu Foundations, and by NIH grant R37/R01 AG11816 to C.K. C.K. is now Vice President of Aging Research at Calico Life Sciences, which supported the research done at Calico. K.A.B. is an Honorary Fellow of the Jane Coffin Childs Memorial Fund.
Author information
Authors and Affiliations
Contributions
K.A.B. and C.K. designed experiments, interpreted data, and wrote the manuscript. K.A.B. performed all experiments.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Pattern of germline protein aggregates.
a–c, Full gonad images of three aggregation-prone proteins in young hermaphrodites and females. Enlarged regions of different parts of the gonad are also shown. Bars, 10 μm.
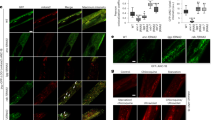
Extended Data Figure 2 Signals from sperm reduce protein aggregation in oocytes.
a–f, Time-lapse images of aggregation-prone proteins in oocytes of females, or females after mating. g–i, Localized fluorescence intensities in oocytes of mated females and non-mated controls. Mean ± s.d. for n = 10 aggregate sites. NS, not significant. ****P < 0.0001. j, k, GFP::RHO-1-aggregation in females and sperm-defective hermaphrodite mutants that still produce MSPs. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. Bars, 10 μm.
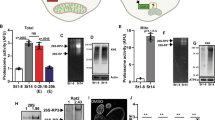
Extended Data Figure 3 V-ATPase suppression of oocyte protein aggregation.
a, Lifespan, mean lifespans in parentheses. b–d, GFP::RHO-1-expressing hermaphrodites with catalytic (V1) or membrane-anchoring (V0) V-ATPase subunits knocked down. Animals with oocyte protein aggregation counted. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. e, Mating after vha-13 knockdown. f, GFP::RHO-1-expressing gsa-1(ce94gf) hermaphrodites following control or vha-13 RNAi. g, Percentage of gsa-1(ce94gf) animals with oocyte protein aggregation. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. h, DMSO- or bafilomycin A1-injected germlines. i, Control or NH4Cl-treated germlines. Bars, 10 μm.
Extended Data Figure 4 Regulation of lysosomal acidity in the germline.
a, Hermaphrodite and female worms stained with LysoSensor Blue DND-99. Gonads are outlined. b, Percentage of puncta that are positive for LysoTracker and/or GFP::VHA-13. Mean ± s.d. from n = 10 proximal oocytes. c, Co-localization (arrows) of GFP::VHA-13 and LysoTracker puncta in an oocyte (enlarged region). d, Proximal oocyte before and after mating in a GFP::VHA-13-expressing female. Bars, 5 μm.
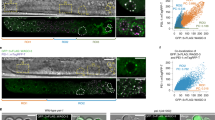
Extended Data Figure 5 Aggregates are not cleared by macroautophagy.
a, GFP::LGG-1-expressing germlines. b, Number of autophagosomes (mean ± s.d.) in the most proximal oocyte. c, Schematic of macroautophagy. d, GFP::LGG-1 after control or lgg-1 RNAi. e,f, Quantification of macroautophagy gene expression by RT-PCR. Normalized expression (mean ± s.d.) was scored for three biological replicates. ****P < 0.0001. The gel source data are shown in Supplementary Figure 1. g, h, GFP::RHO-1-expressing hermaphrodites treated with macroautophagy-gene RNAi. Percentage of animals with oocyte protein aggregation. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. i, Matings after macroautophagy-gene RNAi. j, Time-lapse images of aggregate clearance. Bars, 5 μm.
Extended Data Figure 6 Proteasome involvement in germline proteostasis.
a, LysoTracker-stained dissected germlines. b, GFP::PBS-1 localization. c, d, Schematic and imaging of proteasome sensor UbG76V::GFP. Active proteasomes degrade UbG76V::GFP, unless inhibited by MG132. e–g, GFP::RHO-1-aggregation following control or proteasomal pbs-1 RNAi. The gld-1(q485) mutation precluded aggregation following pbs-1 RNAi. This finding fits the model that the proteasome degrades GLD-1, but not the aggregates, consistent with aggregate engulfment by lysosomes. However, we note that proximal gld-1 germ cells, which form tumors20, could potentially be non-permissive for aggregation. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. Bars, 10 μm.
Extended Data Figure 7 Sperm-induced changes in mitochondrial morphology and ROS levels require V-ATPase function.
a, MitoLS::GFP in germ cells of hermaphrodites. b, Different z-planes for MitoLS::GFP in the same distal germline. c, Proximal:distal MitoLS::GFP fluorescence ratios (mean ± s.d.) for n = 10 germlines. d, e, Proximal oocytes from MitoLS::GFP-expressing or MitoTracker CM-H2TMRos-stained females. f, g, Mitochondria from proximal oocytes in MitoLS::GFP-expressing females before and after mating. Mitochondrial lengths (mean ± s.d.). ****P < 0.0001. h, i, Proximal oocytes from MitoLS::GFP-expressing or MitoTracker CM-H2TMRos-stained hermaphrodites after vha-13 RNAi. Bars, 10 μm.
Extended Data Figure 8 Regulation of mitochondrial membrane potential in the germline.
a, DiOC6(3)-stained germlines from control (ethanol solvent)- or antimycin-treated hermaphrodites. b, Percentage of DiOC6(3)-stained germlines. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. c, Real-colour and heatmap images of DiOC6(3)-stained germlines. d, JC-1-stained mitochondria in the distal and proximal germline of a wild-type hermaphrodite. e, JC-1-stained proximal germline mitochondria following control or vha-13 RNAi. Bars, 5 μm.
Extended Data Figure 9 ATP-synthase inhibition prevents the reduction in mitochondrial membrane potential in proximal oocytes and blocks aggregate clearance.
a, Real-colour and heatmap images of DiOC6(3)-stained germlines after RNAi of genes encoding ATP synthase subunits. b, c, Aggregation-prone proteins in control (ethanol solvent)- or oligomycin-treated hermaphrodites. Percentage of animals with oocyte protein aggregation. Mean ± s.d. from three biological replicates, each of n = 50 animals. ****P < 0.0001. d, LysoTracker reveals lysosome acidification in GFP::RHO-1-expressing hermaphrodites following asb-1 knockdown. Dotted line, intestine. Bars, 10 μm.
Extended Data Figure 10 Activation of germ cell metabolism.
a, Immature germ cells arrest with a high ATP:ADP ratio and a high energy charge, which are reversed in response to sperm signals as ADP levels rise and unlock the ATP synthase. These changes reflect a shift from a resting to an active metabolic state22,23. b, 488 nm-excited fluorescence of PercevalHR and cpmVenus. c, Heatmap of the PercevalHR λhigh/λlow ratio after vha-13 knockdown. d, Peredox fluorescence in a hermaphrodite germline, with line profiles. e, f, GFP::GLD-1 and mitochondrial morphology in ADP- or vehicle-injected female oocytes. Bars, 10 μm.
Supplementary information
Supplementary Figure 1
This file contains the uncropped images of DNA gels. (PDF 722 kb)
Source data
Rights and permissions
About this article
Cite this article
Bohnert, K., Kenyon, C. A lysosomal switch triggers proteostasis renewal in the immortal C. elegans germ lineage. Nature 551, 629–633 (2017). https://doi.org/10.1038/nature24620
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24620
This article is cited by
-
The HEAT repeat protein HPO-27 is a lysosome fission factor
Nature (2024)
-
ATP induces folding of ALS-causing C71G-hPFN1 and nascent hSOD1
Communications Chemistry (2023)
-
Maternal aging increases offspring adult body size via transmission of donut-shaped mitochondria
Cell Research (2023)
-
Vitellogenin accumulation leads to reproductive senescence by impairing lysosomal function
Science China Life Sciences (2023)
-
Mutant C. elegans mitofusin leads to selective removal of mtDNA heteroplasmic deletions across generations to maintain fitness
BMC Biology (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.