Abstract
The branched-chain amino acid (BCAA) pathway and high levels of BCAA transaminase 1 (BCAT1) have recently been associated with aggressiveness in several cancer entities1,2,3,4,5,6. However, the mechanistic role of BCAT1 in this process remains largely uncertain. Here, by performing high-resolution proteomic analysis of human acute myeloid leukaemia (AML) stem-cell and non-stem-cell populations, we find the BCAA pathway enriched and BCAT1 protein and transcripts overexpressed in leukaemia stem cells. We show that BCAT1, which transfers α-amino groups from BCAAs to α-ketoglutarate (αKG), is a critical regulator of intracellular αKG homeostasis. Further to its role in the tricarboxylic acid cycle, αKG is an essential cofactor for αKG-dependent dioxygenases such as Egl-9 family hypoxia inducible factor 1 (EGLN1) and the ten-eleven translocation (TET) family of DNA demethylases7,8,9,10. Knockdown of BCAT1 in leukaemia cells caused accumulation of αKG, leading to EGLN1-mediated HIF1α protein degradation. This resulted in a growth and survival defect and abrogated leukaemia-initiating potential. By contrast, overexpression of BCAT1 in leukaemia cells decreased intracellular αKG levels and caused DNA hypermethylation through altered TET activity. AML with high levels of BCAT1 (BCAT1high) displayed a DNA hypermethylation phenotype similar to cases carrying a mutant isocitrate dehydrogenase (IDHmut), in which TET2 is inhibited by the oncometabolite 2-hydroxyglutarate11,12. High levels of BCAT1 strongly correlate with shorter overall survival in IDHWTTET2WT, but not IDHmut or TET2mut AML. Gene sets characteristic for IDHmut AML13 were enriched in samples from patients with an IDHWTTET2WTBCAT1high status. BCAT1high AML showed robust enrichment for leukaemia stem-cell signatures14,15, and paired sample analysis showed a significant increase in BCAT1 levels upon disease relapse. In summary, by limiting intracellular αKG, BCAT1 links BCAA catabolism to HIF1α stability and regulation of the epigenomic landscape, mimicking the effects of IDH mutations. Our results suggest the BCAA–BCAT1–αKG pathway as a therapeutic target to compromise leukaemia stem-cell function in patients with IDHWTTET2WT AML.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
01 August 2018
In Extended Data Fig. 1a of this Article, the FACS plot depicting the surface phenotype of AML sample DD08 was a duplicate of the plot for AML sample DD06. Supplementary Data 4 has been added to the Supplementary Information of the original Letter to clarify the proteome data acquisition and presentation. The original Letter has been corrected online.
References
Tönjes, M. et al. BCAT1 promotes cell proliferation through amino acid catabolism in gliomas carrying wild-type IDH1. Nat. Med. 19, 901–908 (2013)
Wang, Z. Q. et al. BCAT1 expression associates with ovarian cancer progression: possible implications in altered disease metabolism. Oncotarget 6, 31522–31543 (2015)
Zheng, Y. H. et al. BCAT1, a key prognostic predictor of hepatocellular carcinoma, promotes cell proliferation and induces chemoresistance to cisplatin. Liver Int. 36, 1836–1847 (2016)
Thewes, V. et al. The branched-chain amino acid transaminase 1 sustains growth of antiestrogen-resistant and ERα-negative breast cancer. Oncogene 36, 4124–4134 (2017)
Mayers, J. R. et al. Tissue of origin dictates branched-chain amino acid metabolism in mutant Kras-driven cancers. Science 353, 1161–1165 (2016)
Hattori, A. et al. Cancer progression by reprogrammed BCAA metabolism in myeloid leukaemia. Nature 545, 500–504 (2017)
Loenarz, C. & Schofield, C. J. Expanding chemical biology of 2-oxoglutarate oxygenases. Nat. Chem. Biol. 4, 152–156 (2008)
Kaelin, W. G. Jr & McKnight, S. L. Influence of metabolism on epigenetics and disease. Cell 153, 56–69 (2013)
Epstein, A. C. et al. C. elegans EGL-9 and mammalian homologs define a family of dioxygenases that regulate HIF by prolyl hydroxylation. Cell 107, 43–54 (2001)
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009)
Xu, W. et al. Oncometabolite 2-hydroxyglutarate is a competitive inhibitor of α-ketoglutarate-dependent dioxygenases. Cancer Cell 19, 17–30 (2011)
Dang, L. et al. Cancer-associated IDH1 mutations produce 2-hydroxyglutarate. Nature 462, 739–744 (2009)
Marcucci, G. et al. IDH1 and IDH2 gene mutations identify novel molecular subsets within de novo cytogenetically normal acute myeloid leukemia: a Cancer and Leukemia Group B study. J. Clin. Oncol. 28, 2348–2355 (2010)
Eppert, K. et al. Stem cell gene expression programs influence clinical outcome in human leukemia. Nat. Med. 17, 1086–1093 (2011)
Ng, S. W. et al. A 17-gene stemness score for rapid determination of risk in acute leukaemia. Nature 540, 433–437 (2016)
Sarry, J. E. et al. Human acute myelogenous leukemia stem cells are rare and heterogeneous when assayed in NOD/SCID/IL2Rγc-deficient mice. J. Clin. Invest. 121, 384–395 (2011)
Guan, Y., Gerhard, B. & Hogge, D. E. Detection, isolation, and stimulation of quiescent primitive leukemic progenitor cells from patients with acute myeloid leukemia (AML). Blood 101, 3142–3149 (2003)
Ichihara, A. & Koyama, E. Transaminase of branched chain amino acids. I. Branched chain amino acids-α-ketoglutarate transaminase. J. Biochem. 59, 160–169 (1966)
Jones, M. E. Pyrimidine nucleotide biosynthesis in animals: genes, enzymes, and regulation of UMP biosynthesis. Annu. Rev. Biochem. 49, 253–279 (1980)
Wang, Y., Liu, Y., Malek, S. N., Zheng, P. & Liu, Y. Targeting HIF1α eliminates cancer stem cells in hematological malignancies. Cell Stem Cell 8, 399–411 (2011)
Figueroa, M. E. et al. Leukemic IDH1 and IDH2 mutations result in a hypermethylation phenotype, disrupt TET2 function, and impair hematopoietic differentiation. Cancer Cell 18, 553–567 (2010)
Paschka, P. et al. IDH1 and IDH2 mutations are frequent genetic alterations in acute myeloid leukemia and confer adverse prognosis in cytogenetically normal acute myeloid leukemia with NPM1 mutation without FLT3 internal tandem duplication. J. Clin. Oncol. 28, 3636–3643 (2010)
Kharas, M. G. et al. Musashi-2 regulates normal hematopoiesis and promotes aggressive myeloid leukemia. Nat. Med. 16, 903–908 (2010)
Wouters, B. J. et al. Double CEBPA mutations, but not single CEBPA mutations, define a subgroup of acute myeloid leukemia with a distinctive gene expression profile that is uniquely associated with a favorable outcome. Blood 113, 3088–3091 (2009)
The Cancer Genome Atlas Research Network. Genomic and epigenomic landscapes of adult de novo acute myeloid leukemia. N. Engl. J. Med. 368, 2059–2074 (2013)
Ko, M. et al. Impaired hydroxylation of 5-methylcytosine in myeloid cancers with mutant TET2. Nature 468, 839–843 (2010)
TeSlaa, T. et al. α-Ketoglutarate accelerates the initial differentiation of primed human pluripotent stem cells. Cell Metab. 24, 485–493 (2016)
Carey, B. W., Finley, L. W., Cross, J. R., Allis, C. D. & Thompson, C. B. Intracellular α-ketoglutarate maintains the pluripotency of embryonic stem cells. Nature 518, 413–416 (2015)
Hutson, S. M., Sweatt, A. J. & Lanoue, K. F. Branched-chain amino acid metabolism: implications for establishing safe intakes. J. Nutr. 135 (Suppl.), 1557S–1564S (2005)
Boersema, P. J., Raijmakers, R., Lemeer, S., Mohammed, S. & Heck, A. J. Multiplex peptide stable isotope dimethyl labeling for quantitative proteomics. Nat. Protocols 4, 484–494 (2009)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Smyth, G. K. Linear models and empirical Bayes methods for assessing differential expression in microarray experiments. Stat. Appl. Genet. Mol. Biol. 3, 3 (2004)
Weber, K., Bartsch, U., Stocking, C. & Fehse, B. A multicolor panel of novel lentiviral “Gene Ontology” (LeGO) vectors for functional gene analysis. Mol. Ther. 16, 698–706 (2008)
Hiller, K. et al. MetaboliteDetector: comprehensive analysis tool for targeted and nontargeted GC/MS based metabolome analysis. Anal. Chem. 81, 3429–3439 (2009)
Kochanowski, N. et al. Intracellular nucleotide and nucleotide sugar contents of cultured CHO cells determined by a fast, sensitive, and high-resolution ion-pair RP-HPLC. Anal. Biochem. 348, 243–251 (2006)
Wessel, D. & Flügge, U. I. A method for the quantitative recovery of protein in dilute solution in the presence of detergents and lipids. Anal. Biochem. 138, 141–143 (1984)
Assenov, Y. et al. Comprehensive analysis of DNA methylation data with RnBeads. Nat. Methods 11, 1138–1140 (2014)
Teschendorff, A. E. et al. A beta-mixture quantile normalization method for correcting probe design bias in Illumina Infinium 450 k DNA methylation data. Bioinformatics 29, 189–196 (2013)
Welch, B. On the comparison of several mean values: an alternative approach. Biometrika 38, 330–336 (1951)
Li, S. et al. Distinct evolution and dynamics of epigenetic and genetic heterogeneity in acute myeloid leukemia. Nat. Med. 22, 792–799 (2016)
Acknowledgements
We thank all members of HI-STEM for discussions, M. Milsom and S. Haas for reading the manuscript, A. Ehninger for help with AML sample acquisition, the members of the Central Animal Laboratory at DKFZ for animal husbandry, the members of the DKFZ Flow Cytometry Core Facility for expertise and support, R. Delwel, P. Valk and B. Lowenberg for providing patient survival data for the Erasmus GSE14468 dataset, and A. Lenze for processing cord blood samples. We thank the EMBL Proteomics Core Facility for assistance with mass spectrometry analysis, the microarray unit of the DKFZ Genomics and Proteomics Core Facility for support, and the Metabolomics Core Technology Platform of the Excellence Cluster CellNetworks for support with ultra-performance liquid chromatography-based metabolite quantification. This work was supported by the SFB873 funded by the Deutsche Forschungsgemeinschaft (DFG) (C.L., C.S., and A.T.), the SyTASC consortium funded by the Deutsche Krebshilfe (A.T.) and the Dietmar Hopp Foundation (A.T.), by grant ZUK 49/2 from the DFG (G.P.), and the DFG Heisenberg-Professorship BU 1339/8-1 (L.B.).
Author information
Authors and Affiliations
Contributions
S.R. designed the study and performed experiments; M.F. performed experiments and bioinformatic analyses with conceptual input from S.R. and C.H.; N.K. performed tracing experiments with the help of Y.N. and K.H.; J.H. and J.K. generated and analysed the proteome data; W.W. helped with cloning, generated growth curves and colony-forming unit assays on HL-60 KD cells, and performed western blotting on primary samples; C.L. and A.D.H. provided AML samples, clinical data, and conceptual input; L.B. provided the GSE16432 dataset and conceptual input; G.P. and R.H. performed targeted metabolomics; A.B., C.B., P.Z., A.P., and M.S. helped with mouse and in vitro experiments; M.T., A.E., L.A., P.J., C.S., and S.F. gave conceptual input; S.C. and L.B. performed RNA-sequencing of paired diagnosis/relapse samples; C.T., A.F., J.W., and G.E. provided AML samples, P.L. financial support, and P.W. healthy HSPC samples; A.T. designed with S.R. the overall study and supervised it. B.R. helped to design and supervise parts of the study; S.R., M.F., N.K., B.R., and A.T. interpreted the results; and S.R. wrote the manuscript with M.F., N.K., B.R., and A.T.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. G. Vander Heiden, R. Levine and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Design and quality control of proteome analysis and engraftment capacity of AML populations in NSG mice.
a, Flow cytometric analysis of CD34 and CD38 expression (pre-gated on live, lineage-negative cells) of primary AML samples used for proteome. Experiment was repeated three times with similar results. b, Level of chimaerism based on human CD45 expression. Bone marrow of NSG mice was analysed 10–14 weeks after xenotransplantation of patient samples used for proteome analysis. Each dot represents an individual mouse (n = 56). All engrafted populations were CD33+ in more than 95% of human cells, except for DD13, which aberrantly expressed CD19 (not shown). c, Outline of the proteomic workflow. d, Experimental design of proteomic runs. e, Pearson’s correlation coefficient for each comparison defined in column 4 or 6 of d. f, Sample to sample Pearson’s correlation coefficient comparing CD34+CD38– LSC with CD34–CD38+ non-LSC populations. g, The additional chromosome 13 present in sample HD48 does not affect LSC proteins. Pearson’s correlation coefficient for the log2(fold change) of CD34+CD38− LSC to CD34−CD38+ non-LSC proteins comparing HD48 (46,X,-Y,+13, x axis) with DD13 (46,XX, y axis) for all proteins (left) or proteins encoded on chromosome 13 (right).
Extended Data Figure 2 Differentially expressed proteins between LSC and non-LSC populations.
a, Protein types identified in the proteome analysis. b, Volcano plot of proteins comparing CD34+CD38– LSC with CD34–CD38+ non-LSC populations for individual patients. Coloured dots and numbers represent hits significantly regulated (Padj < 0.01) in this comparison and over-represented in the remaining LSC or non-LSC population (see Methods). c, The log2(fold change) between CD34+CD38– LSC and CD34–CD38+ non-LSC in FLT3ITD, NPM1mut AML (x axis) versus FLT3WT, NPM1WT AML samples (y axis). d, Unsupervised clustering and heat map representation of differentially expressed proteins between LSC and non-LSC populations; one missing value per patient allowed per protein, average log2(fold change) colour-coded.
Extended Data Figure 3 BCAA pathway, knockdown of BCAT1, and subsequent xenotransplantation of primary AML samples.
a, Differentially expressed proteins between LSC and non-LSC populations for indicated AML (n = 6) of KEGG_Valine_Leucine_Isoleucine_Degradation pathway; grey, not quantified or not significantly enriched in LSCs. b, RT–qPCR analysis to determine BCAT1 expression (mean ± s.e.m.) in n = 9 primary AML samples 48 h after transduction with shBCAT1-2 or shNT lentiviral vector. BCAT1 values are relative to ABL and GAPDH expression, normalized to shNT. Two-sided, paired t-test. c, Representative picture of a suspension culture 7 days after transduction of shBCAT1-2 or shNT lentiviral vector of HD72 cells, 20× magnification. Experiment was repeated with at least n = 5 other AML samples showing similar results. d, Representative FACS plots after xenotransplantation of primary AML transduced with shBCAT1-2 or shNT. Top row is pre-gated on live cells; middle and bottom rows are pre-gated on live, human CD45+ cells. Numbers indicate percentage per boxed event. The experiment was repeated with n = 7 AML samples showing similar results. e, Chimaerism of huCD45+mCherry+ cells and relative fraction of myeloid (huCD33+), or lymphoid (huCD19+) cells 12 weeks after xenotransplantation of mobilized peripheral CD34+ HSPCs transduced with either shBCAT1-2 or shNT lentiviral vector. The experiment was repeated with two individual HSPC samples; each dot represents one mouse.
Extended Data Figure 4 BCAA tracing experiments.
a, Fraction of labelled TCA cycle metabolites (mean ± s.e.m.) in HL-60, SKM-1, and MOLM-13 cells after 24 h of tracer supplementation (n = 3 independent cultures). b, Total levels (mean) of indicated TCA cycle metabolites, relative to control (n = 3 independent cultures; two-sided, unpaired t-test, only P values less than 0.05 are shown). c, Fraction of M + 1 labelled amino acids (mean ± s.e.m.) after 24 h of BCAA tracing in HL-60, SKM-1, and MOLM-13 (n = 3 independent cultures; two-sided, unpaired t-test). d, Total levels (mean) of glutamate, relative to control (n = 3 independent cultures; two-sided, unpaired t-test, only P values less than 0.05 are shown).
Extended Data Figure 5 Nucleotide measurements upon BCAT1 knockdown and nucleotide rescue experiments.
a, Abundance of individual nucleotides (mean ± s.e.m.) in HL-60, SKM-1, and MOLM-13 transduced with shBCAT1-2 or shNT lentiviral vector (n = 3 independent cultures; unpaired, two-sided t-test, only P values less than 0.05 are shown). b, Relative cell viability (mean ± s.e.m., measured by CellTiter-Blue assay) of shBCAT1-2 or shNT lentiviral vector transduced cell lines, cultivated in media supplemented with nucleotides or carrier (see Methods) (n = 3 independent cultures, normalized to shNT without carrier). c, Relative cell viability (mean ± s.e.m., measured by CellTiter-Blue assay) of shBCAT1 or shNT transduced cell lines, cultivated in media supplemented with nucleosides (see Methods) (n = 3 independent cultures, normalized to shNT without nucleosides).
Extended Data Figure 6 Cell differentiation upon BCAT1-KD, HIF1α response to exogenous αKG, and validation of knockdown and overexpression constructs.
a, Left: representative FACS plots of HL-60 cells induced with shNT or shBCAT1-2 (top row) and treatment with 2 mM DM-αKG or DMSO (bottom row), respectively. Numbers indicate percentage of cells in CD11b+ gate. Right: quantification of CD11b+ cells (mean, after isotype-subtraction) (n = 3 independent cultures; paired, two-sided t-test). b, GSEA of V$PU.1 gene signature in indicated cell lines transduced with shBCAT1-3 versus shNT expression profile. c, Ratio of cell viability (mean, measured by CellTiter-Blue assay) of shBCAT1-2 versus shNT transduced cells without (control) or with (BCAT1-shRESISTANT) stable overexpression of a mutant-BCAT1 resistant to shBCAT1-2 (n = 3 independent cultures, measured 8 days after transduction). d, GSEA plot of enrichment of V$HIF1 gene signature in shBCAT1-2 versus shNT expression profiles of indicated samples from patients with AML. e, Western blot of indicated cell line treated with octyl-αKG or control (methylacetate). Experiment was repeated three times with similar results. For gel source data, see Supplementary Fig. 1. f–j, Gene expression (mean) of BCAT1 (f–h), HIF1α (i), and EGLN1(j) in indicated cell lines, measured by RT–qPCR relative to the expression of ACTB and GAPDH. shNT and shBCAT1-2 (f–h, n = 2 independent experiments in HL-60, n = 5 in SKM-1, n = 4 in MOLM-13), shcontrol_shNT and shcontrol_shBCAT1-2 (f, n = 4 independent experiments in HL-60), control (i, n = 3 independent experiments in HL-60, n = 5 in SKM-1, n = 5 in MOLM-13), HIF1α-OE (i, n = 2 independent experiments in HL-60, n = 3 in SKM-1, n = 2 in MOLM-13), and shcontrol and shEGLN1 (j, n = 4 independent experiments in HL-60) were all measured 48 h after transduction; HIF1α-OE_shNT and HIF1α-OEsh_BCAT1-2 (f–h, n = 2 independent experiments in HL-60, n = 3 in SKM-1, n = 2 in MOLM-13), shEGLN1_shNT and shEGLN1_shBCAT1-2 (f, n = 4 independent experiments in HL-60) were measured 10 days after transduction to confirm knockdown of BCAT1 in rescued cells, k, l, Western blot of indicated cell lines with stable HIF1α overexpression (k) or shEGLN1 (l). The experiment was repeated three times with similar results. For gel source data, see Supplementary Fig. 1. m, Gene expression (mean) of BCAT1 in BCAT1-OE cell lines 3 days after induction (–DOX and +DOX) by RT–qPCR relative to the expression of ACTB and GAPDH (n = 3 independent cultures).
Extended Data Figure 7 BCAT1 expression and survival in patients with or without mutations in IDH or TET2.
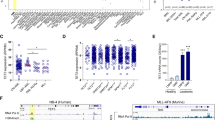
a, Normalized BCAT1 expression in WT (IDHWTTET2WT), IDHmut, or TET2mut of CN-AML samples of the TCGA (WT: n = 52, IDHmut: n = 19, TET2mut: n = 9), GSE16432 (WT: n = 80, IDHmut: n = 35, TET2mut: n = 31), and GSE14468 datasets (WT: n = 147, IDHmut: n = 40). Boxes represent median and 25th–75th percentiles, whiskers are minimum to maximum; unpaired, two-sided t-test. b, Western blot of BCAT1 protein expression in primary AML samples showing IDH1/IDH2 mutational status. The experiment was repeated twice with similar results. For gel source data, see Supplementary Fig. 1. c, Kaplan–Meier plots showing overall survival. TCGA dataset CN-AML, IDHWTTET2WT (left, n = 52) or IDHmut or TET2mut (right, n = 28) are stratified for BCAT1 expression above (red line) or below (blue line) the median. Two-sided log-rank (Mantel–Cox) test.
Extended Data Figure 8 GSEA of IDHWTTET2WTBCAT1high versus IDHWTTET2WTBCAT1low AML samples, BCAT1 expression in paired diagnosis/relapse AML samples, and DNA hypermethylation in AML samples and BCAT1-OE cells.
a, Overlap of hypermethylated probes (P < 0.001, Δmeth >0.25) between IDHWTTET2WTDNMT3AWTBCAT1Q1 (n = 6) versus IDHWTTET2WTDNMT3AWTBCAT1Q4 (n = 6) patients, and IDHmut (n = 13) versus IDHWTTET2WTDNMT3AWTBCAT1Q4 (n = 6) patients, with CN-AML in the TCGA dataset. b, GSEA plots of enrichment of CN-AML from GSE14468 (n = 146), GSE16432 (n = 80), and TCGA (n = 46) patient cohorts comparing IDHWTTET2WTBCAT1high with IDHWTTET2WTBCAT1low expression profiles. ES, enrichment score. c, BCAT1 expression levels (reads per kilobase per million reads) in paired diagnosis/relapse AML samples (n = 19) from GSE83533 (ref. 40). Two-sided paired t-test. d, Histogram of differentially methylated 5mC probes (Δmeth < −0.025 or > 0.025) in MOLM-13 BCAT1-OE cells +DOX versus –DOX cultured for 10 (n = 2 independent cultures) or 20 weeks (n = 3 independent cultures). e, Paired analysis of DNA methylation for 10 or 20 weeks of BCAT1-OE cells. The common probes from a, which were, in addition, more methylated (Δ β > 0) after 20 weeks of BCAT-OE, are shown (n = 779 in HL-60, n = 834 in MOLM-13). Two-sided, paired t-test.
Supplementary information
Supplementary Figure 1
This file contains uncropped scans of western blots with size marker indication. (PDF 1466 kb)
Supplementary Table 1
This file contains patient characteristics and leukaemia-initiating potential of sorted populations after transplantation into NSG mice. (PDF 1273 kb)
Supplementary Data 1
This file contains differentially expressed proteins between LSC and non-LSC populations for all six patients used for proteomic analysis. (XLSX 2782 kb)
Supplementary Data 2
This file contains GSEA for c2cp and hallmark gene sets comparing LSC to non-LSC populations. (XLSX 75 kb)
Supplementary Data 3
Differentially methylated CpGs comparing IDHmut to IDHwtBCAT1Q4 and IDHwtBCAT1Q1 to IDHwtBCAT1Q4 AML patients in the TCGA dataset and overlap to BCAT1-overexpressing AML cell lines. (XLSX 714 kb)
Supplementary Data 4
This file indicates sampling and labelling strategies and lists all identified proteins and their differential expression in LSC compared to non-LSC. (XLSX 5993 kb)
Source data
Rights and permissions
About this article
Cite this article
Raffel, S., Falcone, M., Kneisel, N. et al. BCAT1 restricts αKG levels in AML stem cells leading to IDHmut-like DNA hypermethylation. Nature 551, 384–388 (2017). https://doi.org/10.1038/nature24294
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24294
This article is cited by
-
Iron metabolism: backfire of cancer cell stemness and therapeutic modalities
Cancer Cell International (2024)
-
Metabolic priming by multiple enzyme systems supports glycolysis, HIF1α stabilisation, and human cancer cell survival in early hypoxia
The EMBO Journal (2024)
-
Amino acid metabolism in tumor biology and therapy
Cell Death & Disease (2024)
-
High expression of BCAT1 sensitizes AML cells to PARP inhibitor by suppressing DNA damage response
Journal of Molecular Medicine (2024)
-
The role of bone marrow microenvironment (BMM) cells in acute myeloid leukemia (AML) progression: immune checkpoints, metabolic checkpoints, and signaling pathways
Cell Communication and Signaling (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.