Abstract
Interleukin-1 receptor 8 (IL-1R8, also known as single immunoglobulin IL-1R-related receptor, SIGIRR, or TIR8) is a member of the IL-1 receptor (ILR) family with distinct structural and functional characteristics, acting as a negative regulator of ILR and Toll-like receptor (TLR) downstream signalling pathways and inflammation1. Natural killer (NK) cells are innate lymphoid cells which mediate resistance against pathogens and contribute to the activation and orientation of adaptive immune responses2,3,4. NK cells mediate resistance against haematopoietic neoplasms but are generally considered to play a minor role in solid tumour carcinogenesis5,6,7. Here we report that IL-1R8 serves as a checkpoint for NK cell maturation and effector function. Its genetic blockade unleashes NK-cell-mediated resistance to hepatic carcinogenesis, haematogenous liver and lung metastasis, and cytomegalovirus infection.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
References
Garlanda, C., Dinarello, C. A. & Mantovani, A. The interleukin-1 family: back to the future. Immunity 39, 1003–1018 (2013)
Di Santo, J. P. Natural killer cell developmental pathways: a question of balance. Annu. Rev. Immunol. 24, 257–286 (2006)
Vivier, E. et al. Innate or adaptive immunity? The example of natural killer cells. Science 331, 44–49 (2011)
Bellora, F. et al. Human NK cells and NK receptors. Immunol. Lett. 161, 168–173 (2014)
Guillerey, C., Huntington, N. D. & Smyth, M. J. Targeting natural killer cells in cancer immunotherapy. Nat. Immunol. 17, 1025–1036 (2016)
Stojanovic, A. & Cerwenka, A. Natural killer cells and solid tumors. J. Innate Immun. 3, 355–364 (2011)
Gismondi, A., Stabile, H., Nisti, P. & Santoni, A. Effector functions of natural killer cell subsets in the control of hematological malignancies. Front. Immunol. 6, 567 (2015)
Gulen, M. F. et al. The receptor SIGIRR suppresses Th17 cell proliferation via inhibition of the interleukin-1 receptor pathway and mTOR kinase activation. Immunity 32, 54–66 (2010)
Nold-Petry, C. A. et al. IL-37 requires the receptors IL-18Rα and IL-1R8 (SIGIRR) to carry out its multifaceted anti-inflammatory program upon innate signal transduction. Nat. Immunol. 16, 354–365 (2015)
Garlanda, C., Riva, F., Bonavita, E. & Mantovani, A. Negative regulatory receptors of the IL-1 family. Semin. Immunol. 25, 408–415 (2013)
Cooper, M. A. et al. Human natural killer cells: a unique innate immunoregulatory role for the CD56bright subset. Blood 97, 3146–3151 (2001)
Chiossone, L. et al. Maturation of mouse NK cells is a 4-stage developmental program. Blood 113, 5488–5496 (2009)
Kim, S. et al. In vivo developmental stages in murine natural killer cell maturation. Nat. Immunol. 3, 523–528 (2002)
Takeda, K. et al. Defective NK cell activity and Th1 response in IL-18-deficient mice. Immunity 8, 383–390 (1998)
Ganal, S. C. et al. Priming of natural killer cells by nonmucosal mononuclear phagocytes requires instructive signals from commensal microbiota. Immunity 37, 171–186 (2012)
Gong, J. et al. Inhibition of Toll-like receptors TLR4 and 7 signaling pathways by SIGIRR: a computational approach. J. Struct. Biol. 169, 323–330 (2010)
Marçais, A. et al. The metabolic checkpoint kinase mTOR is essential for IL-15 signaling during the development and activation of NK cells. Nat. Immunol. 15, 749–757 (2014)
Li, C. et al. JNK MAP kinase activation is required for MTOC and granule polarization in NKG2D-mediated NK cell cytotoxicity. Proc. Natl Acad. Sci. USA 105, 3017–3022 (2008)
Peng, H. & Tian, Z. Re-examining the origin and function of liver-resident NK cells. Trends Immunol. 36, 293–299 (2015)
Naugler, W. E. et al. Gender disparity in liver cancer due to sex differences in MyD88-dependent IL-6 production. Science 317, 121–124 (2007)
Dupaul-Chicoine, J. et al. The Nlrp3 inflammasome suppresses colorectal cancer metastatic growth in the liver by promoting natural killer cell tumoricidal activity. Immunity 43, 751–763 (2015)
Lisnic´, B., Lisnic´, V. J. & Jonjic´, S. NK cell interplay with cytomegaloviruses. Curr. Opin. Virol. 15, 9–18 (2015)
Eberl, G., Colonna, M., Di Santo, J. P. & McKenzie, A. N. Innate lymphoid cells: a new paradigm in immunology. Science 348, aaa6566 (2015)
Shih, H. Y. et al. Developmental acquisition of regulomes underlies innate lymphoid cell functionality. Cell 165, 1120–1133 (2016)
Bellora, F. et al. M-CSF induces the expression of a membrane-bound form of IL-18 in a subset of human monocytes differentiating in vitro toward macrophages. Eur. J. Immunol. 42, 1618–1626 (2012)
Martín-Fontecha, A. et al. Induced recruitment of NK cells to lymph nodes provides IFN-gamma for T(H)1 priming. Nat. Immunol. 5, 1260–1265 (2004)
Garlanda, C. et al. Increased susceptibility to colitis-associated cancer of mice lacking TIR8, an inhibitory member of the interleukin-1 receptor family. Cancer Res. 67, 6017–6021 (2007)
Xiao, H. et al. The Toll-interleukin-1 receptor member SIGIRR regulates colonic epithelial homeostasis, inflammation, and tumorigenesis. Immunity 26, 461–475 (2007)
Morvan, M. G. & Lanier, L. L. NK cells and cancer: you can teach innate cells new tricks. Nat. Rev. Cancer 16, 7–19 (2016)
He, G. & Karin, M. NF-κB and STAT3 - key players in liver inflammation and cancer. Cell Res. 21, 159–168 (2011)
Garlanda, C. et al. Intestinal inflammation in mice deficient in Tir8, an inhibitory member of the IL-1 receptor family. Proc. Natl Acad. Sci. USA 101, 3522–3526 (2004)
Bushnell, B. Bbmap: a fast, accurate, splice-aware aligner (Ernest Orlando Lawrence Berkeley National Laboratory, 2014)
Martin, M. Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet.journal 17, http://dx.doi.org/10.14806/ej.17.1.200 (2011)
Dobin, A . et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013)
Gentleman, R. C. et al. Bioconductor: open software development for computational biology and bioinformatics. Genome Biol. 5, R80 (2004)
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010)
Majewski, I. J. et al. Opposing roles of polycomb repressive complexes in hematopoietic stem and progenitor cells. Blood 116, 731–739 (2010)
Ritchie, M. E. et al. limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015)
Mingozzi, F. et al. Prolonged contact with dendritic cells turns lymph node-resident NK cells into anti-tumor effectors. EMBO Mol. Med. 8, 1039–1051 (2016)
Giavazzi, R., Alessandri, G., Spreafico, F., Garattini, S. & Mantovani, A. Metastasizing capacity of tumour cells from spontaneous metastases of transplanted murine tumours. Br. J. Cancer 42, 462–472 (1980)
Wagner, M., Jonjic´, S., Koszinowski, U. H. & Messerle, M. Systematic excision of vector sequences from the BAC-cloned herpesvirus genome during virus reconstitution. J. Virol. 73, 7056–7060 (1999)
Jonjic´, S., Pavic´, I., Lucin, P., Rukavina, D. & Koszinowski, U. H. Efficacious control of cytomegalovirus infection after long-term depletion of CD8+ T lymphocytes. J. Virol. 64, 5457–5464 (1990)
Reddehase, M. J. et al. Interstitial murine cytomegalovirus pneumonia after irradiation: characterization of cells that limit viral replication during established infection of the lungs. J. Virol. 55, 264–273 (1985)
Acknowledgements
We thank N. Polentarutti, G. Benigni, M. Erreni, F. Colombo, V. Juranic´ Lisnic´ and D. Kvestak and Computational and Molecular Biology CRUK MI core facilities for technical assistance, M. Nebuloni for hepatocellular carcinoma histology, A. Doni for STED images, and F. Ficara, R. Carriero and D. Mavilio for discussions. The contributions of the European Commission (ERC project PHII-669415; FP7 project 281608 TIMER; ESA/ITN, H2020-MSCA-ITN-2015-676129), Ministero dell’Istruzione, dell’Università e della Ricerca (MIUR) (project FIRB RBAP11H2R9), Associazione Italiana Ricerca sul Cancro (AIRC IG-19014 and AIRC 5x1000-9962), Fondazione CARIPLO (project 2015-0564), European Regional Development Fund (grant KK.01.1.1.01.0006, to S.J.) and the Italian Ministry of Health are gratefully acknowledged. M.M. received a European Federation of Immunological Sciences short-term fellowship to perform viral infection experiments in the laboratory of S.Jo.
Author information
Authors and Affiliations
Contributions
E.B. and M.M. played a key role in designing and conducting most experiments and drafted the manuscript. F.R., M.B., F.G. and E.M. provided technological support in in vivo experiments. A.P., S.Ja., B.P. and G.B. contributed to the experimental design and in vivo experiments. S.Z. contributed to RNA-seq analysis. S.Jo. and A.S. contributed to the experimental design and supervision of the study. C.G. and A.M. contributed to the experimental design and supervision of the study, and suggested the role of IL-1R8 as a novel checkpoint inhibitor of NK cells.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks M. Karin, M. Smyth and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Expression of IL-1R8 in human and mouse NK cells.
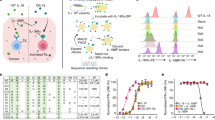
a, b, IL1R8 mRNA expression in human primary NK cells, compared with T and B cells, neutrophils, monocytes and in vitro-derived macrophages (a) and in human primary NK cell maturation stages (CD56brCD16−, CD56brCD16+, CD56dimCD16+), and in the CD56dimCD16− subset (b). c, Representative FACS plot of human NK cell subsets and histograms of IL-1R8 expression in NK cell subsets. d, IL-1R8 protein expression in human bone marrow precursors and mature cells. e, ILR family member (Il1r1, Il1r2, Il1r3, Il1r4, Il1r5, Il1r6, Il1r8) mRNA expression in mouse primary NK cells isolated from the spleen. f, IL-1R8 protein expression in mouse NK cells by confocal microscopy. Scale bars, 10 μm. g, Representative FACS plot of mouse NK cell subsets. a, b, d, *P < 0.05, **P < 0.01, ***P < 0.001. One-way ANOVA. Mean ± s.e.m. a, n = 6 (NK and B cells) or n = 4 donors; b, n = 5 donors; d, n = 4 donors; e, n = 2 mice; f, representative images out of four collected per group. a, b, d–f, One experiment performed.
Extended Data Figure 2 Phenotypic analysis of Il1r8−/− NK cells.
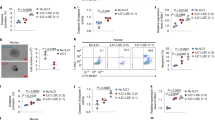
a, b, Representative plot of fluorescence-activated cell sorting of mouse NK cell subsets in Il1r8+/+ and Il1r8−/− mice (a) and histograms of KLRG1 expression in NK cells (b). c, d, NK absolute number and NK cell subsets (DN, CD11blow, DP and CD27low) in bone marrow, spleen and blood of Il1r8+/+ and Il1r8−/− newborn mice at 2 (c) and 3 (d) weeks of age. e, Frequency of bone marrow precursors in Il1r8+/+ and Il1r8−/− mice. f, NKG2D, DNAM-1 and LY49H expression in peripheral NK cells and NK cell subsets of Il1r8+/+ and Il1r8−/− mice. g, Frequency of splenic Perforin+ NK cell subsets upon stimulation in Il1r8+/+ and Il1r8−/− mice. h, i, Peripheral NK cell absolute number (h) and CD27low NK cell frequency (i) in bone marrow chimaeric mice upon reconstitution (9 weeks). j, k, Peripheral NK cell (j) and NK cell subset (k) frequency in competitive chimaeric mice transplanted with 50% of Il1r8+/+ CD45.1 cells and 50% of Il1r8−/− CD45.2 cells upon reconstitution (9 weeks). Upon reconstitution, a defective engraftment (12% instead of 50% engraftment) of Il1r8−/− stem cells was observed in competitive conditions. l, IFNγ production by Il1r8+/+ and Il1r8−/− NK cells upon co-culture with LPS- or CpG-primed Il1r8+/+ and Il1r8−/− dendritic cells. c–l, *P < 0.05, **P < 0.01, ***P < 0.001 between selected relevant comparisons, two-tailed unpaired Student’s t-test. Centre values and error bars, mean ± s.e.m. At least five animals per group were used. c, d, Three pooled experiments; e–l, one experiment was performed.
Extended Data Figure 3 Mechanism of IL-1R8-dependent regulation of NK cells.
a, Splenic CD27low NK cell frequency in wild-type, Il1r8−/−, Il18−/− and Il18−/−/Il1r8−/− mice. b, Peripheral CD27low NK cell frequency in wild-type, Il1r8−/−, Il1r1−/− and Il1r8−/−Il1r1−/− mice (left) and IFNγ production by splenic NK cells after IL-12 and IL-1β or IL-18 stimulation (right). c, d, Splenic CD27low NK cell frequency in Il1r8+/+ and Il1r8−/− mice upon commensal flora depletion (c) and breeding in co-housing conditions (d). e, STED microscopy of human NK cells stimulated with IL-18. Magnification bar, 2 μm. a–d, *P < 0.05, **P < 0.01, ***P < 0.001 between selected relevant comparisons, two-tailed unpaired Student’s t-test; Centre values and error bars, mean ± s.e.m. a, n = 3, 5, or 6 mice; at least five animals per group were used (b–d). a–d, One experiment was performed. e, Representative images out of three collected from two donors.
Extended Data Figure 4 RNA-seq analysis of Il1r8+/+ and Il1r8−/− NK cells.
Metascape analysis of enriched gene pathways of resting and IL-18-activated Il1r8+/+ and Il1r8−/− NK cells. See also Supplementary Table 1 and data deposited in the NCBI Gene Expression Omnibus under accession number GSE105043.
Extended Data Figure 5 NK-cell-mediated resistance to hepatocellular carcinoma and metastasis in IL-1R8-deficient mice.
a, Macroscopic score of liver lesions in female Il1r8+/+ and Il1r8−/− mice 6, 10 and 12 months after diethylnitrosamine (DEN) injection. b, Incidence of hepatocellular carcinoma in Il1r8+/+ and Il1r8−/− female and male mice. c, Frequency of IFNγ+ NK cells in spleen of Il1r8+/+ and Il1r8−/− tumour-bearing mice. d, Macroscopic score of liver lesions in female Il1r8+/+ and Il1r8−/− mice upon NK cell depletion. e, 2-Deoxyglucosone (2-DG) quantification in lungs of Il1r8+/+ and Il1r8−/− tumour-bearing mice upon NK cell depletion. f, Primary tumour growth in Il1r8+/+ and Il1r8−/− mice (25 days after MN/MCA1 cell line injection). g, Number of lung metastases in Il1r8+/+ and Il1r8−/− MN/MCA1 sarcoma-bearing mice upon IFNγ or IL-18 neutralization. h, Volume of lung metastases in Il1r8+/+ and Il1r8−/− MN/MCA1-bearing mice upon depletion of IL-17A or CD4+/CD8+ cells. i, Number of lung metastases in Il1r8+/+ and Il1r8−/−, Il1r1−/−, Il1r1−/−/Il1r8−/− MN/MCA1-bearing mice. j, Number of liver metastases in Il1r8+/+, Il1r8−/−, Il18−/−, Il18−/−Il1r8−/− MC38 colon carcinoma-bearing mice. k, Il1r8+/+ and Il1r8−/− NK cell absolute number 3 or 7 days after adoptive transfer. l, In vivo Il1r8+/+ and Il1r8−/− NK cell proliferation 3 days after adoptive transfer. m, Ex vivo IFNγ production and degranulation upon 4 h stimulation with PMA-ionomycin, IL-12 and IL-18 in adoptively transferred Il1r8+/+ and Il1r8−/− NK cells. n, Volume of lung metastases in Il1r8+/+ MN/MCA1 sarcoma-bearing mice after adoptive transfer of Il1r8+/+ and Il1r8−/− NK cells. a, c–e, g–j, m–n, *P < 0.05, **P < 0.01, ***P < 0.001 between selected relevant comparisons, two-tailed unpaired Student’s t-test or Mann–Whitney U-test. #P < 0.05, ##P < 0.01, Kruskal–Wallis and Dunn’s multiple comparison test. Centre values and error bars, mean ± s.e.m. a, n = 9, 10, 11, 18, 21 mice; b, n = 8–21 mice; c, n = 6 mice; d, n = 10, 12, 13 mice; e, n = 4 (Il1r8−/− isotype) or n = 5; f, n = 10; g, n = 6, 7, 9, 10 mice; h, n = 5, 6, 12 mice; i, n = 6, 8, 10 mice; j, n = 4, 5, 7 mice; k, l, m, n = 3 mice; n, n = 9, 10, 12 mice. Representative experiment out of three (a, b), 2 (d), 6 (f), or one (c, e, g–n) experiments performed. NT, not treated.
Extended Data Figure 6 NK-cell-mediated anti-viral resistance in IL-1R8-deficient mice.
Cytokine serum levels in Il1r8+/+ and Il1r8−/− infected mice (1.5 and 4.5 days after infection). *P < 0.05, **P < 0.01, ***P < 0.001, unpaired Student’s t-test. Centre values and error bars, mean ± s.e.m.; n = 5 mice. One experiment was performed.
Supplementary information
Supplementary Figure
This file shows the murine splenic NK cell gating strategy.
Supplementary Table 1
Gene expression profile of Il1r8+/+ and Il1r8-/-i> NK cells.
Rights and permissions
About this article
Cite this article
Molgora, M., Bonavita, E., Ponzetta, A. et al. IL-1R8 is a checkpoint in NK cells regulating anti-tumour and anti-viral activity. Nature 551, 110–114 (2017). https://doi.org/10.1038/nature24293
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24293
This article is cited by
-
Locally sourced: site-specific immune barriers to metastasis
Nature Reviews Immunology (2023)
-
SIGIRR deficiency contributes to CD4 T cell abnormalities by facilitating the IL1/C/EBPβ/TNF-α signaling axis in rheumatoid arthritis
Molecular Medicine (2022)
-
Macrophages as tools and targets in cancer therapy
Nature Reviews Drug Discovery (2022)
-
The unique role of innate lymphoid cells in cancer and the hepatic microenvironment
Cellular & Molecular Immunology (2022)
-
IL-37 alleviates Coxsackievirus B3-induced viral myocarditis via inhibiting NLRP3 inflammasome-mediated pyroptosis
Scientific Reports (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.