Abstract
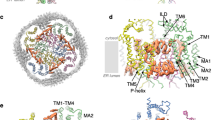
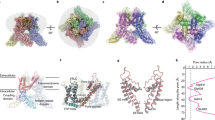
Transient receptor potential mucolipin 1 (TRPML1) is a cation channel located within endosomal and lysosomal membranes. Ubiquitously expressed in mammalian cells1,2, its loss-of-function mutations are the direct cause of type IV mucolipidosis, an autosomal recessive lysosomal storage disease3,4,5,6. Here we present the single-particle electron cryo-microscopy structure of the mouse TRPML1 channel embedded in nanodiscs. Combined with mutagenesis analysis, the TRPML1 structure reveals that phosphatidylinositol-3,5-bisphosphate (PtdIns(3,5)P2) binds to the N terminus of the channel—distal from the pore—and the helix–turn–helix extension between segments S2 and S3 probably couples ligand binding to pore opening. The tightly packed selectivity filter contains multiple ion-binding sites, and the conserved acidic residues form the luminal Ca2+-blocking site that confers luminal pH and Ca2+ modulation on channel conductance. A luminal linker domain forms a fenestrated canopy atop the channel, providing several luminal ion passages to the pore and creating a negative electrostatic trap, with a preference for divalent cations, at the luminal entrance. The structure also reveals two equally distributed S4–S5 linker conformations in the closed channel, suggesting an S4–S5 linker-mediated PtdInsP2 gating mechanism among TRPML channels7,8.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Cheng, X., Shen, D., Samie, M. & Xu, H. Mucolipins: intracellular TRPML1-3 channels. FEBS Lett. 584, 2013–2021 (2010)
Dong, X. P. et al. The type IV mucolipidosis-associated protein TRPML1 is an endolysosomal iron release channel. Nature 455, 992–996 (2008)
Bargal, R. et al. Identification of the gene causing mucolipidosis type IV. Nat. Genet. 26, 118–123 (2000)
Bassi, M. T. et al. Cloning of the gene encoding a novel integral membrane protein, mucolipidin—and identification of the two major founder mutations causing mucolipidosis type IV. Am. J. Hum. Genet. 67, 1110–1120 (2000)
Nilius, B., Owsianik, G., Voets, T. & Peters, J. A. Transient receptor potential cation channels in disease. Physiol. Rev. 87, 165–217 (2007)
Sun, M. et al. Mucolipidosis type IV is caused by mutations in a gene encoding a novel transient receptor potential channel. Hum. Mol. Genet. 9, 2471–2478 (2000)
Dong, X. P. et al. PI(3,5)P2 controls membrane trafficking by direct activation of mucolipin Ca2+ release channels in the endolysosome. Nat. Commun. 1, 38 (2010)
Feng, X. et al. Drosophila TRPML forms PI(3,5)P2-activated cation channels in both endolysosomes and plasma membrane. J. Biol. Chem. 289, 4262–4272 (2014)
Chandra, M. et al. A role for the Ca2+ channel TRPML1 in gastric acid secretion, based on analysis of knockout mice. Gastroenterology 140, 857–867.e1 (2011)
Cheng, X. et al. The intracellular Ca2+ channel MCOLN1 is required for sarcolemma repair to prevent muscular dystrophy. Nat. Med. 20, 1187–1192 (2014)
Puertollano, R. & Kiselyov, K. TRPMLs: in sickness and in health. Am. J. Physiol. Renal Physiol. 296, F1245–F1254 (2009)
Shen, D. et al. Lipid storage disorders block lysosomal trafficking by inhibiting a TRP channel and lysosomal calcium release. Nat. Commun. 3, 731 (2012)
Vergarajauregui, S. & Puertollano, R. Two di-leucine motifs regulate trafficking of mucolipin-1 to lysosomes. Traffic 7, 337–353 (2006)
Zhang, X., Li, X. & Xu, H. Phosphoinositide isoforms determine compartment-specific ion channel activity. Proc. Natl Acad. Sci. USA 109, 11384–11389 (2012)
Xu, H., Delling, M., Li, L., Dong, X. & Clapham, D. E. Activating mutation in a mucolipin transient receptor potential channel leads to melanocyte loss in varitint-waddler mice. Proc. Natl Acad. Sci. USA 104, 18321–18326 (2007)
Venkatachalam, K., Wong, C. O. & Zhu, M. X. The role of TRPMLs in endolysosomal trafficking and function. Cell Calcium 58, 48–56 (2015)
Lee, J. H. et al. Presenilin 1 maintains lysosomal Ca2+ homeostasis via TRPML1 by regulating vATPase-mediated lysosome acidification. Cell Reports 12, 1430–1444 (2015)
Kilpatrick, B. S., Yates, E., Grimm, C., Schapira, A. H. & Patel, S. Endo-lysosomal TRP mucolipin-1 channels trigger global ER Ca2+ release and Ca2+ influx. J. Cell Sci. 129, 3859–3867 (2016)
Cao, Q. et al. BK channels alleviate lysosomal storage diseases by providing positive feedback regulation of lysosomal Ca2+ release. Dev. Cell 33, 427–441 (2015)
Chen, C. C. et al. A small molecule restores function to TRPML1 mutant isoforms responsible for mucolipidosis type IV. Nat. Commun. 5, 4681 (2014)
Rosenthal, P. B. & Henderson, R. Optimal determination of particle orientation, absolute hand, and contrast loss in single-particle electron cryomicroscopy. J. Mol. Biol. 333, 721–745 (2003)
Li, M. et al. Structural basis of dual Ca2+/pH regulation of the endolysosomal TRPML1 channel. Nat. Struct. Mol. Biol. 24, 205–213 (2017)
Liao, M., Cao, E., Julius, D. & Cheng, Y. Structure of the TRPV1 ion channel determined by electron cryo-microscopy. Nature 504, 107–112 (2013)
Gao, Y., Cao, E., Julius, D. & Cheng, Y. TRPV1 structures in nanodiscs reveal mechanisms of ligand and lipid action. Nature 534, 347–351 (2016)
Shen, P. S. et al. The structure of the polycystic kidney disease channel PKD2 in lipid nanodiscs. Cell 167, 763–773.e11 (2016)
Nilius, B. & Owsianik, G. The transient receptor potential family of ion channels. Genome Biol. 12, 218 (2011)
Grimm, C. et al. A helix-breaking mutation in TRPML3 leads to constitutive activity underlying deafness in the varitint-waddler mouse. Proc. Natl Acad. Sci. USA 104, 19583–19588 (2007)
Kim, H. J. et al. Gain-of-function mutation in TRPML3 causes the mouse varitint-waddler phenotype. J. Biol. Chem. 282, 36138–36142 (2007)
Nagata, K. et al. The varitint-waddler (Va) deafness mutation in TRPML3 generates constitutive, inward rectifying currents and causes cell degeneration. Proc. Natl Acad. Sci. USA 105, 353–358 (2008)
Dong, X. P. et al. Activating mutations of the TRPML1 channel revealed by proline-scanning mutagenesis. J. Biol. Chem. 284, 32040–32052 (2009)
Morales-Perez, C. L., Noviello, C. M. & Hibbs, R. E. Manipulation of subunit stoichiometry in heteromeric membrane proteins. Structure 24, 797–805 (2016)
Zheng, S. Q. et al. MotionCor2: anisotropic correction of beam-induced motion for improved cryo-electron microscopy. Nat. Methods 14, 331–332 (2017)
Zhang, K. Gctf: Real-time CTF determination and correction. J. Struct. Biol. 193, 1–12 (2016)
Scheres, S. H. RELION: implementation of a Bayesian approach to cryo-EM structure determination. J. Struct. Biol. 180, 519–530 (2012)
Bai, X. C., Rajendra, E., Yang, G., Shi, Y. & Scheres, S. H. Sampling the conformational space of the catalytic subunit of human γ-secretase. eLife 4, e11182 (2015)
Kucukelbir, A., Sigworth, F. J. & Tagare, H. D. Quantifying the local resolution of cryo-EM density maps. Nat. Methods 11, 63–65 (2014)
Emsley, P., Lohkamp, B., Scott, W. G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D 66, 486–501 (2010)
Afonine, P. V., Headd, J. J., Terwilliger, T. C. & Adams, P. D. New tool: phenix.real_space_refine. Computational Crystallography Newsletter 4, 43–44 (2013)
Adams, P. D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D 66, 213–221 (2010)
Amunts, A. et al. Structure of the yeast mitochondrial large ribosomal subunit. Science 343, 1485–1489 (2014)
Chen, V. B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D 66, 12–21 (2010)
Smart, O. S., Neduvelil, J. G., Wang, X., Wallace, B. A. & Sansom, M. S. HOLE: a program for the analysis of the pore dimensions of ion channel structural models. J. Mol. Graph. 14, 354–360, 376 (1996)
The PyMOL Molecular Graphics System v.1.8 (Schrödinger, LLC, 2015)
Pettersen, E. F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004)
Grimm, C. et al. Small molecule activators of TRPML3. Chem. Biol. 17, 135–148 (2010)
Acknowledgements
We thank N. Nguyen for manuscript preparation. Single-particle cryo-EM data were collected at the University of Texas Southwestern (UTSW) Medical Center Cryo-Electron Microscopy Facility. We thank D. Nicastro and Z. Chen for support in facility access and data acquisition. Negatively stained sample screening was performed at the UTSW Electron Microscopy core. This work was supported in part by the Howard Hughes Medical Institute (Y.J.) and by grants from the National Institutes of Health (GM079179 to Y.J.; NS062792 and AR060837 to H.X.) and the Welch Foundation (grant I-1578 to Y.J.). X.B. is supported by the Cancer Prevention and Research Initiative of Texas and Virginia Murchison Linthicum Scholar in Medical Research fund.
Author information
Authors and Affiliations
Contributions
Q.C. and J.S. prepared the samples; Q.C., J.S., J.G. and X.B. performed data acquisition, image processing and structure determination; W.Z. performed electrophysiology; H.X. provided DNA materials and constructive advice; all authors participated in research design, data analysis, and manuscript preparation.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks S. Hansen, C. Ulens and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Publisher's note: Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data figures and tables
Extended Data Figure 1 Ligand activation of TRPML1 overexpressed in HEK293 cells.
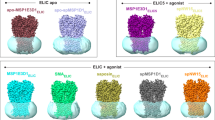
a, Macroscopic currents of plasma membrane-localized TRPML1–4A in an inside-out patch in the presence and absence of ligands in bath solution (cytosolic). TRPML1 can be activated by PtdIns(3,5)P2 and mucolipin synthetic agonist 1 (ML-SA1), yielding inwardly rectified cation currents. PtdIns(3,5)P2 and ML-SA1 activations are synergistic, indicating non-overlapping activation sites between the two ligands. b, Macroscopic currents of TRPML1–4A in the whole-cell configuration. The pipette (cytosolic) solution contained 50 μM PtdIns(3,5)P2 (black trace). Addition of 10 μM ML-SA1 into the bath solution (extracellular/luminal side) yielded a much larger current, suggesting that ML-SA1 can also activate the channel from the luminal side. c, Sample traces of single-channel currents recorded at −120 mV in an inside-out excised patch showing PtdIns(3,5)P2 and ML-SA1 activation. The patch contained multiple channels (N ≥ 5). N, total number of channels; Po, single-channel open probability; NPo, total open probability (the number of channels multiplied by single-channel open probability); C, closed state; O, open state, with the number indicating open events from multiple channels. d, Sample traces of single-channel currents recorded at −120 mV in an inside-out excised patch showing PtdIns(4,5)P2 (in bath solution) inhibition of TRPML1. e, Single-channel recordings of N-terminal truncation mutation of TRPML1–4A in an inside-out patch. Deletion of the poly-basic domain abolishes PtdIns(3,5)P2 activation, confirming its participation in PtdInsP2 binding; the ML-SA1 activation remains intact in this mutant, confirming distinct activation sites between PtdInsP2 and the small-molecule agonist.
Extended Data Figure 2 Structure determination of mouse TRPML1 in nanodiscs.
a, Representative micrograph of TRPML1 in a nanodisc. b, Two-dimensional class averages. c, Euler angle distribution of particles used in the final three-dimensional reconstruction, with the heights of the cylinders corresponding to the number of particles. d, Gold-standard FSC curves of the final 3D reconstructions. e, Final density maps coloured by local resolution.
Extended Data Figure 4 Data collection, structure refinement and model validation.
a, Data collection and model refinement statistics. b, FSC curves for cross-validation between the maps and the models. Curves for model versus summed map in blue (full), for model versus half map in green (work), and for model versus half map not used for refinement in black (free).
Extended Data Figure 5 Sample electron microscopy density maps (blue mesh) for various parts of the channel.
The maps are low-pass filtered to 3.59 Å and sharpened with a temperature factor of −120 Å2.
Extended Data Figure 6 Sequence alignment of TRPML channels.
Secondary structure assignments are based on the mouse TRPML1 structure. Purple crosses mark the basic residues important for PtdInsP2 binding and the blue dot marks the location of V432P mutation in TRPML1.
Extended Data Figure 7 Luminal Ca2+ and pH modulation of TRPML1.
a, Sample traces (I–V curves) of whole-cell currents from the constitutively active TRPML1(V432P) mutant recorded with various luminal (bath solution) Ca2+ concentrations [Ca2+] and pH values. Inset shows normalized channel currents at −120 mV. Data are mean ± s.e.m. of five measurements. 0 mM [Ca2+] here refers to nominally Ca2+-free medium in the recording. b, Sample traces of whole-cell currents of the TRPML1(V432P/D472N) mutant recorded at various luminal (bath solution) [Ca2+] and pH values. Inset shows normalized channel currents at −120 mV. Data are mean ± s.e.m. of five measurements. c, Luminal [Ca2+]-dependent blockage of inward currents in TRPML1(V432P) and TRPML1(V432P/D472N) measured at −120 mV and pH 7.4. In summary, 1 mM Ca2+—close to the lysosomal Ca2+ concentration—can markedly reduce the channel current; lowering the pH can partially alleviate Ca2+ blockage, probably by protonating the Ca2+-binding acidic residues; in the absence of Ca2+, however, lower pH by itself has an inhibitory effect on channel conductance. Neutralizing Asp472 with Asn diminishes the luminal Ca2+ blocking.
Extended Data Figure 8 Structural comparison between the S1–S2 linker domains of TRPML1 and PKD2, a member of the TRPP family.
a, Side view of TRPML1 luminal linker domain atop the channel with open side windows. The front subunit is highlighted in blue. b, Side view of PKD2 polycystin domain. The polycystin domain has an extra hairpin loop (red) that clogs the side window, making the central hole the only extracellular passage to the filter. There is an extra helix–turn motif (magenta) between S3 and S4 in PKD2 that extends upright and provides extra contact between the polycystin domain and the transmembrane domain. c, The luminal pore loop (magenta) between α1 and α2 points downwards in TRPML1, generating a funnel-shaped central hole with a constriction of 12 Å. d, The luminal pore loop (red) in PKD2 points upwards and generates a central hole with an inverted funnel shape. The front and rear subunits are removed in panels c and d for clarity.
Supplementary information
Rights and permissions
About this article
Cite this article
Chen, Q., She, J., Zeng, W. et al. Structure of mammalian endolysosomal TRPML1 channel in nanodiscs. Nature 550, 415–418 (2017). https://doi.org/10.1038/nature24035
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature24035
This article is cited by
-
Unexpected inhibition of the lipid kinase PIKfyve reveals an epistatic role for p38 MAPKs in endolysosomal fission and volume control
Cell Death & Disease (2024)
-
Nanoscale one-dimensional close packing of interfacial alkali ions driven by water-mediated attraction
Nature Nanotechnology (2024)
-
Regulating ion affinity and dehydration of metal-organic framework sub-nanochannels for high-precision ion separation
Nature Communications (2024)
-
TRP (transient receptor potential) ion channel family: structures, biological functions and therapeutic interventions for diseases
Signal Transduction and Targeted Therapy (2023)
-
Mutation of TRPML1 Channel and Pathogenesis of Neurodegeneration in Haimeria
Molecular Neurobiology (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.